

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0001.6
(381 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
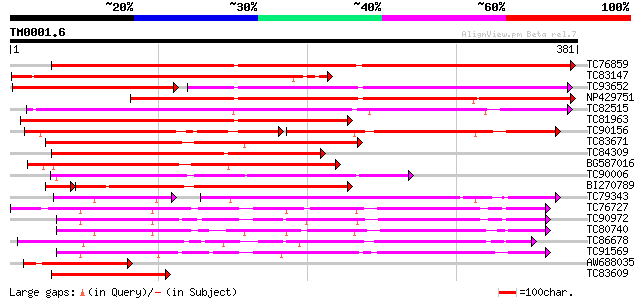
Score E
Sequences producing significant alignments: (bits) Value
TC76859 Enod8.1 [Medicago truncatula]; early nodule-specific pro... 335 2e-92
TC83147 similar to PIR|T48618|T48618 early nodule-specific prote... 298 2e-81
TC93652 Enod8.2 [Medicago truncatula] 192 9e-80
NP429751 NP429751|AF463407.1|AAL68830.1 Enod8.3 [Medicago trunca... 274 4e-74
TC82515 similar to GP|3688284|emb|CAA09694.1 lanatoside 15'-O-ac... 271 4e-73
TC81963 Enod8-like protein [Medicago truncatula] 244 5e-65
TC90156 similar to PIR|B86227|B86227 hypothetical protein [impor... 152 4e-63
TC83671 similar to PIR|T09416|T09416 coil protein PO22 microspo... 218 3e-57
TC84309 similar to GP|9294302|dbj|BAB02204.1 nodulin-like protei... 204 4e-53
BG587016 similar to PIR|T09416|T09 coil protein PO22 microspore... 203 8e-53
TC90006 similar to PIR|A96590|A96590 hypothetical protein T22H22... 193 1e-49
BI270789 similar to PIR|T09416|T09 coil protein PO22 microspore... 170 2e-47
TC79343 weakly similar to PIR|E86411|E86411 protein F1K23.18 [im... 104 3e-37
TC76727 similar to GP|10638955|emb|CAB81548. putative proline-ri... 133 1e-31
TC90972 similar to GP|10638955|emb|CAB81548. putative proline-ri... 129 2e-30
TC80740 similar to GP|10177228|dbj|BAB10602. GDSL-motif lipase/h... 120 9e-28
TC86678 similar to PIR|E96579|E96579 hypothetical protein T18A20... 114 6e-26
TC91569 similar to GP|18252233|gb|AAL61949.1 putative APG protei... 105 2e-23
AW688035 similar to PIR|T48618|T486 early nodule-specific protei... 104 6e-23
TC83609 similar to GP|5295941|dbj|BAA81842.1 ESTs AU075322(C1110... 103 1e-22
>TC76859 Enod8.1 [Medicago truncatula]; early nodule-specific protein
Length = 1350
Score = 335 bits (858), Expect = 2e-92
Identities = 169/354 (47%), Positives = 224/354 (62%), Gaps = 2/354 (0%)
Frame = +2
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C FPAI++FG SN DTGG++AAF P PYGE++ + + R DGR+I+DFIA LPY
Sbjct: 140 CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPY 319
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
LS YLNSLG+N+ HGANFA+GGSTI + +SPFSL +Q++QFK+F ++TK +
Sbjct: 320 LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIR 499
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FRMMNFDQMRESMPDIVNQLASAV 207
+ + + +P + FSKALY FDIGQNDL++G F Q+ ++PDIVN +
Sbjct: 500 DQG--GVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENI 673
Query: 208 KNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDR 267
KNIY LG R+FWIH T P GC PV L N P+ D YGC K N ++ FN +LK+
Sbjct: 674 KNIYNLGARSFWIHGTGPKGCAPVIL---ANFPSAIKDSYGCAKQYNEVSQYFNFKLKEA 844
Query: 268 VVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVN-DTHIWCGTIGTA 326
+ +LR+ L AAITYVD+Y KY L +N + GF P CCGY + + CG
Sbjct: 845 LAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNIGVGCGASINI 1024
Query: 327 NGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQACYR 380
NG + +C+ PS + WDGVHY EAAN V ++IL G F DPP + +ACYR
Sbjct: 1025NGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYR 1186
>TC83147 similar to PIR|T48618|T48618 early nodule-specific protein-like -
Arabidopsis thaliana, partial (43%)
Length = 653
Score = 298 bits (764), Expect = 2e-81
Identities = 159/222 (71%), Positives = 177/222 (79%), Gaps = 6/222 (2%)
