

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0593a.3
(562 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
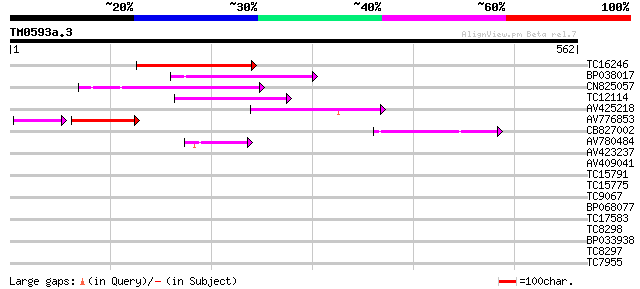
Score E
Sequences producing significant alignments: (bits) Value
TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired re... 175 2e-44
BP038017 106 9e-24
CN825057 92 2e-19
TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transp... 90 1e-18
AV425218 86 1e-17
AV776853 75 1e-16
CB827002 51 6e-07
AV780484 41 4e-04
AV423237 33 0.094
AV409041 33 0.12
TC15791 similar to UP|Q9ZV01 (Q9ZV01) At2g39070 protein, partial... 30 1.0
TC15775 similar to UP|WR17_ARATH (Q9SJA8) Probable WRKY transcri... 29 1.8
TC9067 similar to UP|WR15_ARATH (O22176) Probable WRKY transcrip... 28 3.0
BP068077 28 3.9
TC17583 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired re... 27 6.7
TC8298 homologue to UP|Q9SXP4 (Q9SXP4) DNA-binding protein NtWRK... 27 8.8
BP033938 27 8.8
TC8297 similar to UP|Q40828 (Q40828) WRKY3, partial (27%) 27 8.8
TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcrip... 27 8.8
>TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired response
protein-like, partial (4%)
Length = 660
Score = 175 bits (443), Expect = 2e-44
Identities = 78/119 (65%), Positives = 96/119 (80%)
Frame = +2
Query: 126 DSLKLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQG 185
D + LYGVR+C+I L++G KGGHES+GF K DL NH+ K KR + GDA +ALSYL+G
Sbjct: 2 DGMNLYGVRACHIMALMLGPKGGHESLGFTKTDLSNHIAKPKRERIQNGDAAAALSYLEG 181
Query: 186 KGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNF 244
K + DPMFF+KFT++G+E+LENLFWCDG SR+DY FGDVI F STYKKNKYNKP+V F
Sbjct: 182 KADNDPMFFYKFTKTGDESLENLFWCDGVSRMDYNVFGDVIAFDSTYKKNKYNKPLVVF 358
>BP038017
Length = 570
Score = 106 bits (265), Expect = 9e-24
Identities = 49/146 (33%), Positives = 83/146 (56%)
Frame = +2
Query: 160 YNHVDKKKRISLGAGDAVSALSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDY 219
+ +V ++ +LG DA + L+Y + K+P F++ E + N+FW D SR Y
Sbjct: 125 FTYVKSSQKRTLGR-DAQNLLNYFKKMQGKNPGFYYAIQLDDENRMINVFWADARSRSAY 301
Query: 220 QAFGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEA 279
FGD ++F + Y+ N+Y P F+G NHH + +F CAL+ DE+ ++ W+ + A
Sbjct: 302 NYFGDAVIFDTMYRPNQYQVPFAPFTGVNHHGQNVLFGCALLLDESESSFTWLFRTWLSA 481
Query: 280 MFGKQPKAVVIDGDKSMRAAVKVVFP 305
M + P ++ D D++++AAV VFP
Sbjct: 482 MNDRPPVSITTDQDRAIQAAVAQVFP 559
>CN825057
Length = 721
Score = 92.0 bits (227), Expect = 2e-19
Identities = 57/184 (30%), Positives = 87/184 (46%)
Frame = +1
Query: 69 VTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGDKAQVDSL 128
