

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
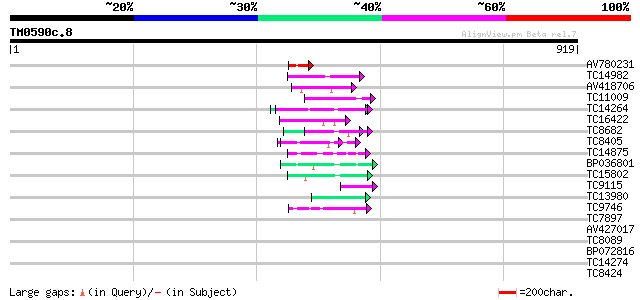
Query= TM0590c.8
(919 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV780231 58 6e-09
TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%) 51 9e-07
AV418706 48 8e-06
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 47 1e-05
TC14264 similar to UP|Q84NF9 (Q84NF9) Histone H1 (Fragment), par... 46 3e-05
TC16422 similar to PIR|T05653|T05653 amino acid transport protei... 46 3e-05
TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate... 44 9e-05
TC8405 weakly similar to UP|O04083 (O04083) Lysophospholipase is... 44 2e-04
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 44 2e-04
BP036801 43 3e-04
TC15802 similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cucumber... 43 3e-04
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 42 3e-04
TC13980 similar to UP|Q9SGT0 (Q9SGT0) T6H22.18 protein (F14J16.3... 42 6e-04
TC9746 41 7e-04
TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-li... 40 0.001
AV427017 40 0.001
TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinog... 40 0.002
BP072816 40 0.002
TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.3... 40 0.002
TC8424 weakly similar to GB|BAC22265.1|24414015|AP003803 arm rep... 39 0.003
>AV780231
Length = 530
Score = 58.2 bits (139), Expect = 6e-09
Identities = 31/40 (77%), Positives = 33/40 (82%)
Frame = -3
Query: 453 SKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPAS 492
S PGAQPTTAAA KGKNV EASVAAA EPT VPAS+P +
Sbjct: 477 SNPGAQPTTAAAR-KGKNVNEASVAAAIEPTTVPASTPVT 361
Score = 37.7 bits (86), Expect = 0.008
Identities = 17/17 (100%), Positives = 17/17 (100%)
Frame = -3
Query: 307 AKSSFARKYFRVAVHPA 323
AKSSFARKYFRVAVHPA
Sbjct: 528 AKSSFARKYFRVAVHPA 478
>TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%)
Length = 1036
Score = 50.8 bits (120), Expect = 9e-07
Identities = 33/125 (26%), Positives = 58/125 (46%), Gaps = 1/125 (0%)
Frame = +3
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAV-PASSPASAGATAAFAAESSIGATA 509
G++ P P A P ++ +T PTA+ A +P++A AA +A S+ G ++
Sbjct: 39 GSTLPPKPPPPQTAPPPPTQLSPPQRPNSTAPTALLTALNPSTAAPAAAASAPSASG*SS 218
Query: 510 ASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSC 569
+S+ +++SS + + P +PP P + P++ P PSPP+ S
Sbjct: 219 SSS------SSSSSSAVLASPSTSSTAPTTPPSPSPPSNSPTSTSPPPQPSPPNSTSPSP 380
Query: 570 PGAAT 574
P T
Sbjct: 381 PQTPT 395
Score = 36.2 bits (82), Expect = 0.024
Identities = 31/118 (26%), Positives = 51/118 (42%), Gaps = 15/118 (12%)
Frame = +3
Query: 459 PTTAAAAPKGKNVAEASVAAATEP-TAVPA---SSPASAGATAAFAAESSIGATAASAGV 514
PT + + + A ++ A P TA PA S+P+++G +++ ++ SS A AS
Sbjct: 90 PTQLSPPQRPNSTAPTALLTALNPSTAAPAAAASAPSASG*SSSSSSSSSSSAVLASPST 269
Query: 515 NATKAAA-----------SSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSP 561