Frame = +1
Query: 2 GLRPLFIAFFLSCTLCVNSVELKNSPP-CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGE 60
GL +F+ FF + CV VELK+S P C+FPAIYNFGDSNSDTGGISAAF PIP PYG+
Sbjct: 1 GLSEVFVVFFFFLS-CVKCVELKDSSPSCSFPAIYNFGDSNSDTGGISAAFVPIPSPYGQ 177
Query: 61 SFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETI 120
F KP RD DGR+I+D+IA+KL PYLSAYLNSLGTNYRHGANFATGGSTIR+QNETI
Sbjct: 178 GFFHKPFGRDSDGRVILDYIADKLKWPYLSAYLNSLGTNYRHGANFATGGSTIRKQNETI 357
Query: 121 FQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDL 180
FQYGISPFSLD+Q VQF QFKARTKQLYQEA +LERSKLPVPEEF+KALYTFDIGQNDL
Sbjct: 358 FQYGISPFSLDIQIVQFNQFKARTKQLYQEANNSLERSKLPVPEEFAKALYTFDIGQNDL 537
Query: 181 SVGFRMMNF-----DQMRESMPDIVNQLASAVKNIYELGGRT 217
SVGFR F + S P V+ +S KNIYE GGR+
Sbjct: 538 SVGFRKDEF*PNS*NHAGYS*P--VSLCSS--KNIYEQGGRS 651
>TC93652 Enod8.2 [Medicago truncatula]
Length = 1195
Score = 192 bits (488), Expect(2) = 9e-80
Identities = 108/263 (41%), Positives = 148/263 (56%), Gaps = 4/263 (1%)
Frame = +2
Query: 120 IFQYGI-SPFSLDMQFV-QFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQ 177
+++ GI SPFSL +Q K F ++T + + + + +P + FSKALYTFDIGQ
Sbjct: 401 LYRNGIFSPFSLQIQGTFNSKIFISKTNLIRDQG--GVFATLIPKEDYFSKALYTFDIGQ 574
Query: 178 NDLSVG-FRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYK 236
NDL G F Q+ ++PDIVN +KNIY LG R+FWIH+T P GC P L
Sbjct: 575 NDLIGGYFGNKTIKQVNATVPDIVNNFIVNIKNIYNLGARSFWIHSTVPSGCTPTILA-- 748
Query: 237 HNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNT 296
N P+ D YGC K N ++ FN +LK + +LR +LP AAITYVD+Y+ Y L N
Sbjct: 749 -NFPSAIKDSYGCAKQYNEVSQYFNLKLKKALAQLRVDLPLAAITYVDIYSPNYSLFQNP 925
Query: 297 KNEGFVDPLKICCGYHVN-DTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAAN 355
K GF P CCGY + + CG NG + +C+ PS + WDG H+ E
Sbjct: 926 KKYGFELPHVACCGYGGKYNIRVGCGETLNINGTKIEAGSCKNPSTRIIWDGSHFTERRY 1105
Query: 356 HWVANRILNGSFTDPPTLITQAC 378
V ++I G+F+DPP + +AC
Sbjct: 1106KIVFDQISTGAFSDPPISLNRAC 1174
Score = 122 bits (307), Expect(2) = 9e-80
Identities = 58/111 (52%), Positives = 79/111 (70%)
Frame = +3
Query: 3 LRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESF 62
LR + + + LC+ + + + C FPAI++FG SN DTGG++AAF+ P PYGE++
Sbjct: 51 LRHMSLVSLIVLILCIITPPIFATRNCDFPAIFSFGASNVDTGGLAAAFQAPPSPYGETY 230
Query: 63 PQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTI 113
+ + R DGR+I+DFIA+ LPYLS YLNSLG+N+ HGANFATGGSTI
Sbjct: 231 FHRSTGRFSDGRIILDFIAQSFGLPYLSPYLNSLGSNFTHGANFATGGSTI 383
>NP429751 NP429751|AF463407.1|AAL68830.1 Enod8.3 [Medicago truncatula]
Length = 902
Score = 274 bits (701), Expect = 4e-74
Identities = 144/304 (47%), Positives = 187/304 (61%), Gaps = 5/304 (1%)
Frame = +3
Query: 82 EKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFK 141
+ LPYLS YLNSLG+N+ HGANFAT GSTI+ N I SPFSL +Q +QFK F
Sbjct: 3 QSFGLPYLSPYLNSLGSNFTHGANFATAGSTIKIPNSIIPNGMFSPFSLQIQSIQFKDFI 182
Query: 142 ARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGF-RMMNFDQMRESMPDIV 200
+ K + + + + +P + +SKALYTFDIGQNDL+ GF Q+ ++PDIV
Sbjct: 183 PKAKFIRDQG--GVFATLIPKEDYYSKALYTFDIGQNDLTAGFFGNKTIQQVNTTVPDIV 356
Query: 201 NQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEF 260
+KNIY LG R+FWIHNT PIGC+P+ L N P+ D YGC K N ++ F
Sbjct: 357 KSFIDNIKNIYNLGARSFWIHNTGPIGCVPLILA---NFPSAIKDRYGCAKQYNEVSQYF 527
Query: 261 NKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCG----YHVNDT 316
N +LK+ + +LR +LP AAITYVD+Y+ KY L N K GF PL CCG Y+ N
