

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0002.6
(260 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
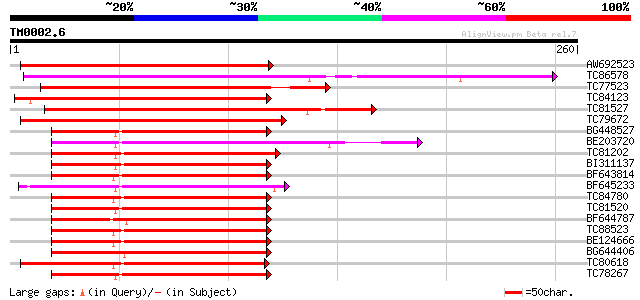
Score E
Sequences producing significant alignments: (bits) Value
AW692523 similar to GP|9954116|gb| tuber-specific and sucrose-re... 199 7e-52
TC86578 similar to PIR|C85440|C85440 myb-related protein [import... 186 1e-47
TC77523 homologue to GP|9954112|gb|AAG08959.1| tuber-specific an... 183 5e-47
TC84123 similar to PIR|T47712|T47712 MYB transcription factor-li... 177 4e-45
TC81527 similar to GP|9954116|gb|AAG08961.1| tuber-specific and ... 172 2e-43
TC79672 similar to GP|6979341|gb|AAF34434.1| myb-like protein {O... 137 3e-33
BG448527 similar to PIR|T51509|T51 probable transcription factor... 115 2e-26
BE203720 similar to PIR|T51666|T51 myb-related transcription fac... 114 3e-26
TC81202 homologue to GP|17380966|gb|AAL36295.1 putative transcri... 113 6e-26
BI311137 similar to GP|17380966|gb putative transcription factor... 109 9e-25
BF643814 similar to GP|19386839|db putative myb-related protein ... 109 9e-25
BF645233 similar to PIR|T03762|T03 myb-related transcription fac... 107 4e-24
TC84780 homologue to PIR|F85021|F85021 probable transcription fa... 107 4e-24
TC81520 similar to PIR|D86470|D86470 hypothetical protein AAD460... 107 5e-24
BF644787 similar to GP|11994692|db MYB-like DNA-binding domain p... 106 8e-24
TC88523 similar to PIR|T05690|T05690 myb-related transcription f... 106 8e-24
BE124666 similar to PIR|JQ0957|JQ0 myb-related protein 330 - gar... 106 1e-23
BG644406 homologue to PIR|T07398|T07 myb-related transcription f... 105 1e-23
TC80618 homologue to PIR|T06455|T06455 Myb26 protein - garden pe... 105 2e-23
TC78267 weakly similar to PIR|B85064|B85064 MYB-like protein [im... 105 2e-23
>AW692523 similar to GP|9954116|gb| tuber-specific and sucrose-responsive
element binding factor {Solanum tuberosum}, partial
(39%)
Length = 660
Score = 199 bits (507), Expect = 7e-52
Identities = 93/116 (80%), Positives = 103/116 (88%)
Frame = +1
Query: 6 DSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQ 65
+S++ GG + D+VKGSWS EDETL+KLV ++G RNWSVIS IPGRSGKSCRLRWCNQ
Sbjct: 202 NSNSNGGDENGDKVKGSWSKTEDETLVKLVKENGARNWSVISNEIPGRSGKSCRLRWCNQ 381
Query: 66 LSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRR 121
LSP VQH PFTP ED+III+AHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRR
Sbjct: 382 LSPTVQHLPFTPAEDSIIIEAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRR 549
>TC86578 similar to PIR|C85440|C85440 myb-related protein [imported] -
Arabidopsis thaliana, partial (42%)
Length = 1283