++T +S +P N T SPP+ P S P+PSP
Sbjct: 270 SSTAPTTPPSPSPPSNSPTSTSPPPQPSPPNSTSPSPPQ--TPTRKTSHSPTYPLPSP 437
Score = 28.1 bits (61), Expect = 6.6
Identities = 19/49 (38%), Positives = 21/49 (42%), Gaps = 7/49 (14%)
Frame = +3
Query: 546 PPSPPSTHDAGPMP---SPPHQGEKSCPGAATT----SEAAQIEQAPAP 587
PP PP A P P SPP + + P A T S AA A AP
Sbjct: 51 PPKPPPPQTAPPPPTQLSPPQRPNSTAPTALLTALNPSTAAPAAAASAP 197
>AV418706
Length = 374
Score = 47.8 bits (112), Expect = 8e-06
Identities = 33/115 (28%), Positives = 52/115 (44%), Gaps = 11/115 (9%)
Frame = +1
Query: 458 QPTTAAAAPKGKNV----AEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAG 513
QP+ A AP + A T P VP P +A A AA SS G++++S+
Sbjct: 22 QPSPPAMAPPPPTSHYRRQNPNSTAPTAPPTVPVLHPLAAAAATVAAAHSSAGSSSSSSS 201
Query: 514 VNATKA----AASSDTPIGDKEKENETPKSPPRQDA---PPSPPSTHDAGPMPSP 561
+++ A +++S T + + PP PPSPPS++ P P+P
Sbjct: 202 SSSSSAPPAPSSTSSTTLNALPSPSRHSNFPPSNSPPPPPPSPPSSNSHSPPPTP 366
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 47.0 bits (110), Expect = 1e-05
Identities = 32/117 (27%), Positives = 57/117 (48%), Gaps = 3/117 (2%)
Frame = -2
Query: 479 ATEPTAVPASSPASA-GATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP 537
A P+++ SSP+S+ G+ + A+ SS AT + K+++SS + + +
Sbjct: 354 AFSPSSLSLSSPSSSSGSKSLDASSSSAAATVRRTRYSVPKSSSSSSSSSSSSSSSSSSS 175
Query: 538 KSPPRQDAPPSPP--STHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSS 592
SPP P +PP S+ + SPP PGA+++S ++ +P+ SS
Sbjct: 174 GSPPPPRPPSTPPSSSSSSSSSSSSPP-------PGASSSSSSSPSSSSPSSSSSSS 25
Score = 34.3 bits (77), Expect = 0.092
Identities = 29/136 (21%), Positives = 59/136 (43%), Gaps = 8/136 (5%)
Frame = -2
Query: 478 AATEPTAVPASSPASA-------GATAAFAAES-SIGATAASAGVNATKAAASSDTPIGD 529
+++ P + +SSP + T AF+ S S+ + ++S+G + A++SS
Sbjct: 435 SSSSPPSSESSSPEPSLSDLFLFDGTPAFSPSSLSLSSPSSSSGSKSLDASSSSAAATVR 256
Query: 530 KEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEV 589
+ + + S + S S+ + P PP P +S ++ + +P
Sbjct: 255 RTRYSVPKSSSSSSSSSSSSSSSSSSSGSPPPPRP-----PSTPPSSSSSSSSSSSSPPP 91
Query: 590 GSSSYYNMLPNAIEPS 605
G+SS + P++ PS
Sbjct: 90 GASSSSSSSPSSSSPS 43
>TC14264 similar to UP|Q84NF9 (Q84NF9) Histone H1 (Fragment), partial (76%)
Length = 1277
Score = 45.8 bits (107), Expect = 3e-05
Identities = 46/165 (27%), Positives = 67/165 (39%), Gaps = 3/165 (1%)
Frame = +2
Query: 424 KRGGAENQAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPT 483
K A +A++ APK +KA++ T+K A+P AAA PK K A+A AA +
Sbjct: 548 KPAAAATKAKAKPAPK----SKAAATKTTAK--AKP-AAAAKPKAKPAAKAKPAAKAKAV 706
Query: 484 AVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQ 543
A PA + AS A A + + V K AA + + + ++ P +
Sbjct: 707 AAPAKAKASPAKPKAKAKSKTAPRMNVNPPVPKAKPAAKAKPAARPAKASRTSTRTTPGK 886
Query: 544 DAPPSPPSTHDAGPMPSPPHQGEKSCP---GAATTSEAAQIEQAP 585
PP P P P K P G A T ++ AP