Sbjct: 528 NLKLKEALAQLRKDLPLAAITYVDIYSPKYSLFQNPKKYGFELPLVACCGNGGKYNYN-I 704
Query: 317 HIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRILNGSFTDPPTLITQ 376
CG NG + +C+KPS + WDG HY EAAN V ++I NG+FTDPP + +
Sbjct: 705 RAGCGATININGTNTVVGSCKKPSTRIIWDGTHYTEAANKIVFDQISNGAFTDPPIPLNR 884
Query: 377 ACYR 380
ACY+
Sbjct: 885 ACYK 896
>TC82515 similar to GP|3688284|emb|CAA09694.1 lanatoside
15'-O-acetylesterase {Digitalis lanata}, partial (76%)
Length = 1230
Score = 271 bits (692), Expect = 4e-73
Identities = 150/379 (39%), Positives = 218/379 (56%), Gaps = 12/379 (3%)
Frame = +1
Query: 12 LSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDC 71
L+C + ++S + C F I+NFGDSNSDTGG +AF P PYG ++ + P R
Sbjct: 85 LNCIM-ISSFIRSSYSKCDFQGIFNFGDSNSDTGGFYSAFPAQPIPYGMTYFKTPVGRSS 261
Query: 72 DGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLD 131
DGRLIVDF+AE L LPYLS YL S+G++Y HGANFAT ST+ ++F G+SPF+L
Sbjct: 262 DGRLIVDFLAEALGLPYLSPYLQSIGSDYTHGANFATSASTVLLPTTSLFVSGLSPFALQ 441
Query: 132 MQFVQFKQFKARTKQLYQ----EAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMM 187
+Q Q +QF+A+ ++ + T + K+P P+ F K++Y F IGQND +
Sbjct: 442 IQLRQMQQFRAKVHDFHKRDPLKPSTCASKIKIPSPDIFGKSIYMFYIGQNDFTSKIAAS 621
Query: 188 -NFDQMRESMPDIVNQLASAVKNI-YELGGRTFWIHNTAPIGCLPVNLFYKHNLP--AGY 243
+ ++ +P I+ Q+ASA+K + Y GGRTF + N P+GC P Y LP +
Sbjct: 622 GGINGLKNYLPQIIYQIASAIKELYYAQGGRTFMVLNLGPVGCYP---GYLVELPHTSSD 792
Query: 244 LDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVD 303
L+ +GC+ N ++NK LK+ + + R L +A++ YVD +A L + + G
Sbjct: 793 LNEHGCIITYNNAVDDYNKLLKETLTQTRKSLSDASLIYVDTNSALMELFRHPTSYGLKH 972
Query: 304 PLKICCGYHVNDTHI----WCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVA 359
K CCG+ D + CG ++ +ACE P YVSWDG+H+ EAAN +A
Sbjct: 973 STKACCGHGGGDYNFDPKALCG--------NMLASACEDPQNYVSWDGIHFTEAANKIIA 1128
Query: 360 NRILNGSFTDPPTLITQAC 378
ILNGS +DPP L+ + C
Sbjct: 1129MAILNGSLSDPPFLLHKLC 1185
>TC81963 Enod8-like protein [Medicago truncatula]
Length = 783
Score = 244 bits (622), Expect = 5e-65
Identities = 125/225 (55%), Positives = 157/225 (69%), Gaps = 2/225 (0%)
Frame = +3
Query: 8 IAFFLSCTLCVNSV-ELKNSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKP 66
+ FF+ ++ +V E ++ C F AI+NFGDSNSDTGG++A+F PPYGE++ +P
Sbjct: 111 VTFFVVLSIATTTVIESSSNSECNFRAIFNFGDSNSDTGGLAASFVAPKPPYGETYFHRP 290
Query: 67 SARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGIS 126
+ R DGRLIVDFIA+ LPYLSAYL+SLGTN+ HGANFAT STIR I Q G S
Sbjct: 291 NGRFSDGRLIVDFIAQSFGLPYLSAYLDSLGTNFSHGANFATTSSTIRPPPSIIPQGGFS 470
Query: 127 PFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVG-FR 185
PF LD+Q+ QF+ FK RT+ + Q+ L S +P E FSKALYTFDIGQNDL G F
Sbjct: 471 PFYLDVQYTQFRDFKPRTQFIRQQG--GLFASLMPKEEYFSKALYTFDIGQNDLGAGFFG 644
Query: 186 MMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLP 230
M Q+ S+P+I+N + VK+IY LGGR+FWIHNT PIGCLP
Sbjct: 645 NMTIQQVNASVPEIINSFSKNVKDIYNLGGRSFWIHNTGPIGCLP 779
>TC90156 similar to PIR|B86227|B86227 hypothetical protein [imported] -
Arabidopsis thaliana, partial (84%)
Length = 1421
Score = 152 bits (385), Expect(2) = 4e-63
Identities = 76/190 (40%), Positives = 114/190 (60%), Gaps = 6/190 (3%)
Frame = +1
Query: 187 MNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLP--VNLFYKHNLPAGYL 244
+++ Q+ + +P ++ ++ +AVK++Y GGR FW+HNT P GCLP + L K +L
Sbjct: 574 LSYVQVIKRIPTVITEIENAVKSLYNEGGRKFWVHNTGPFGCLPKLIALSXKKDL----- 738