V + C A V+ KW + F EHNH L PA HF R + +K +D L
Sbjct: 166 VKKTDCKACMHVK-RKPDGKWIIHEFIKEHNHELLPALAYHF-RIHRNVKLAEKNNMDIL 339
Query: 129 KLYGVRSCNITGLLMGQKGGHESVGFLKKDLYNHVDKKKRISLGAGDAVSALSYLQGKGE 188
R+ + + Q GG ++ L DL + K + +++ GDA L Y + +
Sbjct: 340 HAVSERTRKMYVEMSRQSGGCLNIESLVGDLNDQFKKGQYLAMDEGDAQVMLEYFKHIQK 519
Query: 189 KDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYN 248
++P FF+ + E+ L N+FW D S DY +F DV+ F ++Y K+ P F G N
Sbjct: 520 ENPNFFYSIDLNEEQRLRNIFWIDAKSINDYLSFNDVVSFDTSYIKSNEKLPFAPFVGVN 699
Query: 249 HHTE 252
HH +
Sbjct: 700 HHCQ 711
>TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transposase-like
{Oryza sativa (japonica cultivar-group);}, partial (8%)
Length = 488
Score = 89.7 bits (221), Expect = 1e-18
Identities = 43/116 (37%), Positives = 68/116 (58%)
Frame = +1
Query: 164 DKKKRISLGAGDAVSALSYLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFG 223
+K+++ SL G+A L Y Q K ++ F+ + E+ + N+FW D +DY FG
Sbjct: 10 EKRRQRSLLHGEAGYILQYFQRKLVENSPFYHAYQLDDEDQITNVFWVDARMLIDYGYFG 189
Query: 224 DVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEA 279
D++ STY + N+P+ FSG+NHH + IF AL+ DET E+Y W+ ++ EA
Sbjct: 190 DMVSLDSTYCTHSSNRPLAVFSGFNHHRKAVIFGAALLYDETTESY*WLFESFLEA 357
>AV425218
Length = 419
Score = 86.3 bits (212), Expect = 1e-17
Identities = 34/138 (24%), Positives = 75/138 (53%), Gaps = 4/138 (2%)
Frame = +2
Query: 239 KPVVNFSGYNHHTETTIFACALVCDETIETYKWVLKALDEAMFGKQPKAVVIDGDKSMRA 298
+P+ G NHH ++ +F CAL+ E E++ W+ ++L M G P+ ++ D ++M+
Sbjct: 2 RPLATLVGVNHHGQSVLFGCALLSSEDSESFVWLFQSLLHCMSGVPPQGIITDHSEAMKK 181
Query: 299 AVKVVFPNATHRLCGWHIQQNCLEKI----KIPDFLNEFKTLIYGNFTPERFETKWLQVI 354
A++ V P+ HR C +I + +K+ + + + ++Y + FE W +++
Sbjct: 182 AIETVLPSTRHRWCLSYIMKKLPQKLLGYAQYESIRHHLQNVVYDAVVIDEFERNWKKIV 361
Query: 355 EKYGIGEEKWIKQMYETR 372
E +G+ + +W+ +++ R
Sbjct: 362 EDFGLEDNEWLNELFLER 415
>AV776853
Length = 625
Score = 75.5 bits (184), Expect(2) = 1e-16
Identities = 37/67 (55%), Positives = 43/67 (63%)
Frame = +1
Query: 62 RIKEAKPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFIPSFRGLTDGD 121
R ++ KPVT +C AK RV K KWKVV FE HNH LTPA VHFIP +R + D D
Sbjct: 415 RKRDHKPVTHTNCLAKLRVHLDYKIGKWKVVSFEECHNHELTPARFVHFIPPYRVMNDAD 594
Query: 122 KAQVDSL 128
KA+ D L
Sbjct: 595 KAKGDML 615
Score = 27.7 bits (60), Expect(2) = 1e-16
Identities = 19/53 (35%), Positives = 29/53 (53%)
Frame = +2
Query: 4 LCFVLRRMLLNFTNTMPISKVLVQGKMMSVVMQTVILLVVNWCVIGKV*DMRS 56
L V++R +N M KVL+ GKM+ V + +L VN IG+ * ++S
Sbjct: 239 LTLVVKRKHINSIKHMQNIKVLL*GKMILVEIIMGMLTCVNLSAIGRG*GVKS 397
>CB827002