Sbjct: 887 KVPPPPK------PAPKKAATPVKKAPAKSGKAKTVKSPAKRTAP 1003
Score = 43.5 bits (101), Expect = 2e-04
Identities = 44/157 (28%), Positives = 64/157 (40%), Gaps = 1/157 (0%)
Frame = +2
Query: 432 AESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGK-NVAEASVAAATEPTAVPASSP 490
A ++K ++ + ++ KP A+ AAAA K K A S AAAT+ TA + P
Sbjct: 464 AAASKPAAAKKKPQPAAAKSKPKPKAKAKPAAAATKAKAKPAPKSKAAATKTTA--KAKP 637
Query: 491 ASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPP 550
A+A A A + A A A KA AS P K K+ PR + P P
Sbjct: 638 AAAAKPKAKPAAKAKPAAKAKAVAAPAKAKASPAKP-----KAKAKSKTAPRMNVNPPVP 802
Query: 551 STHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAP 587
A P + K+ + T+ ++ P P
Sbjct: 803 KAKPAA-KAKPAARPAKASRTSTRTTPGKKVPPPPKP 910
>TC16422 similar to PIR|T05653|T05653 amino acid transport protein homolog
F22I13.20 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (38%)
Length = 685
Score = 45.8 bits (107), Expect = 3e-05
Identities = 35/136 (25%), Positives = 60/136 (43%), Gaps = 21/136 (15%)
Frame = +3
Query: 438 PKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPA---SSPASAG 494
P + +S + S P ++P+ +++P + AS+ A++EP PA SSP+ +
Sbjct: 135 PTLEKTHLSSPNPHLSPPNSKPSPTSSSPSSAPESSASLTASSEPAGSPASSCSSPSPSS 314
Query: 495 ATAAFAAESSIGA-------------TAASAGVNATKAAASSDT-----PIGDKEKENET 536
T A + S + A +A SA +A ++AAS T P D +
Sbjct: 315 PTTA*CSSSPLAASSIPSLGTPNSDPSATSASPSAARSAASPSTP*LSSPRLDSASATSS 494
Query: 537 PKSPPRQDAPPSPPST 552
P +P +PP+T
Sbjct: 495 SSRPLSASSPATPPAT 542
>TC8682 similar to UP|HPPD_SOLSC (Q9ARF9) 4-hydroxyphenylpyruvate
dioxygenase (4HPPD) (HPD) (HPPDase) , partial (29%)
Length = 576
Score = 44.3 bits (103), Expect = 9e-05
Identities = 32/135 (23%), Positives = 51/135 (37%), Gaps = 3/135 (2%)
Frame = +1
Query: 444 TKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPT---AVPASSPASAGATAAFA 500
+K S++ S + PT K +S A T PT A P +S +
Sbjct: 172 SKPGSNSSGSTASSGPTQNQTGSP*KGSTTSSSGAPTPPTPHAASPGASACRSSQNQTSP 351
Query: 501 AESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPS 560
E+++ +SA + ++ P +P SPP +P PP+ + P PS
Sbjct: 352 PETTLTPPTSSAPATSASSSPPHTLPKSPSPPPPPSPLSPPPPASPSPPPTASPSAPSPS 531
Query: 561 PPHQGEKSCPGAATT 575
K P +TT
Sbjct: 532 KSPTPSKPSPPQSTT 576
Score = 42.7 bits (99), Expect = 3e-04
Identities = 32/115 (27%), Positives = 47/115 (40%), Gaps = 5/115 (4%)
Frame = +1
Query: 478 AATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP 537
A++ PT SP T++ A + AAS G +A +++ + +P ET
Sbjct: 205 ASSGPTQNQTGSP*KGSTTSSSGAPTPPTPHAASPGASACRSSQNQTSP-------PETT 363
Query: 538 KSPPRQDAPP-----SPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAP 587
+PP AP SPP T P P PP P A+ + AP+P
Sbjct: 364 LTPPTSSAPATSASSSPPHTLPKSPSPPPPPSPLSPPPPASPSPPPTASPSAPSP 528
>TC8405 weakly similar to UP|O04083 (O04083) Lysophospholipase isolog,