Query: 245 DPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDP 304
D +GC+ N A FN+ L KLRTEL +A + YVD+YA K LI+N GF +P
Sbjct: 739 DSFGCLSSYNSAARLFNEALYHSSQKLRTELKDATLVYVDIYAIKNDLITNATKYGFTNP 918
Query: 305 LKICCGY----HVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVAN 360
L +CCG+ + D + CG G C++ S YVSWDG+HY EAAN W+A+
Sbjct: 919 LMVCCGFGGPPYNFDARVTCGQPGY--------QVCDEGSRYVSWDGIHYTEAANTWIAS 1074
Query: 361 RILNGSFTDP 370
+IL+ +++ P
Sbjct: 1075KILSTAYSTP 1104
Score = 107 bits (266), Expect(2) = 4e-63
Identities = 74/183 (40%), Positives = 111/183 (60%), Gaps = 9/183 (4%)
Frame = +3
Query: 11 FLSCTLCVN----SVELKNSPPCAF-PAI-YNFGDSNSDTGG-ISAAFEPIPPPYGESFP 63
++S T+C+ S+ L S C+ PA+ + FGDSNSDTGG +S P+ P G +F
Sbjct: 57 WISMTICIFFSSVSLALSVSSGCSSKPAVVFVFGDSNSDTGGLVSGLGFPVNLPNGRTFF 236
Query: 64 QKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSL-GTNYRHGANFATGGSTIRRQNETIFQ 122
+ + R DGRL++DF+ + LN +L+ YL+S+ G+ + +GANFA GS+ T+ +
Sbjct: 237 HRSTGRLSDGRLVIDFLCQSLNTRFLTPYLDSMSGSTFTNGANFAVVGSS------TLPK 398
Query: 123 YGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEE-FSKALYTFDIGQNDLS 181
Y PFSL++Q +QF+ FKAR+ QL A +K + ++ F ALY DIGQNDL+
Sbjct: 399 Y--LPFSLNIQVMQFQHFKARSLQL------ATSGAKNMINDQGFRDALYLIDIGQNDLA 554
Query: 182 VGF 184
F
Sbjct: 555 DSF 563
>TC83671 similar to PIR|T09416|T09416 coil protein PO22
microspore/pollen-specific - alfalfa, partial (60%)
Length = 671
Score = 218 bits (555), Expect = 3e-57
Identities = 113/218 (51%), Positives = 146/218 (66%), Gaps = 5/218 (2%)
Frame = +3
Query: 25 NSPPCAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKL 84
+S C +PAIYNFGDSNSDTG I A + + PP G +F S R DGRLI+D I E+L
Sbjct: 39 SSDKCVYPAIYNFGDSNSDTGTIYATYTSVQPPNGITFFGNISGRASDGRLIIDSITEEL 218
Query: 85 NLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKART 144
LPYLSAYLNS+G+NYRHGANFA G++IR + G F+L +Q QF FK+ T
Sbjct: 219 KLPYLSAYLNSVGSNYRHGANFAVSGASIRPR-------GYHLFNLGLQVSQFILFKSHT 377
Query: 145 KQLYQEAKTALE----RSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIV 200
K L+ + +S LP PE+FSKALYT DIGQNDL+ GF+ + +Q++ S+P+I+
Sbjct: 378 KILFNQLSNNRTEPSLKSGLPRPEDFSKALYTIDIGQNDLAHGFQYTSEEQVQRSIPEIL 557
Query: 201 NQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLF-YKH 237
+ + +VK +Y G R FWIHNT PIGCLP N + YKH
Sbjct: 558 SNFSQSVKQLYNEGARVFWIHNTGPIGCLPFNYYTYKH 671
>TC84309 similar to GP|9294302|dbj|BAB02204.1 nodulin-like protein protein
{Arabidopsis thaliana}, partial (47%)
Length = 672
Score = 204 bits (520), Expect = 4e-53
Identities = 100/185 (54%), Positives = 137/185 (74%), Gaps = 1/185 (0%)
Frame = +2
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C+FPAI+NFGDSNSDTGG+SAAF PPP G +F Q P+ R DGRLI+DF+A+ L+LPY
Sbjct: 119 CSFPAIFNFGDSNSDTGGLSAAFGQAPPPNGITFFQTPAGRFSDGRLIIDFLAQNLSLPY 298
Query: 89 LSAYLNSLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLY 148
LSAYL+S+G+N+ +GANFAT GSTIR QN T Q G SP SLD+Q +Q+ FKAR+ +
Sbjct: 299 LSAYLDSVGSNFSNGANFATAGSTIRPQNTTKSQSGYSPISLDVQLIQYSDFKARS--IL 472
Query: 149 QEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRM-MNFDQMRESMPDIVNQLASAV 207
K + LP E FS+ALYTFDIGQNDL+ G+++ M +Q++ +PD++ Q + +
Sbjct: 473 VRKKGGVFMKLLPKEEYFSEALYTFDIGQNDLTAGYKLNMTTEQVKAYIPDVLGQFSDVI 652
Query: 208 KNIYE 212
+++Y+
Sbjct: 653 RSVYK 667
>BG587016 similar to PIR|T09416|T09 coil protein PO22
microspore/pollen-specific - alfalfa, partial (55%)
Length = 696
Score = 203 bits (517), Expect = 8e-53