Length = 534
Score = 50.8 bits (120), Expect = 6e-07
Identities = 31/128 (24%), Positives = 61/128 (47%)
Frame = +1
Query: 361 EEKWIKQMYETRQMWATAFMRENFFAGIRTTSLCEGINSFIKSYVLCKNSILDFIYNFER 420
+ +W+ +Y + + A +R+ FFA + T + +NS+ YV ++ F +E+
Sbjct: 31 DHEWLS-LYSSCRQGAPVHLRDTFFAEMSITQRSDSMNSYFDGYVNASTNLNQFFKLYEK 207
Query: 421 AVEEYRHNELASDFKSRYGEPVLITALSHIEQGASKLLTLNMFREMRHMIQRALNLNLVE 480
A+E E+ +D+ + PVL T S +E+ AS+L T +F + + L +
Sbjct: 208 ALESRNEKEVRADYDTMNTLPVLRTP-SPMEKQASELYTRKIFMRFQEELVGTLTFMASK 384
Query: 481 RSEIGTTV 488
+ G +
Sbjct: 385 ADDDGEVI 408
>AV780484
Length = 529
Score = 41.2 bits (95), Expect = 4e-04
Identities = 22/69 (31%), Positives = 39/69 (55%), Gaps = 2/69 (2%)
Frame = -1
Query: 174 GDAVSALS--YLQGKGEKDPMFFFKFTRSGEENLENLFWCDGTSRLDYQAFGDVIVFYST 231
G +V A++ ++ +GE +P FF+ +L ++FW D RLDY+ +++ ++
Sbjct: 217 GRSVKAMTEYFVSTQGE-NPNFFYAIDLDLNRHLTSVFWVDIKGRLDYETSMMLVLIHTH 41
Query: 232 YKKNKYNKP 240
Y KNKY P
Sbjct: 40 YLKNKYKIP 14
>AV423237
Length = 310
Score = 33.5 bits (75), Expect = 0.094
Identities = 16/61 (26%), Positives = 34/61 (55%)
Frame = +2
Query: 212 DGTSRLDYQAFGDVIVFYSTYKKNKYNKPVVNFSGYNHHTETTIFACALVCDETIETYKW 271
D TSR++Y FGD ++F +T + + + + + G + I+ AL+ +E+ ++ W
Sbjct: 17 DATSRMNYSYFGDAVIFDTTI--DNHIESICSSWG*SSWATCVIWLVALIANESESSFVW 190
Query: 272 V 272
+
Sbjct: 191 L 193
>AV409041
Length = 349
Score = 33.1 bits (74), Expect = 0.12
Identities = 15/39 (38%), Positives = 20/39 (50%)
Frame = +3
Query: 67 KPVTRVSCPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPA 105
+P ++ C A V+ + KW V F EHNH L PA
Sbjct: 225 RPSPKIGCKASMHVK-RRQDGKWYVYSFVKEHNHELLPA 338
>TC15791 similar to UP|Q9ZV01 (Q9ZV01) At2g39070 protein, partial (91%)
Length = 556
Score = 30.0 bits (66), Expect = 1.0
Identities = 12/18 (66%), Positives = 14/18 (77%)
Frame = +2
Query: 319 NCLEKIKIPDFLNEFKTL 336
NC+EK+ IP FLN KTL
Sbjct: 482 NCIEKVPIPQFLNGSKTL 535
>TC15775 similar to UP|WR17_ARATH (Q9SJA8) Probable WRKY transcription
factor 17 (WRKY DNA-binding protein 17), partial (53%)
Length = 1284
Score = 29.3 bits (64), Expect = 1.8
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = +3
Query: 74 CPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAH 107
CPA+ V +V +E EH+HG+ A H
Sbjct: 831 CPARKHVERATDDPAMLIVTYEDEHDHGIQTAMH 932
>TC9067 similar to UP|WR15_ARATH (O22176) Probable WRKY transcription
factor 15 (WRKY DNA-binding protein 15), partial (31%)
Length = 660
Score = 28.5 bits (62), Expect = 3.0
Identities = 14/38 (36%), Positives = 19/38 (49%)
Frame = +3
Query: 74 CPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFI 111