25331-24357 (At1g11090), partial (18%)
Length = 552
Score = 43.5 bits (101), Expect = 2e-04
Identities = 36/130 (27%), Positives = 59/130 (44%), Gaps = 1/130 (0%)
Frame = +3
Query: 440 RRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVP-ASSPASAGATAA 498
R+R T ++ T P + TA ++P + + S + P++ P A++P AG++
Sbjct: 87 RKRNTTPPKESATPNPTSTLPTARSSP---SPSSPSTPKSKPPSS*PTATAPTPAGSSRR 257
Query: 499 FAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPM 558
A+ S GAT +S ++ AA P + SPP P SP S+ A
Sbjct: 258 SASTSPPGATPSSPPTSSATAA-----PTASAATSATSTASPP----PRSPSSSTSAAAS 410
Query: 559 PSPPHQGEKS 568
P+PP + S
Sbjct: 411 PTPPFRPSSS 440
Score = 41.6 bits (96), Expect = 6e-04
Identities = 35/114 (30%), Positives = 49/114 (42%), Gaps = 6/114 (5%)
Frame = +3
Query: 434 STKAPKRRRLTKASSDAGTSKPGAQ-PTTAAAAPKGKNVAEASVAAATEPTAVPASSPAS 492
ST R + +S SKP + PT A P G + AS T P SSP +
Sbjct: 138 STLPTARSSPSPSSPSTPKSKPPSS*PTATAPTPAGSSRRSAS----TSPPGATPSSPPT 305
Query: 493 AGATAAFAAESSIGATAASAGV-----NATKAAASSDTPIGDKEKENETPKSPP 541
+ ATAA A ++ AT+ ++ ++T AAAS P + SPP
Sbjct: 306 SSATAAPTASAATSATSTASPPPRSPSSSTSAAASPTPPFRPSSSASPWAGSPP 467
Score = 32.3 bits (72), Expect = 0.35
Identities = 27/110 (24%), Positives = 41/110 (36%), Gaps = 6/110 (5%)
Frame = +3
Query: 490 PASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSP 549
P ++GAT + +A + A SS +P +++ P S P AP
Sbjct: 63 PLTSGATCRKRNTTPPKESATPNPTSTLPTARSSPSPSSPSTPKSKPPSS*PTATAPTPA 242
Query: 550 ------PSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
ST G PS P + A+ + +A +P P SSS
Sbjct: 243 GSSRRSASTSPPGATPSSPPTSSATAAPTASAATSATSTASPPPRSPSSS 392
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 43.5 bits (101), Expect = 2e-04
Identities = 41/136 (30%), Positives = 59/136 (43%)
Frame = +1
Query: 450 AGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATA 509
A T P + P T+ + P AS A+T P P +SP+S+ +TAA +
Sbjct: 121 APTI*PPSNPKTSTSTP-----TPASPPASTSPLTPPTTSPSSSTSTAA---------PS 258
Query: 510 ASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSC 569
ASA + T A+S T +TP S P A SP ST P +P G+ S
Sbjct: 259 ASAPLTTTPTTATSTT-----SPRRQTPSSSPSTTA--SPRSTPSPSPTTTP---GKPSS 408
Query: 570 PGAATTSEAAQIEQAP 585
A+T+ ++ P
Sbjct: 409 GLPASTTRGSRTTPIP 456
Score = 28.9 bits (63), Expect = 3.8
Identities = 20/62 (32%), Positives = 27/62 (43%), Gaps = 4/62 (6%)
Frame = +1
Query: 536 TPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAA----TTSEAAQIEQAPAPEVGS 591
TP SPP +P +PP+T SP + P A+ TT A +P + S
Sbjct: 172 TPASPPASTSPLTPPTT-------SPSSSTSTAAPSASAPLTTTPTTATSTTSPRRQTPS 330
Query: 592 SS 593
SS
Sbjct: 331 SS 336
Score = 28.9 bits (63), Expect = 3.8
Identities = 23/76 (30%), Positives = 31/76 (40%), Gaps = 5/76 (6%)
Frame = +1
Query: 535 ETPKSPPRQDAPPSPPSTHDAGPMP-SPPHQGEKSCP----GAATTSEAAQIEQAPAPEV 589
+TP P PPS P T + P P SPP P +++TS AA AP