Score = 186 bits (471), Expect = 1e-47
Identities = 112/259 (43%), Positives = 146/259 (56%), Gaps = 14/259 (5%)
Frame = +2
Query: 7 SSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQL 66
++N DR+KG WSP+EDE+L KLV ++GPRNWS+IS IPGRSGKSCRLRWCNQL
Sbjct: 29 NTNAPSSTKMDRIKGPWSPEEDESLTKLVDRYGPRNWSLISRAIPGRSGKSCRLRWCNQL 208
Query: 67 SPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLP 126
SP V+HR F+P ED III+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+ +
Sbjct: 209 SPQVEHRAFSPEEDNIIIRAHAQFGNKWATIARLLSGRTDNAIKNHWNSTLKRKCSSFIG 388
Query: 127 LPLASPSPSP---SPSPPLPFSMAKRPHGYELLDRYDGLMDQGYPFKRHCVEKDDEESRD 183
+P P S S P S P G +L D G+ + + + +
Sbjct: 389 SDDRDFNPQPLKRSASVGAPGS----PSGSDLSD--SGVQQTVFRPVPIRLPVETTSEPE 550
Query: 184 GVEEAETMTEEEVLASSVPTSL-----------SLFPMGEKKMVAVAEKEEKKMDMQEGL 232
V E + E++ +S+ SL S P+ V V+E+ G
Sbjct: 551 PVTVPEAVEEDDGPLTSLSLSLPGVDAAAAVLVSPAPITAVTAVTVSEEAGIGALNLSGE 730
Query: 233 MRTMMQRMIAEEVRAYFDK 251
+MQ MI EVR+Y ++
Sbjct: 731 FMAVMQEMIKNEVRSYMEE 787
>TC77523 homologue to GP|9954112|gb|AAG08959.1| tuber-specific and
sucrose-responsive element binding factor {Solanum
tuberosum}, partial (43%)
Length = 1513
Score = 183 bits (465), Expect = 5e-47
Identities = 88/133 (66%), Positives = 101/133 (75%)
Frame = +2
Query: 15 DRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRP 74
+ DR+KG WSP+EDE L KLV +HGPRNWS+IS IPGRSGKSCRLRWCNQLSP V+HR
Sbjct: 212 EMDRIKGPWSPEEDEALQKLVEKHGPRNWSIISKSIPGRSGKSCRLRWCNQLSPQVEHRA 391
Query: 75 FTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSP 134
FTP ED II+AHA GNKWATI+RLL GRTDNAIKNHWNSTL+R+ +S
Sbjct: 392 FTPEEDDTIIRAHARFGNKWATIARLLSGRTDNAIKNHWNSTLKRK--------CSSIMI 547
Query: 135 SPSPSPPLPFSMA 147
S SPPL S++
Sbjct: 548 DDSESPPLKRSVS 586
>TC84123 similar to PIR|T47712|T47712 MYB transcription factor-like protein
- Arabidopsis thaliana, partial (27%)
Length = 682
Score = 177 bits (449), Expect = 4e-45
Identities = 82/123 (66%), Positives = 92/123 (74%), Gaps = 5/123 (4%)
Frame = +1
Query: 3 GGGDSS-----NGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKS 57
G GD GG G +VKG WSP+ED L +LV Q G RNWS+I+ GI GRSGKS
Sbjct: 139 GSGDEGLAALGEGGEGSSGSKVKGPWSPEEDAILSRLVSQFGARNWSLIARGIAGRSGKS 318
Query: 58 CRLRWCNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTL 117
CRLRWCNQL P V+ +PFT ED II+ AHA+HGNKWA I+RLLPGRTDNAIKNHWNSTL
Sbjct: 319 CRLRWCNQLDPSVKRKPFTDEEDRIIVAAHAIHGNKWAAIARLLPGRTDNAIKNHWNSTL 498
Query: 118 RRR 120
RRR
Sbjct: 499 RRR 507
>TC81527 similar to GP|9954116|gb|AAG08961.1| tuber-specific and
sucrose-responsive element binding factor {Solanum
tuberosum}, partial (56%)
Length = 765
Score = 172 bits (435), Expect = 2e-43