Identities = 112/225 (49%), Positives = 144/225 (63%), Gaps = 15/225 (6%)
Frame = +3
Query: 13 SCTLCVNSV---ELKNSPP--------CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGES 61
SCTL SV + +SP C +PAIYNFGDSNSDTG +A + + PP G S
Sbjct: 39 SCTLVFQSVCDIYIHSSPK*ECL*LKKCEYPAIYNFGDSNSDTGAANAIYTAVTPPNGIS 218
Query: 62 FPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETIF 121
+ + R DGRLI+DFI+E+L LPYLSAYLNS+G+NYRHGANFA GG++IR
Sbjct: 219 YFGSTTGRASDGRLIIDFISEELKLPYLSAYLNSIGSNYRHGANFAVGGASIR------- 377
Query: 122 QYGISPFSLDMQFVQFKQFKARTK----QLYQEAKTALERSKLPVPEEFSKALYTFDIGQ 177
G SP L +Q QF FK+ TK QL + +S LP EEFSKALYT DIGQ
Sbjct: 378 PGGYSPIFLGLQVSQFILFKSHTKILFNQLSDNRTESPFKSGLPRNEEFSKALYTIDIGQ 557
Query: 178 NDLSVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHN 222
NDL++G + + +Q++ S+PDI++Q + AV+ +Y G R FWIHN
Sbjct: 558 NDLAIGLQNTSEEQVKRSIPDILSQFSQAVQQLYNEGARVFWIHN 692
>TC90006 similar to PIR|A96590|A96590 hypothetical protein T22H22.20
[imported] - Arabidopsis thaliana, partial (63%)
Length = 858
Score = 193 bits (490), Expect = 1e-49
Identities = 112/252 (44%), Positives = 145/252 (57%), Gaps = 8/252 (3%)
Frame = +3
Query: 28 PCA------FPAIYNFGDSNSDTGG-ISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFI 80
PCA FPA++NFGDSNSDTG ++A FE + PP G ++ PS R DGRLI+DF+
Sbjct: 90 PCAKSIHLDFPAVFNFGDSNSDTGTLVTAGFESLYPPNGHTYFHLPSGRYSDGRLIIDFL 269
Query: 81 AEKLNLPYLSAYLNSLGT-NYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQ 139
+ L+LP+L+AYL+SLG N+R G NFA GSTI + I PFS +Q QF +
Sbjct: 270 MDALDLPFLNAYLDSLGLPNFRKGCNFAAAGSTILPATAS----SICPFSFGIQVSQFLK 437
Query: 140 FKARTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDI 199
FKAR +L ++ +P + F K LY FDIGQNDL+ F DQ+ S+P I
Sbjct: 438 FKARALELLSGKGRKFDKY-VPSEDIFEKGLYMFDIGQNDLAGAFYSKTLDQVLASIPTI 614
Query: 200 VNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVE 259
+ + S +K +Y+ G R FWIHNT P+GCL N+ K LD GCV N
Sbjct: 615 LLEFESGIKRLYDEGARYFWIHNTGPLGCLAQNV-AKFGTDPSKLDELGCVSGHNQAVKT 791
Query: 260 FNKQLKDRVVKL 271
FN QL KL
Sbjct: 792 FNLQLHALCSKL 827
>BI270789 similar to PIR|T09416|T09 coil protein PO22
microspore/pollen-specific - alfalfa, partial (62%)
Length = 657
Score = 170 bits (430), Expect(2) = 2e-47
Identities = 90/186 (48%), Positives = 117/186 (62%)
Frame = +1
Query: 45 GGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGA 104
G A F PP G SF S R DGRLI+D+I E+L +PYLSAYLNS+G+NYR+GA
Sbjct: 76 GTAYATFLCNQPPNGISFGNI-SGRASDGRLIIDYITEELKVPYLSAYLNSVGSNYRYGA 252
Query: 105 NFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPE 164
NFA GG++IR + G SPF L +Q QF QFK+ T+ L+ +S LP PE
Sbjct: 253 NFAAGGASIRPGS------GFSPFHLGLQVDQFIQFKSHTRILFNNGTEPSLKSGLPRPE 414
Query: 165 EFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTA 224
+F ALYT DIG NDL+ GF + +Q++ S P+I+ + AVK +Y + R FWIHN
Sbjct: 415 DFCTALYTIDIGLNDLASGFLHASEEQVQMSFPEILGHFSKAVKQLYNVXARVFWIHNVG 594
Query: 225 PIGCLP 230
P+GC P
Sbjct: 595 PVGCCP 612
Score = 37.4 bits (85), Expect(2) = 2e-47
Identities = 15/20 (75%), Positives = 17/20 (85%)
Frame = +3
Query: 25 NSPPCAFPAIYNFGDSNSDT 44
+S C +PAIYNFGDSNSDT
Sbjct: 15 SSHECVYPAIYNFGDSNSDT 74
>TC79343 weakly similar to PIR|E86411|E86411 protein F1K23.18 [imported] -
Arabidopsis thaliana, partial (52%)
Length = 1420
Score = 104 bits (259), Expect(2) = 3e-37
Identities = 80/248 (32%), Positives = 116/248 (46%), Gaps = 6/248 (2%)
Frame = +3