Sbjct: 103 QTPPLPAPTI*PPSNPKTSTSTPTPASPPASTSPLTPPTTSPSSSTSTAAPSASAPLTTT 282
Query: 590 GSSSYYNMLPNAIEPS 605
+++ P PS
Sbjct: 283 PTTATSTTSPRRQTPS 330
>BP036801
Length = 540
Score = 42.7 bits (99), Expect = 3e-04
Identities = 46/162 (28%), Positives = 65/162 (39%), Gaps = 4/162 (2%)
Frame = +2
Query: 439 KRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSP----ASAG 494
K + L +SS S A P AAA +G + A A++P + + +S
Sbjct: 8 KNKPLLLSSSP*SPSCCSASPQHAAA--QGGTGFSYNAA*ASQPCTCRS*TTTRWSSSTA 181
Query: 495 ATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHD 554
T+A S A AAS V ++ + TP + SPP A TH
Sbjct: 182 PTSAHPTSPSPTAAAASTPVTWPSSSTAPPTP--------SSTTSPPTLSALSPSKPTHG 337
Query: 555 AGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYYN 596
A P PSPP S P A+TT+ + PA S++ N
Sbjct: 338 APPAPSPP-TAPSSKPAASTTATTSSAPSPPAHNTTSATGSN 460
>TC15802 similar to UP|Q9XIV1 (Q9XIV1) mRNA expressed in cucumber hypocotyls
(Arabinogalactan protein), partial (26%)
Length = 795
Score = 42.7 bits (99), Expect = 3e-04
Identities = 37/144 (25%), Positives = 53/144 (36%), Gaps = 7/144 (4%)
Frame = +2
Query: 451 GTSKPGAQPTTAAAAPKGKNVAEASVA-------AATEPTAVPASSPASAGATAAFAAES 503
G P A PTT+ A P VA AAT P A P+++ A A AT
Sbjct: 101 GGQSPTAAPTTSPATPSATTPVSTPVAPVASPKPAATTPAASPSTATAPAPATTTPVTPV 280
Query: 504 SIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPH 563
+ AA + AA +P ++P +P P + P T P P+P
Sbjct: 281 TSPPPAAVPVASPPPAAVPVSSPPA------KSPPAPAPTAVPVAAPVTTPEVPAPAPSK 442
Query: 564 QGEKSCPGAATTSEAAQIEQAPAP 587
+ + P + + AP P
Sbjct: 443 TKKDAAPAPSPVVPDSPPAGAPGP 514
Score = 38.1 bits (87), Expect = 0.006
Identities = 37/146 (25%), Positives = 53/146 (35%), Gaps = 1/146 (0%)
Frame = +2
Query: 437 APKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGAT 496
+PK T A+S + + P A TT P A A+ P AVP SSP +
Sbjct: 191 SPKPAATTPAASPSTATAP-APATTTPVTPVTSPPPAAVPVASPPPAAVPVSSPPAKSPP 367
Query: 497 AAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETP-KSPPRQDAPPSPPSTHDA 555
A + A + V A P K K++ P SP D+PP+
Sbjct: 368 APAPTAVPVAAPVTTPEVPA---------PAPSKTKKDAAPAPSPVVPDSPPAGAPGPSD 520
Query: 556 GPMPSPPHQGEKSCPGAATTSEAAQI 581
P P G + A T + ++
Sbjct: 521 AVSPGPDGAGTANDESGAETMRSLKM 598
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 42.4 bits (98), Expect = 3e-04
Identities = 20/59 (33%), Positives = 28/59 (46%)
Frame = +1
Query: 537 PKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSSYY 595
P SPP AP +P + +P+PPH G+ATTS +A + P S +Y
Sbjct: 337 PPSPPTPSAPLTP*APTAPSTLPTPPHSAAPPSSGSATTSASATTRLSTPPTTNRSQFY 513
Score = 28.9 bits (63), Expect = 3.8
Identities = 20/70 (28%), Positives = 26/70 (36%), Gaps = 12/70 (17%)
Frame = -2
Query: 526 PIGDKEKENETPKSPPRQDAPP-SPPSTHDAGPMPSPPHQGEKSC-----------PGAA 573
P +K K P+ +PP SPPS+ P PH + C P
Sbjct: 861 PPTEKANPQHGTKCSPKSTSPPHSPPSSPPQSSPPPSPHHAKPLCEHTQRQHQQPSPAPP 682