Identities = 87/158 (55%), Positives = 106/158 (67%), Gaps = 6/158 (3%)
Frame = +2
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
DR+KG WS +ED L +LV QHG RNWS+IS I GRSGKSCRLRWCNQLSP V+HRPF+
Sbjct: 146 DRIKGPWSAEEDRILTRLVEQHGARNWSLISRYIKGRSGKSCRLRWCNQLSPTVEHRPFS 325
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPS- 135
ED II AHA +GN+WATI+RLLPGRTDNA+KNHWNSTL+RR G + +A
Sbjct: 326 SQEDETIIAAHAQYGNRWATIARLLPGRTDNAVKNHWNSTLKRRAGCGGSVTVAGGGNEG 505
Query: 136 -----PSPSPPLPFSMAKRPHGYELLDRYDGLMDQGYP 168
P PL ++ P G + DR + + D +P
Sbjct: 506 GQFTFPLEDDPLT-ALTLAPPGNGVEDREEMVGDHRFP 616
>TC79672 similar to GP|6979341|gb|AAF34434.1| myb-like protein {Oryza
sativa}, partial (46%)
Length = 1391
Score = 137 bits (346), Expect = 3e-33
Identities = 61/122 (50%), Positives = 81/122 (66%)
Frame = +2
Query: 6 DSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQ 65
D S+ + +G W P EDE L +LV Q+G +NW+ I+ + GRSGKSCRLRW NQ
Sbjct: 143 DDSSSAAEDTKSCPRGHWRPAEDEKLRQLVEQYGAQNWNSIAEKLQGRSGKSCRLRWFNQ 322
Query: 66 LSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESL 125
L P + RPFT E+ ++ AH +HGNKWA I+RL PGRTDNA+KNHW+ + R++ E
Sbjct: 323 LDPRINRRPFTEEEEERLLAAHRIHGNKWALIARLFPGRTDNAVKNHWHVIMARKQREQS 502
Query: 126 PL 127
L
Sbjct: 503 KL 508
>BG448527 similar to PIR|T51509|T51 probable transcription factor (MYB9) -
Arabidopsis thaliana, partial (40%)
Length = 633
Score = 115 bits (287), Expect = 2e-26
Identities = 52/103 (50%), Positives = 71/103 (68%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+ +EDE L+ + +HG NW +S AG+ R GKSCRLRW N L PD++ FT
Sbjct: 79 KGPWTQEEDEKLIDYINKHGHGNWGTLSKRAGL-NRCGKSCRLRWTNYLRPDIKRGKFTD 255
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ +II H+V GNKW+ I+ LPGRTDN IKN+WN+ +R++
Sbjct: 256 EEERVIINLHSVLGNKWSKIAAHLPGRTDNEIKNYWNTNIRKK 384
>BE203720 similar to PIR|T51666|T51 myb-related transcription factor MYB59
[imported] - Arabidopsis thaliana, partial (48%)
Length = 514
Score = 114 bits (286), Expect = 3e-26
Identities = 65/173 (37%), Positives = 90/173 (51%), Gaps = 3/173 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+ QED L VG G R W I+ +G+ RSGKSCRLRW N L PD++ TP
Sbjct: 22 KGPWTEQEDYKLAYFVGLFGDRRWDFIAKVSGL-SRSGKSCRLRWVNYLHPDLKRGKMTP 198
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGESLPLPLASPSPSPS 137
E+ ++++ H+ GN+W+ I+R LPGRTDN IKN+W + +R++R + S S SP
Sbjct: 199 QEERLVMELHSKWGNRWSRIARKLPGRTDNEIKNYWRTHMRKKRAQEKKHATPSISSSPD 378
Query: 138 PSPPLPFS-MAKRPHGYELLDRYDGLMDQGYPFKRHCVEKDDEESRDGVEEAE 189
S S + HG + H K+ EE V+E E
Sbjct: 379 QSSSCQSSHFSNSNHGVD----------------SHASNKESEEENYQVQEIE 489
>TC81202 homologue to GP|17380966|gb|AAL36295.1 putative transcription