Query: 129 SLDMQFVQFKQFKARTKQL--YQEAKTALERSKLPVPEEFSKALYTF-DIGQNDLSVG-F 184
SL V F + T QL ++E +L S E F+ +L+ +IG ND + F
Sbjct: 483 SLKRGVVGFSTNYSLTVQLNWFKELLPSLCNSSKNCHEVFANSLFLMGEIGGNDFNYPLF 662
Query: 185 RMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYL 244
+ +++ +P +++ + SA+ + +LG RT I P+GC + L
Sbjct: 663 IRRSIVEIKTYVPHVISAITSAINELIDLGARTLMIPGNFPLGCNVIYLTKYETTDKSQY 842
Query: 245 DPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDP 304
D GC+K N A +N++L+ + +LR P A I Y D Y A L N GF
Sbjct: 843 DSAGCLKWLNEFAEFYNQELQYELHRLRRIHPHATIIYADYYNALLPLYQNPTKFGFTG- 1019
Query: 305 LKICCGY--HVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRI 362
LK CCG N CG G VF AC+ PS Y+ WDGVH EAA +A+ I
Sbjct: 1020LKNCCGMGGSYNFGSGSCGKPG------VF--ACDDPSQYIGWDGVHLTEAAYRLIADGI 1175
Query: 363 LNGSFTDP 370
+NG + P
Sbjct: 1176INGPCSVP 1199
Score = 68.9 bits (167), Expect(2) = 3e-37
Identities = 37/93 (39%), Positives = 55/93 (58%), Gaps = 10/93 (10%)
Frame = +2
Query: 30 AFPAIYNFGDSNSDTGGISAAFEPIP-----PPYGESFPQKPSARDCDGRLIVDFIAEKL 84
++ +I++FGDS +DTG + + +P PPYG+++ PS R DGRLI+DFIAE L
Sbjct: 188 SYSSIFSFGDSIADTGNLYLSSQPPSDHCFFPPYGQTYFHHPSGRCSDGRLIIDFIAESL 367
Query: 85 NLPYLSAYLNSLG-----TNYRHGANFATGGST 112
+P + YL + + GANFA G+T
Sbjct: 368 GIPMVKPYLGIKNGVLEDNSAKEGANFAVIGAT 466
>TC76727 similar to GP|10638955|emb|CAB81548. putative proline-rich protein
APG isolog {Cicer arietinum}, partial (92%)
Length = 1919
Score = 133 bits (335), Expect = 1e-31
Identities = 110/379 (29%), Positives = 158/379 (41%), Gaps = 16/379 (4%)
Frame = -2
Query: 1 MGLRPLFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGG---ISAAFEPIPPP 57
M + + +SC L S + PAI FGDS D G + F+ PP
Sbjct: 1171 MNSKETLVLLIVSCFLTCGSF----AQDTLVPAIMTFGDSAVDVGNNDYLPTLFKANYPP 1004
Query: 58 YGESFPQK-PSARDCDGRLIVDFIAEKLNLP-YLSAYLN--SLGTNYRHGANFATGGSTI 113
YG F K P+ R C+G+L DF AE L + AYL+ + G N GANFA+ S
Sbjct: 1003 YGRDFTNKQPTGRFCNGKLATDFTAETLGFTSFAPAYLSPQASGKNLLLGANFASAASGY 824
Query: 114 RRQNETIFQYGISPFSLDMQFVQFKQFKARTKQLYQEAKTALERSKLPVPEEFSKALYTF 173
+ T+ L Q FK+++ + Q+ K A +LY
Sbjct: 823 DEKAATLNH----AIPLSQQLEYFKEYQGKLAQVAGSKKAA---------SIIKDSLYVL 683
Query: 174 DIGQNDLSVGF-------RMMNFDQMRESMPDIVNQLASAVKNIYELGGRTFWIHNTAPI 226
G +D + + + DQ + D + +K +Y LG R + + P+
Sbjct: 682 SAGSSDFVQNYYTNPWINQAITVDQYSSYLLD---SFTNFIKGVYGLGARKIGVTSLPPL 512
Query: 227 GCLPV--NLFYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVD 284
GCLP LF H GCV N A FNK++ L+ +LP I D
Sbjct: 511 GCLPAARTLFGYHE--------NGCVARINTDAQGFNKKVSSAASNLQKQLPGLKIVIFD 356
Query: 285 LYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVS 344
+Y Y L+ N N GF + K CCG + +T T N K + C + YV
Sbjct: 355 IYKPLYDLVQNPSNFGFAEAGKGCCGTGLVET-----TSLLCNPKSL--GTCSNATQYVF 197
Query: 345 WDGVHYAEAANHWVANRIL 363
WD VH +EAAN +A+ ++
Sbjct: 196 WDSVHPSEAANQVLADNLI 140
>TC90972 similar to GP|10638955|emb|CAB81548. putative proline-rich protein
APG isolog {Cicer arietinum}, partial (88%)
Length = 1258
Score = 129 bits (324), Expect = 2e-30
Identities = 107/346 (30%), Positives = 156/346 (44%), Gaps = 14/346 (4%)
Frame = +3
Query: 32 PAIYNFGDSNSDTGG---ISAAFEPIPPPYGESFP-QKPSARDCDGRLIVDFIAEKLNLP 87
PAI FGDS D G + F+ PPYG F KP+ R C+G+L D AE L
Sbjct: 105 PAIVTFGDSAVDVGNNDYLFTLFKANYPPYGRDFVXHKPTGRFCNGKLATDITAETLGFK 284
Query: 88 -YLSAYLN--SLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKART 144
Y AYL+ + G N GANFA+ S + I + I L Q +K+++++