Query: 574 TTSEAAQIEQ 583
T S A Q+ Q
Sbjct: 681 TQSPACQL*Q 652
>TC13980 similar to UP|Q9SGT0 (Q9SGT0) T6H22.18 protein (F14J16.33), partial
(12%)
Length = 565
Score = 41.6 bits (96), Expect = 6e-04
Identities = 29/101 (28%), Positives = 39/101 (37%), Gaps = 5/101 (4%)
Frame = +2
Query: 490 PASAGATAAFAAESSIGATAASAGVNATKAAAS---SDTPIGDKEKENETPKSPPRQDAP 546
PAS +A A+ I S+ + T + S S +P G P PP Q P
Sbjct: 242 PASTTPSATRASSLGIPNPPTSSSASKTPSPCSPPTSSSPTGSSSHSTSNPPHPPPQQQP 421
Query: 547 PSPPSTHDAGPMPSPP--HQGEKSCPGAATTSEAAQIEQAP 585
SPP P P+ P H + P A S A++ P
Sbjct: 422 FSPPPPPPPPPSPTAPSSHPRLPAAPAAGRISSASKSSTTP 544
>TC9746
Length = 637
Score = 41.2 bits (95), Expect = 7e-04
Identities = 40/144 (27%), Positives = 63/144 (42%), Gaps = 9/144 (6%)
Frame = +2
Query: 452 TSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAAS 511
TSKP QP P + A +S A+ +EP + P +SPA G A + SS + S
Sbjct: 182 TSKPKCQPP-----PHPSSTATSSPASNSEPPSGP-NSPARWGT*APTSPSSS--PSL*S 337
Query: 512 AGVNATKAAASSDTPIGDKEKENETP-KSPPRQDAPPSPPSTHDAGP--------MPSPP 562
+ + ++S + + +P +S P +PPSP H P PSPP
Sbjct: 338 TTLTSPPHSSSPHSTTSPLASSSASPCQSSP*SPSPPSPSPKHPHSPSPRSPPPASPSPP 517
Query: 563 HQGEKSCPGAATTSEAAQIEQAPA 586
+ + P + +S A + Q+ A
Sbjct: 518 YFSSSAPPVSCPSSTATSLCQSSA 589
>TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-like protein
2 precursor (Phytocyanin-like protein), partial (31%)
Length = 1482
Score = 40.4 bits (93), Expect = 0.001
Identities = 49/200 (24%), Positives = 69/200 (34%), Gaps = 22/200 (11%)
Frame = +3
Query: 431 QAESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAV----- 485
++ + +PK + + AG P P + +P G V +A T P+A
Sbjct: 549 KSSPSPSPKASPPSGSPPTAGEPSPAGTPPSVLPSPAGAPVPAGGPSAGTPPSASTSPSP 728
Query: 486 -----------------PASSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIG 528
P+SSP S + + G AT + + D+P G
Sbjct: 729 SPVSAPPAASPAPGAESPSSSPTS---------------NSPAGGPTATSPSPAGDSPAG 863
Query: 529 DKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPE 588
SP D PPS +GP SP G S A T S ++ AP P
Sbjct: 864 -----GPPAPSPTTGDTPPSA-----SGPDVSP--SGTPSSLPADTPSSSSNNSTAPPPG 1007
Query: 589 VGSSSYYNMLPNAIEPSEFL 608
G SS + PS FL
Sbjct: 1008NGGSS--------VVPSSFL 1043
Score = 31.2 bits (69), Expect = 0.78
Identities = 20/57 (35%), Positives = 25/57 (43%)
Frame = +3
Query: 537 PKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAPEVGSSS 593
P PP+ PSP ++ +G SPP GE S P S AP P G S+
Sbjct: 534 PSPPPKSSPSPSPKASPPSG---SPPTAGEPS-PAGTPPSVLPSPAGAPVPAGGPSA 692
>AV427017
Length = 429
Score = 40.4 bits (93), Expect = 0.001
Identities = 33/121 (27%), Positives = 47/121 (38%)
Frame = +3
Query: 459 PTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNATK 518
PTTA + + T A AS+PASA A+AA AA S ++A+S A
Sbjct: 9 PTTAPPFRRRNRNPTTAQPTTTTAEAAAASAPASAAASAAAAAASPTASSASS----ARS 176