factor {Arabidopsis thaliana}, partial (40%)
Length = 1506
Score = 113 bits (283), Expect = 6e-26
Identities = 55/107 (51%), Positives = 73/107 (67%), Gaps = 2/107 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG WSP+EDE L+ + +HG WS + AG+ R GKSCRLRW N L PD++ F+
Sbjct: 98 KGLWSPEEDEKLLNYITKHGHGCWSSVPKLAGLQ-RCGKSCRLRWINYLRPDLKRGAFSQ 274
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRRRGES 124
E+ +II+ HAV GN+W+ I+ LPGRTDN IKN WNS L++R +S
Sbjct: 275 QEENLIIELHAVLGNRWSQIAAQLPGRTDNEIKNLWNSCLKKRLRQS 415
Score = 32.3 bits (72), Expect = 0.19
Identities = 20/72 (27%), Positives = 37/72 (50%), Gaps = 8/72 (11%)
Frame = +2
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQ----- 71
D +G++S QE+ +++L G R WS I+A +PGR+ + W + L ++
Sbjct: 248 DLKRGAFSQQEENLIIELHAVLGNR-WSQIAAQLPGRTDNEIKNLWNSCLKKRLRQSGID 424
Query: 72 ---HRPFTPTED 80
H+P + E+
Sbjct: 425 PNTHKPLSEVEN 460
>BI311137 similar to GP|17380966|gb putative transcription factor
{Arabidopsis thaliana}, partial (35%)
Length = 761
Score = 109 bits (273), Expect = 9e-25
Identities = 52/103 (50%), Positives = 71/103 (68%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG WSP+EDE L+ + +HG WS + AG+ R GKSCRLRW N L PD++ F+
Sbjct: 187 KGLWSPEEDEKLLNYITKHGHGCWSSVPKLAGLQ-RCGKSCRLRWINYLRPDLKRGAFSE 363
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ II+ HA+ GN+W+ I+ LPGRTDN IKN WNS+L+++
Sbjct: 364 QEENSIIELHALLGNRWSQIAAQLPGRTDNEIKNLWNSSLKKK 492
Score = 37.4 bits (85), Expect = 0.006
Identities = 26/89 (29%), Positives = 45/89 (50%), Gaps = 12/89 (13%)
Frame = +1
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQ----- 71
D +G++S QE+ ++++L G R WS I+A +PGR+ + W + L ++
Sbjct: 337 DLKRGAFSEQEENSIIELHALLGNR-WSQIAAQLPGRTDNEIKNLWNSSLKKKLKQKGID 513
Query: 72 ---HRPFTPTED----TIIIQAHAVHGNK 93
H+PF+ D I IQ +V N+
Sbjct: 514 PNTHQPFSENNDNDTPNIQIQKTSVGSNE 600
>BF643814 similar to GP|19386839|db putative myb-related protein {Oryza
sativa (japonica cultivar-group)}, partial (49%)
Length = 545
Score = 109 bits (273), Expect = 9e-25
Identities = 52/103 (50%), Positives = 71/103 (68%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG+WS +EDE L+ + Q G W + +AG+ R GKSCRLRW N L PD++ FT
Sbjct: 28 KGAWSKEEDERLINYIKQQGEGCWRSLPKAAGL-ARCGKSCRLRWINYLRPDLKRGNFTH 204
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED +II HA+ GNKW+ I++ LPGRTDN IKN+WN+ ++R+
Sbjct: 205 EEDELIISLHAMVGNKWSQIAQKLPGRTDNEIKNYWNTHIKRK 333
Score = 33.1 bits (74), Expect = 0.11
Identities = 17/62 (27%), Positives = 31/62 (49%)
Frame = +1
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D +G+++ +EDE ++ L G + WS I+ +PGR+ + W + + R