CPA+ V + VV +E EHNH L+ A + I
Sbjct: 216 CPARKHVERALDDAAMLVVTYEGEHNHALSAADATNLI 329
>BP068077
Length = 368
Score = 28.1 bits (61), Expect = 3.9
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = -3
Query: 201 GEENLENLFWCDGTSRLDYQAFGDVIVFYSTY 232
G NL NLFWC +S D+ D+ + Y Y
Sbjct: 108 GSRNLWNLFWCTISSLFDFST--DI*IGYQCY 19
>TC17583 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired response
protein, mutator-like transposase-like protein,
phytochrome A signaling protein-like, partial (3%)
Length = 666
Score = 27.3 bits (59), Expect = 6.7
Identities = 10/25 (40%), Positives = 16/25 (64%)
Frame = +3
Query: 217 LDYQAFGDVIVFYSTYKKNKYNKPV 241
+DY+ FGDV+ +TY N +P+
Sbjct: 12 MDYEYFGDVVSLDTTYSTNNAYRPL 86
>TC8298 homologue to UP|Q9SXP4 (Q9SXP4) DNA-binding protein NtWRKY3,
partial (16%)
Length = 514
Score = 26.9 bits (58), Expect = 8.8
Identities = 13/38 (34%), Positives = 18/38 (47%)
Frame = +3
Query: 74 CPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFI 111
CPA+ V +V +E EH H +T A V F+
Sbjct: 192 CPARKHVERAQDDPNMLIVTYEGEHRHLITTTAAVGFM 305
>BP033938
Length = 479
Score = 26.9 bits (58), Expect = 8.8
Identities = 11/27 (40%), Positives = 13/27 (47%)
Frame = -2
Query: 494 ICRHPNTQLSSWSFMTRIRTCLIVIVG 520
+C NTQ+ W F I CL I G
Sbjct: 319 LCEAKNTQIEGWWFFLMIHECLF*IKG 239
>TC8297 similar to UP|Q40828 (Q40828) WRKY3, partial (27%)
Length = 617
Score = 26.9 bits (58), Expect = 8.8
Identities = 13/38 (34%), Positives = 18/38 (47%)
Frame = +1
Query: 74 CPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAHVHFI 111
CPA+ V +V +E EH H +T A V F+
Sbjct: 262 CPARKHVERAQDDPNMLIVTYEGEHRHLITTTAAVGFM 375
>TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcription
factor 11 (WRKY DNA-binding protein 11), partial (27%)
Length = 579
Score = 26.9 bits (58), Expect = 8.8
Identities = 12/34 (35%), Positives = 15/34 (43%)
Frame = +3
Query: 74 CPAKFRVRFHAKSSKWKVVLFEPEHNHGLTPAAH 107
CPA+ V +V +E EH H L A H
Sbjct: 183 CPARKHVERATDDPTMLIVTYEGEHRHALQAAMH 284
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,954,968
Number of Sequences: 28460
Number of extensions: 136807
Number of successful extensions: 878
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 874
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 877
length of query: 562
length of database: 4,897,600
effective HSP length: 95
effective length of query: 467
effective length of database: 2,193,900
effective search space: 1024551300
effective search space used: 1024551300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0593a.3