Sbjct: 285 SYAPAYLSPQATGKNLLIGANFASAASGYD-EKAAILNHAIP---LSQQLKYYKEYQSKL 452
Query: 145 KQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLSVGFRMMNFDQMRESMPD-----I 199
++ K A ALY G +D + +N + PD +
Sbjct: 453 SKIAGSKKAA---------SIIKGALYLLSGGSSDFIQNY-YVNPLINKVVTPDQYSAYL 602
Query: 200 VNQLASAVKNIYELGGRTFWIHNTAPIGCLPVN--LFYKHNLPAGYLDPYGCVKDQNVMA 257
V+ +S VK++Y+LG R + + P+GCLP LF H GCV N A
Sbjct: 603 VDTYSSFVKDLYKLGARKIGVTSLPPLGCLPATRTLFGFHEK--------GCVTRINNDA 758
Query: 258 VEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTH 317
FNK++ VKL+ +LP I ++Y Y L+ + GF + K CCG + +T
Sbjct: 759 QGFNKKINSATVKLQKQLPGLKIVVFNIYKPLYELVQSPSKFGFAEARKGCCGTGIVET- 935
Query: 318 IWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRIL 363
T N K + C + YV WD VH +EAAN +A+ ++
Sbjct: 936 ----TSLLCNQKSL--GTCSNATQYVFWDSVHPSEAANQILADALI 1055
>TC80740 similar to GP|10177228|dbj|BAB10602. GDSL-motif
lipase/hydrolase-like protein {Arabidopsis thaliana},
partial (94%)
Length = 1167
Score = 120 bits (301), Expect = 9e-28
Identities = 101/350 (28%), Positives = 150/350 (42%), Gaps = 18/350 (5%)
Frame = +3
Query: 32 PAIYNFGDSNSDTGGISAAFEPIP---PPYGESFP-QKPSARDCDGRLIVDFIAEKLNLP 87
PA++ FGDS D + + + PPYG F Q P+ R C+G+L DF AE L
Sbjct: 186 PALFIFGDSVVDARNNNNLYTIVKSNFPPYGRDFNNQMPTGRFCNGKLAADFTAENLGFT 365
Query: 88 -YLSAYLN--SLGTNYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKART 144
Y AYLN N +GANFA+G S ++ SL+ Q
Sbjct: 366 TYPPAYLNLQEKRKNLLNGANFASGASGYFDPTAKLYH----AISLEQQL---------- 503
Query: 145 KQLYQEAKTALE--RSKLPVPEEFSKALYTFDIGQNDLSVGF-------RMMNFDQMRES 195
+ Y+E + L K S A+Y G +D + ++ DQ +
Sbjct: 504 -EHYKECQNILVGVAGKSNASSIISGAIYLVRAGSSDFVQNYYINPLLYKVFTADQFSDI 680
Query: 196 MPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLP--VNLFYKHNLPAGYLDPYGCVKDQ 253
+ + ++N+Y LG R + P+GCLP + LF H+ CV
Sbjct: 681 L---MQHYTIFIQNLYALGARKIGVTTLPPLGCLPAAITLFGSHSNE--------CVDRL 827
Query: 254 NVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHV 313
N A+ FN +L L+ EL + +D+Y + L++ GF + K CCG +
Sbjct: 828 NNDALNFNTKLNTTSQNLQKELSNLTLAVLDIYQPLHDLVTKPTENGFYEARKACCGTGL 1007
Query: 314 NDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRIL 363
+T I C KD G C + YV WDG H +EAAN+ +A+ +L
Sbjct: 1008IETSILC-------NKDSIG-TCANATEYVFWDGFHTSEAANNVLADDLL 1133
>TC86678 similar to PIR|E96579|E96579 hypothetical protein T18A20.15
[imported] - Arabidopsis thaliana, partial (43%)
Length = 1341
Score = 114 bits (285), Expect = 6e-26
Identities = 100/361 (27%), Positives = 159/361 (43%), Gaps = 12/361 (3%)
Frame = +2
Query: 6 LFIAFFLSCTLCVNSVELKNSPPCAFPAIYNFGDSNSDTGGIS----AAFEPIP-PPYGE 60
+F FL + + K + A++ FGDS D G + F+ PYGE
Sbjct: 101 IFFKVFLIIAIISQTFGSKTDYYRSNKALFIFGDSFLDAGNNNYINTTTFDQANFLPYGE 280
Query: 61 SFPQKPSARDCDGRLIVDFIAEKLNLPYLSAYLNSLGTNYRHGANFATGGSTIRRQNETI 120
++ P+ R DGRLI DFIAE +N+P + +L Y +G NFA+GG+ +
Sbjct: 281 TYFNFPTGRFSDGRLISDFIAEYVNIPLVPPFLQPDNNKYYNGVNFASGGAGALVET--- 451
Query: 121 FQYGISPFSLDMQFVQFKQFKA--RTKQLYQEAKTALERSKLPVPEEFSKALYTFDIGQN 178
FQ + PF Q + FK+ R K ++KT L S A+Y F IG N
Sbjct: 452 FQGSVIPFK--TQAINFKKVTTWLRHKLGSSDSKTLL-----------SNAVYMFSIGSN 592
Query: 179 DLSVGFRMMNFDQMR-----ESMPDIVNQLASAVKNIYELGGRTFWIHNTAPIGCLPVNL 233
D F + N D ++ E + ++ S +K I++ G + F I N P+GCLP
Sbjct: 593 DYLSPF-LTNSDVLKHYSHTEYVAMVIGNFTSTIKEIHKRGAKKFVILNLPPLGCLPGTR 769
Query: 234 FYKHNLPAGYLDPYGCVKDQNVMAVEFNKQLKDRVVKLRTELPEAAITYVDLYAAKYGLI 293
+ C+++ + +A N+ L + +++L+ +L + D + +I