Sbjct: 178 DLKRGNFTHEEDELIISLHAMVGNK-WSQIAQKLPGRTDNEIKNYWNTHIKRKLYSRGID 354
Query: 77 PT 78
PT
Sbjct: 355 PT 360
>BF645233 similar to PIR|T03762|T03 myb-related transcription factor - rice,
partial (20%)
Length = 643
Score = 107 bits (268), Expect = 4e-24
Identities = 58/127 (45%), Positives = 77/127 (59%), Gaps = 3/127 (2%)
Frame = +1
Query: 5 GDSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRW 62
GD++ GG GG+ KG W+ EDE L+ V ++G NW+ + G+ R GKSCRLRW
Sbjct: 259 GDTT-GGEGGEGGLKKGPWTAAEDEILVAHVQKYGEGNWNSVRKCTGL-ARCGKSCRLRW 432
Query: 63 CNQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR-R 121
N L PD++ T E+ II+ H GNKWA ++ LLPGRTDN IKN WN+ ++R R
Sbjct: 433 ANHLRPDLRKGAITAEEERRIIELHHKMGNKWAQMAALLPGRTDNEIKNFWNTRCKKRGR 612
Query: 122 GESLPLP 128
LP
Sbjct: 613 ANXTNLP 633
>TC84780 homologue to PIR|F85021|F85021 probable transcription factor
[imported] - Arabidopsis thaliana, partial (39%)
Length = 971
Score = 107 bits (268), Expect = 4e-24
Identities = 51/103 (49%), Positives = 71/103 (68%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG WSP+EDE L+ + ++G WS + AG+ R GKSCRLRW N L PD++ F+
Sbjct: 142 KGLWSPEEDEKLLNHITKYGHGCWSSVPKQAGLQ-RCGKSCRLRWINYLRPDLKRGTFSQ 318
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ +II+ H+V GN+W+ I+ LPGRTDN IKN WNS L+++
Sbjct: 319 EEENLIIELHSVLGNRWSQIAAQLPGRTDNEIKNLWNSCLKKK 447
Score = 32.3 bits (72), Expect = 0.19
Identities = 17/61 (27%), Positives = 33/61 (53%)
Frame = +1
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D +G++S +E+ +++L G R WS I+A +PGR+ + W + L ++ +
Sbjct: 292 DLKRGTFSQEEENLIIELHSVLGNR-WSQIAAQLPGRTDNEIKNLWNSCLKKKLRQKGID 468
Query: 77 P 77
P
Sbjct: 469 P 471
>TC81520 similar to PIR|D86470|D86470 hypothetical protein AAD46010.1
[imported] - Arabidopsis thaliana, partial (39%)
Length = 1259
Score = 107 bits (267), Expect = 5e-24
Identities = 48/103 (46%), Positives = 70/103 (67%), Gaps = 2/103 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+P+ED+ L++ + QHG +W + AG+ R GKSCRLRW N L PD++ F+
Sbjct: 95 KGPWTPEEDQKLVEHIQQHGHGSWRALPKLAGL-NRCGKSCRLRWTNYLRPDIKRGKFSQ 271
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ I+ H++ GNKW+ I+ LPGRTDN IKN WN+ L+++
Sbjct: 272 EEEQTILHLHSILGNKWSAIATHLPGRTDNEIKNFWNTHLKKK 400
Score = 32.3 bits (72), Expect = 0.19
Identities = 25/100 (25%), Positives = 42/100 (42%)
Frame = +2
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D +G +S +E++T++ L G + WS I+ +PGR+ + W L + F
Sbjct: 245 DIKRGKFSQEEEQTILHLHSILGNK-WSAIATHLPGRTDNEIKNFWNTHLKKKLIQMGFD 421
Query: 77 PTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNST 116
P H + +TI LL I +H N +
Sbjct: 422 P-------MTHQPRTDLVSTIPYLLALANMTEIIDHQNQS 520
