

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0627.15
(557 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
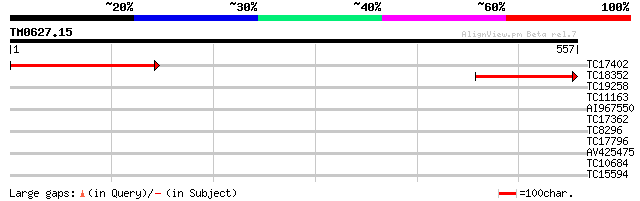
TC17402 weakly similar to UP|O23240 (O23240) Actin interacting p... 311 2e-85
TC18352 similar to UP|O23240 (O23240) Actin interacting protein,... 201 2e-52
TC19258 weakly similar to GB|AAG30907.1|11120512|AF303980 cytoki... 37 0.011
TC11163 similar to UP|Q94AX4 (Q94AX4) AT5g06580/F15M7_11, partia... 37 0.011
AI967550 34 0.071
TC17362 similar to UP|Q942K4 (Q942K4) OJ1111_G12.11 protein, par... 30 0.79
TC8296 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MSJ1... 28 3.0
TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog ... 28 3.0
AV425475 28 3.9
TC10684 28 5.1
TC15594 similar to UP|CYTI_VIGUN (Q06445) Cysteine proteinase in... 27 6.7
>TC17402 weakly similar to UP|O23240 (O23240) Actin interacting protein,
partial (12%)
Length = 494
Score = 311 bits (796), Expect = 2e-85
Identities = 147/147 (100%), Positives = 147/147 (100%)
Frame = +3
Query: 1 MNRNTRATLRLLTPFFNPRRNPSLLPSTPHFDENWWHSQTNIAHGEHRVSSLFNPLRFSV 60
MNRNTRATLRLLTPFFNPRRNPSLLPSTPHFDENWWHSQTNIAHGEHRVSSLFNPLRFSV
Sbjct: 54 MNRNTRATLRLLTPFFNPRRNPSLLPSTPHFDENWWHSQTNIAHGEHRVSSLFNPLRFSV 233
Query: 61 KNQQMGVGISTGIQHKCYASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLST 120
KNQQMGVGISTGIQHKCYASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLST
Sbjct: 234 KNQQMGVGISTGIQHKCYASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLST 413
Query: 121 VNTDWMHKYEGSSKLLLQPWNTDQVSQ 147
VNTDWMHKYEGSSKLLLQPWNTDQVSQ
Sbjct: 414 VNTDWMHKYEGSSKLLLQPWNTDQVSQ 494
>TC18352 similar to UP|O23240 (O23240) Actin interacting protein, partial
(18%)
Length = 547
Score = 201 bits (512), Expect = 2e-52
Identities = 98/100 (98%), Positives = 99/100 (99%)
Frame = +2
Query: 458 GNAANVVGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKA 517
GNAANVVGYGHL DGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKA
Sbjct: 2 GNAANVVGYGHLADGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKA 181
Query: 518 NEILYSKSLETVQLMASIKNLLDPNHILNPYKVLPHSLIA 557
NEILYSKSLETVQLMASIK+LLDPNHILNPYKVLPHSLIA
Sbjct: 182 NEILYSKSLETVQLMASIKSLLDPNHILNPYKVLPHSLIA 301
>TC19258 weakly similar to GB|AAG30907.1|11120512|AF303980 cytokinin oxidase
{Arabidopsis thaliana;} , partial (7%)
Length = 417
Score = 36.6 bits (83), Expect = 0.011
Identities = 26/103 (25%), Positives = 49/103 (47%), Gaps = 1/103 (0%)
Frame = +1
Query: 103 QEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLA-VVP 161
Q+I + D + LS +TD+ H + L+L+P + +S ++K NSR+ +
Sbjct: 82 QQIRPLTSFQSDPKTLSEASTDFGHIVHKTPTLVLEPSSVSDISDLIKQSNSRATPFTIA 261
Query: 162 QGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAG 204
G+ G V+V+++ +N+ F SGI++ G
Sbjct: 262 ARGHAHSYLGQAMAEGGVVVNMTELNR---FRNGSGIVITCGG 381
>TC11163 similar to UP|Q94AX4 (Q94AX4) AT5g06580/F15M7_11, partial (9%)
Length = 590
Score = 36.6 bits (83), Expect = 0.011
Identities = 17/49 (34%), Positives = 28/49 (56%)
Frame = +3
Query: 504 GSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
G+ + HG+G K + E ++ M IK LDP++I+NP K++P
Sbjct: 12 GTCTGXHGVGTGKMKYLEEELGEEALRTMKKIKAALDPHNIMNPGKLIP 158
>AI967550
Length = 412
Score = 33.9 bits (76), Expect = 0.071
Identities = 36/145 (24%), Positives = 58/145 (39%), Gaps = 4/145 (2%)
Frame = +2
Query: 153 NSRSLAVVPQGGNTGLVGGSVPVFDE----VIVSLSSMNKVISFDKVSGILVCEAGCILE 208
NS + + +GG G S V DE VI+ + ++N V S D + E G L