Query: 519 AAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQGEKSCPGAATTSEA 578
+ SS +P + A P P S + P + P T+E+
Sbjct: 177 SPPSSSSPPSPASSSG*SSAPTSSNSASPKPHSLNSPTPTTLSTTISPSTSPSETLTAES 356
Query: 579 A 579
A
Sbjct: 357 A 359
Score = 39.3 bits (90), Expect = 0.003
Identities = 35/120 (29%), Positives = 59/120 (49%)
Frame = +3
Query: 432 AESTKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPA 491
A +T P RRR + + T++P AAAA + A ++ AAA PTA ASS
Sbjct: 6 APTTAPPFRRR----NRNPTTAQPTTTTAEAAAASAPASAAASAAAAAASPTASSASSAR 173
Query: 492 SAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPS 551
S+ ++ SS + A+S+G ++ +++S +P + TP + +P + PS
Sbjct: 174 SSPPSS-----SSPPSPASSSG*SSAPTSSNSASP-KPHSLNSPTPTTLSTTISPSTSPS 335
Score = 35.4 bits (80), Expect = 0.041
Identities = 33/131 (25%), Positives = 49/131 (37%), Gaps = 10/131 (7%)
Frame = +3
Query: 480 TEPTAVPA-----SSPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKEN 534
T PT P +P +A T AE++ + ASA +A AAAS +
Sbjct: 3 TAPTTAPPFRRRNRNPTTAQPTTT-TAEAAAASAPASAAASAAAAAASPTA-----SSAS 164
Query: 535 ETPKSPPRQDAPPSPPSTHDAGPMP-----SPPHQGEKSCPGAATTSEAAQIEQAPAPEV 589
SPP +PPSP S+ P + P + P T S +P+ +
Sbjct: 165 SARSSPPSSSSPPSPASSSG*SSAPTSSNSASPKPHSLNSPTPTTLSTTISPSTSPSETL 344
Query: 590 GSSSYYNMLPN 600
+ S P+
Sbjct: 345 TAESASTTTPS 377
>TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (43%)
Length = 890
Score = 39.7 bits (91), Expect = 0.002
Identities = 42/161 (26%), Positives = 58/161 (35%), Gaps = 7/161 (4%)
Frame = +2
Query: 434 STKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPAS----- 488
S + P R + SS P PTT+ A + S + PT+ P S
Sbjct: 155 SARQPWRLSRQRCSSSPWLLPPQT-PTTSPAFSPSTPSSPPSTTTSPSPTSPPRSTNGPQ 331
Query: 489 SPASAGATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQD-APP 547
SP++ T + S T S + + SS T P+S R APP
Sbjct: 332 SPSAPSTTPPWTTSSQ--NTHQSPPSRTSSPSTSSST--------TSAPRSSTRSPTAPP 481
Query: 548 SPPS-THDAGPMPSPPHQGEKSCPGAATTSEAAQIEQAPAP 587
SPP T P +PP A + A + AP+P
Sbjct: 482 SPPQCTKPPAPPRAPPDSSTSPTSAAGKSDSAPRTTTAPSP 604
>BP072816
Length = 492
Score = 39.7 bits (91), Expect = 0.002
Identities = 41/151 (27%), Positives = 62/151 (40%), Gaps = 5/151 (3%)
Frame = +1
Query: 447 SSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIG 506
SS + +S P P T AP +T P++ P+++ A +T AA +S
Sbjct: 118 SSSSSSSSPLPSPPTTLHAP------------STSPSSAPSATAAPPPSTPTPAANTSSK 261
Query: 507 ATAASAGVNATKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMP---SPPH 563
+ +S + ASS +PP+ PPS PS+ ++ P P S PH
Sbjct: 262 PSTSSKPTTSAAPPASS------------LLSTPPKPAGPPSNPSSTNS-PAPASTSAPH 402
Query: 564 QGEKS--CPGAATTSEAAQIEQAPAPEVGSS 592
K+ P A TS A P+P+ S
Sbjct: 403 AASKAHGSPKVA*TSPACN-NSKPSPQTPHS 492
Score = 33.1 bits (74), Expect = 0.20
Identities = 30/106 (28%), Positives = 48/106 (44%)
Frame = +1