>BF644787 similar to GP|11994692|db MYB-like DNA-binding domain protein
{Arabidopsis thaliana}, partial (53%)
Length = 682
Score = 106 bits (265), Expect = 8e-24
Identities = 49/103 (47%), Positives = 70/103 (67%), Gaps = 2/103 (1%)
Frame = +3
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPG--RSGKSCRLRWCNQLSPDVQHRPFTP 77
KG W+ +ED L+ V HG W+ + A + G R+GKSCRLRW N L PD++ TP
Sbjct: 234 KGPWTAEEDRLLIDYVRLHGEGRWNSV-ARLTGLKRNGKSCRLRWVNYLRPDLKRGQITP 410
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E++II++ HA GN+W+TI+R LPGRTDN IKN+W + +++
Sbjct: 411 QEESIILELHARWGNRWSTIARSLPGRTDNEIKNYWRTHFKKK 539
>TC88523 similar to PIR|T05690|T05690 myb-related transcription factor MYB4
- Arabidopsis thaliana, partial (54%)
Length = 1292
Score = 106 bits (265), Expect = 8e-24
Identities = 50/103 (48%), Positives = 71/103 (68%), Gaps = 2/103 (1%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG+W+ +ED+ L+ + HG W + +AG+ R GKSCRLRW N L PD++ FT
Sbjct: 310 KGAWTKEEDDRLISYIRAHGEGCWRSLPKAAGLL-RCGKSCRLRWINYLRPDLKRGNFTE 486
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED +II+ H++ GNKW+ I+ LPGRTDN IKN+WN+ +RR+
Sbjct: 487 EEDELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRK 615
Score = 34.3 bits (77), Expect = 0.049
Identities = 17/61 (27%), Positives = 33/61 (53%)
Frame = +1
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D +G+++ +EDE ++KL G + WS+I+ +PGR+ + W + + +R
Sbjct: 460 DLKRGNFTEEEDELIIKLHSLLGNK-WSLIAGRLPGRTDNEIKNYWNTHIRRKLLNRGID 636
Query: 77 P 77
P
Sbjct: 637 P 639
>BE124666 similar to PIR|JQ0957|JQ0 myb-related protein 330 - garden
snapdragon, partial (48%)
Length = 536
Score = 106 bits (264), Expect = 1e-23
Identities = 50/103 (48%), Positives = 70/103 (67%), Gaps = 2/103 (1%)
Frame = +3
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG+W+ +EDE L+ + HG W + +AG+ R GKSCRLRW N L PD++ FT
Sbjct: 66 KGAWTKEEDERLINYIKLHGEGCWRSLPKAAGLL-RCGKSCRLRWINYLRPDLKRGNFTE 242
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
ED +II H++ GNKW+ I+ LPGRTDN IKN+WN+ ++R+
Sbjct: 243 QEDDLIINLHSLLGNKWSLIAARLPGRTDNEIKNYWNTHIKRK 371
Score = 33.9 bits (76), Expect = 0.064
Identities = 23/82 (28%), Positives = 39/82 (47%), Gaps = 10/82 (12%)
Frame = +3
Query: 17 DRVKGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQHRPFT 76
D +G+++ QED+ ++ L G + WS+I+A +PGR+ + W + + R
Sbjct: 216 DLKRGNFTEQEDDLIINLHSLLGNK-WSLIAARLPGRTDNEIKNYWNTHIKRKLYSRGVD 392
Query: 77 P----------TEDTIIIQAHA 88
P T TII A+A
Sbjct: 393 PQTHRSLNDSTTSTTIIPPANA 458
>BG644406 homologue to PIR|T07398|T07 myb-related transcription factor THM6 -
tomato, partial (42%)
Length = 762
Score = 105 bits (263), Expect = 1e-23