Sbjct: 770 IIQSQ------GKGSCLEELSSLASIHNQALYEVLLELQKQLRGFKFSLYDFNSDLSHMI 931
Query: 294 SNTKNEGFVDPLKICCGYHVNDTHIWCGTIGTANGKDVFGNACEKPSMYVSWDGVHYAEA 353
++ GF + CCG C G G+ F C+KP+ V WD H E+
Sbjct: 932 NHPLKYGFKEGKSACCGSGPFRGEYSC---GGKRGEKHF-ELCDKPNESVFWDSYHLTES 1099
Query: 354 A 354
A
Sbjct: 1100A 1102
>TC91569 similar to GP|18252233|gb|AAL61949.1 putative APG protein
{Arabidopsis thaliana}, partial (60%)
Length = 1398
Score = 105 bits (263), Expect = 2e-23
Identities = 91/343 (26%), Positives = 152/343 (43%), Gaps = 11/343 (3%)
Frame = +1
Query: 32 PAIYNFGDSNSDTGG---ISAAFEPIPPPYGESFPQ-KPSARDCDGRLIVDFIAEKLNLP 87
PA+ FGDS+ D+G IS + PYG +P+ R +GR+ DFI+E +
Sbjct: 130 PAVIVFGDSSVDSGNNNMISTFLKSNFRPYGRDIDGGRPTGRFSNGRIPPDFISEAFGIK 309
Query: 88 YL-SAYLNSLGT--NYRHGANFATGGSTIRRQNETIFQYGISPFSLDMQFVQFKQFKART 144
L AYL+ T ++ G FA+ G+ N T + P +++F +K+++ +
Sbjct: 310 SLIPAYLDPAYTIDDFVTGVCFASAGTGY--DNATSAILNVIPLWKEVEF--YKEYQDKL 477
Query: 145 KQLYQEAKTALERSKLPVPEEFSKALYTFDIGQNDLS---VGFRMMNFDQMRESMPDIVN 201
K E K+ E S+ALY +G ND GF + F D +
Sbjct: 478 KAHIGEEKSI---------EIISEALYIISLGTNDFLGNYYGFTTLRFRYTISQYQDYLI 630
Query: 202 QLA-SAVKNIYELGGRTFWIHNTAPIGCLPVNLFYKHNLPAGYLDPYGCVKDQNVMAVEF 260
+A + ++ +Y LG R I P+GCLP+ N+ G+ + C + N++A+EF
Sbjct: 631 GIAENFIRQLYSLGARKLAITGLIPMGCLPLERAI--NIFGGF---HRCYEKYNIVALEF 795
Query: 261 NKQLKDRVVKLRTELPEAAITYVDLYAAKYGLISNTKNEGFVDPLKICCGYHVNDTHIWC 320
N +L++ + KL ELP+ ++Y +I+ G + K CC + C
Sbjct: 796 NVKLENMISKLNKELPQLKALSANVYDLFNDIITRPSFYGIEEVEKACCSTGTIEMSYLC 975
Query: 321 GTIGTANGKDVFGNACEKPSMYVSWDGVHYAEAANHWVANRIL 363
+ C+ S Y+ WD H E N ++N ++
Sbjct: 976 NKMNLM--------TCKDASKYMFWDAFHPTEKTNRIISNYLI 1080
>AW688035 similar to PIR|T48618|T486 early nodule-specific protein-like -
Arabidopsis thaliana, partial (13%)
Length = 596
Score = 104 bits (259), Expect = 6e-23
Identities = 52/74 (70%), Positives = 58/74 (78%), Gaps = 1/74 (1%)
Frame = +2
Query: 10 FFLSCTLCVNSVELKNSPP-CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSA 68
FFLSC CV ELK+S P C+FPAIYNFGDSNSDTGGISAAF PIP PYG+ F KP
Sbjct: 32 FFLSCVKCV---ELKDSSPSCSFPAIYNFGDSNSDTGGISAAFVPIPSPYGQGFFHKPFG 202
Query: 69 RDCDGRLIVDFIAE 82
RD DGR+I+DFI +
Sbjct: 203 RDSDGRVILDFIGK 244
>TC83609 similar to GP|5295941|dbj|BAA81842.1 ESTs AU075322(C11109)
D22430(C11109) correspond to a region of the predicted
gene.~Similar to, partial (22%)
Length = 437
Score = 103 bits (257), Expect = 1e-22
Identities = 46/80 (57%), Positives = 60/80 (74%)
Frame = +2
Query: 29 CAFPAIYNFGDSNSDTGGISAAFEPIPPPYGESFPQKPSARDCDGRLIVDFIAEKLNLPY 88
C F AI+NFGDSNSDTGG AAF PYG ++ KP+ R DGRL++DFIA+ + +P+
Sbjct: 191 CDFKAIFNFGDSNSDTGGFYAAFPAESGPYGMTYFNKPAGRASDGRLVIDFIAQAIGIPF 370
Query: 89 LSAYLNSLGTNYRHGANFAT 108
LS YL S+G+ Y+HGAN+AT
Sbjct: 371 LSPYLQSIGSYYKHGANYAT 430
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.138 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,424,568
Number of Sequences: 36976
Number of extensions: 180929
Number of successful extensions: 1092
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 995
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1006
length of query: 381
length of database: 9,014,727
effective HSP length: 98
effective length of query: 283
effective length of database: 5,391,079
effective search space: 1525675357
effective search space used: 1525675357
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0001.6