Sbjct: 2 NSARVEIRVRGGGHSYEGTS-SVADEGTLFVIIDMMNLNHV-SVDMETETAWVEGGATLG 175
Query: 209 NIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTV 268
+ + G+ + +GG++ GL +YG +V+ V A+G +
Sbjct: 176 ETYYAISQASSVHGFSAGSSPTVGVGGHIGGGGVGLMSRKYGLAADNVVDALLVDANGQL 355
Query: 269 LDMLKTLRKDNTGYDLKHLFIGSEG 293
LD K+ G D+ G G
Sbjct: 356 LD------KETMGEDVFWAIRGGRG 412
>TC17362 similar to UP|Q942K4 (Q942K4) OJ1111_G12.11 protein, partial (7%)
Length = 432
Score = 30.4 bits (67), Expect = 0.79
Identities = 14/23 (60%), Positives = 16/23 (68%)
Frame = -3
Query: 533 ASIKNLLDPNHILNPYKVLPHSL 555
AS+ LL PNH LNP+ PHSL
Sbjct: 409 ASLSCLLYPNHHLNPHAPSPHSL 341
>TC8296 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MSJ1_4
{Arabidopsis thaliana;}, partial (55%)
Length = 1276
Score = 28.5 bits (62), Expect = 3.0
Identities = 24/76 (31%), Positives = 38/76 (49%)
Frame = +3
Query: 157 LAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDN 216
+ ++ TG+ GG EV++S MN VI +GI+V EAG + I+ +
Sbjct: 462 IVMITDPAETGITGGEAAA--EVMIS---MNVVIGTVGETGIIVAEAGAAVPVWITKV-V 623
Query: 217 EGFIMPLDLGAKGSCQ 232
EG +M + L G+ Q
Sbjct: 624 EGGVMMMSLVEAGADQ 671
>TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog F21C20.180
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 594
Score = 28.5 bits (62), Expect = 3.0
Identities = 27/116 (23%), Positives = 50/116 (42%), Gaps = 3/116 (2%)
Frame = +1
Query: 200 VCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGV 259
V +AG L + + + + G + +GG++S G L +YG +V+
Sbjct: 4 VVQAGATLGELYYGIWQKSKVHGFPAGVCPTVGVGGHISGGGYGNMLRKYGLSVDNVIDA 183
Query: 260 EAVLADGTVLDMLKTLRKDNTGYDLKHLFI-GSEGSLGIVTK--VAILTPPKLSSV 312
+ V G +LD + + G DL G GS G++ V ++ P++ +V
Sbjct: 184 QIVDVQGRILD------RKSMGEDLFWAIRGGGGGSFGVILSYTVKLVKVPEIVTV 333
>AV425475
Length = 344
Score = 28.1 bits (61), Expect = 3.9
Identities = 17/69 (24%), Positives = 31/69 (44%)
Frame = +3
Query: 202 EAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEA 261
+AG + + + + + G S +GG++ A G + +YG +VL
Sbjct: 45 QAGATIGEVYYRISEKSAVHGFPAGLCTSLGVGGHIIGGAYGSMMRKYGLGADNVLDARI 224
Query: 262 VLADGTVLD 270
V A+G +LD
Sbjct: 225 VDANGRILD 251
>TC10684
Length = 564
Score = 27.7 bits (60), Expect = 5.1
Identities = 25/108 (23%), Positives = 48/108 (44%)
Frame = +3
Query: 109 KNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGL 168
K +++ EE+L + M + GS + + + + V+ I Y S A+ G+
Sbjct: 120 KGLIRQEEELEDLKCYVMSFHPGSLQRCARLRSKEAVNLIESY----SFALFNHEGS--- 278
Query: 169 VGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDN 216
GSV DE++ S SS+ +++ G + E ++ + DN
Sbjct: 279 --GSVEKDDEILTSFSSLKRLVLEAVAFGSFLWETEDYIDMLYKLKDN 416
>TC15594 similar to UP|CYTI_VIGUN (Q06445) Cysteine proteinase inhibitor
(Cystatin), complete
Length = 703
Score = 27.3 bits (59), Expect = 6.7
Identities = 10/14 (71%), Positives = 11/14 (78%)
Frame = +3
Query: 17 NPRRNPSLLPSTPH 30
NP NP+LLPS PH
Sbjct: 54 NPTENPNLLPSQPH 95
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,311,270
Number of Sequences: 28460
Number of extensions: 124132
Number of successful extensions: 610
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 609
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 610
length of query: 557
length of database: 4,897,600
effective HSP length: 95
effective length of query: 462
effective length of database: 2,193,900
effective search space: 1013581800
effective search space used: 1013581800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0627.15