Identities = 49/102 (48%), Positives = 65/102 (63%), Gaps = 1/102 (0%)
Frame = +1
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIP-GRSGKSCRLRWCNQLSPDVQHRPFTPT 78
KG W+P+ED L+ V +HGP NW + R KSCRLRW N L P ++ FT
Sbjct: 184 KGPWTPEEDIMLVSFVQEHGPGNWRTVPTHTGLRRCSKSCRLRWTNYLRPGIKRGSFTDQ 363
Query: 79 EDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ +IIQ A+ GNKWA I+ LP RTDN IKN+WN+ L+++
Sbjct: 364 EEKMIIQLQALLGNKWAAIASYLPERTDNDIKNYWNTHLKKK 489
>TC80618 homologue to PIR|T06455|T06455 Myb26 protein - garden pea, partial
(57%)
Length = 799
Score = 105 bits (261), Expect = 2e-23
Identities = 51/116 (43%), Positives = 71/116 (60%), Gaps = 2/116 (1%)
Frame = +2
Query: 6 DSSNGGGGGDRDRVKGSWSPQEDETLMKLVGQHGPRNWSVI--SAGIPGRSGKSCRLRWC 63
D D D KG W+ +ED L+ + HG W+ + SAG+ R+GKSCRLRW
Sbjct: 53 DKKECSSSQDPDVRKGPWTMEEDLILINYIANHGEGVWNSLAKSAGLK-RTGKSCRLRWL 229
Query: 64 NQLSPDVQHRPFTPTEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRR 119
N L PDV+ TP E +II+ HA GN+W+ I++ LPGRTDN IKN+W + +++
Sbjct: 230 NYLRPDVRRGNITPEEQLLIIELHAKWGNRWSKIAKHLPGRTDNEIKNYWRTRIQK 397
Score = 28.5 bits (62), Expect = 2.7
Identities = 13/52 (25%), Positives = 29/52 (55%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVISAGIPGRSGKSCRLRWCNQLSPDVQ 71
+G+ +P+E +++L + G R WS I+ +PGR+ + W ++ ++
Sbjct: 254 RGNITPEEQLLIIELHAKWGNR-WSKIAKHLPGRTDNEIKNYWRTRIQKHIK 406
>TC78267 weakly similar to PIR|B85064|B85064 MYB-like protein [imported] -
Arabidopsis thaliana, partial (35%)
Length = 869
Score = 105 bits (261), Expect = 2e-23
Identities = 49/103 (47%), Positives = 71/103 (68%), Gaps = 2/103 (1%)
Frame = +2
Query: 20 KGSWSPQEDETLMKLVGQHGPRNWSVIS--AGIPGRSGKSCRLRWCNQLSPDVQHRPFTP 77
KG+W+ +ED L+ V ++G NW + AG+ R GKSCRLRW N L P+++ FT
Sbjct: 134 KGTWTAEEDRKLIAYVTRYGFWNWRQLPKFAGL-SRCGKSCRLRWLNYLRPNIRRGNFTK 310
Query: 78 TEDTIIIQAHAVHGNKWATISRLLPGRTDNAIKNHWNSTLRRR 120
E+ +II+ H GNKW+TI+ LPGRTDN +KNHW+++L++R
Sbjct: 311 DEEELIIRMHKKLGNKWSTIAAELPGRTDNEVKNHWHTSLKKR 439
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.133 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,971,929
Number of Sequences: 36976
Number of extensions: 141699
Number of successful extensions: 4946
Number of sequences better than 10.0: 222
Number of HSP's better than 10.0 without gapping: 2307
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4318
length of query: 260
length of database: 9,014,727
effective HSP length: 94
effective length of query: 166
effective length of database: 5,538,983
effective search space: 919471178
effective search space used: 919471178
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0002.6