Query: 434 STKAPKRRRLTKASSDAGTSKPGAQPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASA 493
+T AP T A++ + ++PTT+AA P AS +T P PA P++
Sbjct: 205 ATAAPPPSTPTPAANTSSKPSTSSKPTTSAAPP-------ASSLLSTPPK--PAGPPSNP 357
Query: 494 GATAAFAAESSIGATAASAGVNATKAAASSDTPIGDKEKENETPKS 539
+T + A S+ AAS + K A +S + + +TP S
Sbjct: 358 SSTNSPAPASTSAPHAASKAHGSPKVA*TSPA-CNNSKPSPQTPHS 492
Score = 31.6 bits (70), Expect = 0.59
Identities = 19/64 (29%), Positives = 29/64 (44%), Gaps = 4/64 (6%)
Frame = +1
Query: 534 NETPKSPPRQDAPPSPPSTHDAG--PMPSPPH--QGEKSCPGAATTSEAAQIEQAPAPEV 589
+ T +PP P PS+ + P+PSPP + P +A ++ AA P P
Sbjct: 67 SNTHHNPPLSHQPCHSPSSSSSSSSPLPSPPTTLHAPSTSPSSAPSATAAPPPSTPTPAA 246
Query: 590 GSSS 593
+SS
Sbjct: 247 NTSS 258
>TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.35
{Arabidopsis thaliana;} , partial (16%)
Length = 1208
Score = 39.7 bits (91), Expect = 0.002
Identities = 35/133 (26%), Positives = 51/133 (38%), Gaps = 22/133 (16%)
Frame = +3
Query: 479 ATEPTAVPASSPASAGATAAFAAESSIGATAASAG----------------VNATKAAAS 522
+T P++ P+ S S G A + SS G+T + V+ + +
Sbjct: 78 STSPSSPPSISRTSTGKPATSSRTSSSGSTPTAVSPPNPTTPATPPPSGTSVSPSLSLTP 257
Query: 523 SDTPIGDKEKENETPKSPPRQDAPP----SPPSTHDAGPMPS--PPHQGEKSCPGAATTS 576
S TP P PP +PP SP ST P S P + P A +TS
Sbjct: 258 SLTPFSPSRSSTPNPPIPPNPSSPPSASSSPTSTTSMIPPASANSPSSAPPAAPTARSTS 437
Query: 577 EAAQIEQAPAPEV 589
+A + AP P +
Sbjct: 438 SSASL-VAPNPSI 473
>TC8424 weakly similar to GB|BAC22265.1|24414015|AP003803 arm repeat
containing protein homolog-like {Oryza sativa (japonica
cultivar-group);} , partial (49%)
Length = 1586
Score = 39.3 bits (90), Expect = 0.003
Identities = 32/141 (22%), Positives = 54/141 (37%), Gaps = 12/141 (8%)
Frame = -3
Query: 458 QPTTAAAAPKGKNVAEASVAAATEPTAVPASSPASAGATAAFAAESSIGATAASAGVNA- 516
+P + AP + + S+A +P PA+SP + +A S TA SA ++
Sbjct: 819 EPQPSC*APPSPSPPDQSLAPPQDP---PAASPLQRSPPSPCSASQSTACTALSAHPSSP 649
Query: 517 --TKAAASSDTPIGDKEKENETPKSPPRQDAPPSPPSTHDAGPMPSPPHQG--------- 565
+A+ + +++ + + PP Q + P + P PS H+
Sbjct: 648 YWNHLSAAKPAAVSPRKRRSSSCSPPPTQGSAKHTPRSASKSPSPSSAHKSTTSQLRHWL 469
Query: 566 EKSCPGAATTSEAAQIEQAPA 586
SC T A APA
Sbjct: 468 SASCSHH*VTGSTAPSPHAPA 406
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.128 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,286,333
Number of Sequences: 28460
Number of extensions: 256865
Number of successful extensions: 4967
Number of sequences better than 10.0: 393
Number of HSP's better than 10.0 without gapping: 2907
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4023
length of query: 919
length of database: 4,897,600
effective HSP length: 99
effective length of query: 820
effective length of database: 2,080,060
effective search space: 1705649200
effective search space used: 1705649200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0590c.8


