

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.5
(441 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
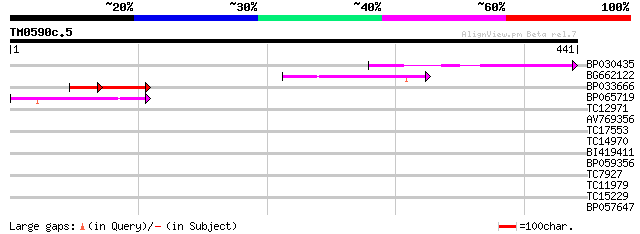
Score E
Sequences producing significant alignments: (bits) Value
BP030435 119 1e-27
BG662122 83 8e-17
BP033666 39 9e-08
BP065719 51 3e-07
TC12971 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14... 30 1.0
AV769356 28 2.3
TC17553 weakly similar to UP|AAP20176 (AAP20176) Kinase substrat... 28 2.3
TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) prec... 28 2.3
BI419411 28 2.3
BP059356 28 3.0
TC7927 similar to UP|Q93YR3 (Q93YR3) HSP associated protein like... 28 3.9
TC11979 similar to UP|Q9LVH4 (Q9LVH4) GTPase activating protein-... 28 3.9
TC15229 weakly similar to UP|O60585 (O60585) Ser/Arg-related nuc... 27 6.7
BP057647 27 8.8
>BP030435
Length = 533
Score = 119 bits (297), Expect = 1e-27
Identities = 72/162 (44%), Positives = 87/162 (53%)
Frame = +1
Query: 280 CEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKREEERADTKGKK 339
C +H++ GH T++C TLQRE+ KLI G K +
Sbjct: 103 CVYHQAMGHITEECRTLQREVGKLIATG----------------------------KPTR 198
Query: 340 KQESAAIATKGADDTFARHSGPPVGTINTIAGGFGGGGDTHAARKRHVRAVNSVHEVAFG 399
Q A KG ++TIAGGFGGGG T AARKR+ RAVN+V EV FG
Sbjct: 199 IQGGNAPVPKG---------------VHTIAGGFGGGGITSAARKRYARAVNTVTEVLFG 333
Query: 400 FVHPDITISMADFEGIKPHKDDPIVVQLRMNSFNVRRVLLDQ 441
F HP IT S DF IKPH D+PIV+ LR+N NV+RVLLDQ
Sbjct: 334 FSHPVITFSSDDFIRIKPHLDNPIVILLRVNQLNVQRVLLDQ 459
>BG662122
Length = 386
Score = 83.2 bits (204), Expect = 8e-17
Identities = 50/126 (39%), Positives = 74/126 (58%), Gaps = 11/126 (8%)
Frame = +2
Query: 213 KEQLYPRRDDYEQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPR 272
+EQLYPRR+ ++RRP Q R+REE DM +N ++D+L+ A++V + PR
Sbjct: 11 REQLYPRREALDRRRP*QPMGSRRREE-DMELNAHLTDILQDVKAAHMVGKSGQS*PPPR 187
Query: 273 DA-NPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAG----------YQGNRQGQWRNGGD 321
+ K CE+ RS HD DDC+TL+REI+KLI+ G QG+RQ R GD
Sbjct: 188 RGIDTTK*CEYRRSVVHDIDDCFTLKREIEKLIKMGRLKQYDRGSRQQGDRQQGGRLQGD 367
Query: 322 QNKAHK 327
+ + ++
Sbjct: 368 RQQGNR 385
>BP033666
Length = 342
Score = 38.9 bits (89), Expect(2) = 9e-08
Identities = 19/40 (47%), Positives = 25/40 (62%)
Frame = -1
Query: 70 FSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARY 109
F + FL FSAN+ Q T DL+NI QQ GE ++ Y + Y
Sbjct: 273 FFN*FLTXFSANQTQKATSADLFNICQQVGERVRPYFSLY 154
Score = 33.9 bits (76), Expect(2) = 9e-08
Identities = 15/26 (57%), Positives = 17/26 (64%)
Frame = -2
Query: 47 TFKSTAMAWFTTLPRGSISNFRDFSS 72
+FK AM WF P SISNF DFS+
Sbjct: 341 SFKGVAMTWFIKQPPYSISNFTDFST 264
>BP065719
Length = 567
Score = 51.2 bits (121), Expect = 3e-07
Identities = 29/111 (26%), Positives = 55/111 (49%), Gaps = 2/111 (1%)
Frame = +3
Query: 1 IPDNMKTLVLDSYSGDSDPK--DHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTT 58
+P K +SGDS +H+ + + +A ++ +K + FPS+ A WFTT
Sbjct: 162 LPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLTKNAFTWFTT 341
Query: 59 LPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARY 109
L S+ + F QF + + V+ DL +++++ ES+ +Y+ R+
Sbjct: 342 LAPRSVHTWAQLERIFHEQFFRGECK-VSXKDLASVKRKPAESIDDYLNRF 491
>TC12971 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;}, partial (7%)
Length = 721
Score = 29.6 bits (65), Expect = 1.0
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = +3
Query: 319 GGDQNKAHKREEERADTKGKKKQE 342
GGD+ ++ REEE D K K K+E
Sbjct: 261 GGDEIPSNNREEEEGDNKSKNKKE 332
>AV769356
Length = 574
Score = 28.5 bits (62), Expect = 2.3
Identities = 19/59 (32%), Positives = 26/59 (43%), Gaps = 1/59 (1%)
Frame = -2
Query: 19 PKDHLLYFNTKMV-IIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLV 76
PK + F T ++ + A + P TFK + W T L S SN RD K L+
Sbjct: 546 PKGDVYSFGTVLLEFVTGERATQVAKAPETFKGNLVEWITQLT--SHSNLRDAIDKSLL 376
>TC17553 weakly similar to UP|AAP20176 (AAP20176) Kinase substrate HASPP28
(Fragment), partial (61%)
Length = 754
Score = 28.5 bits (62), Expect = 2.3
Identities = 32/164 (19%), Positives = 65/164 (39%), Gaps = 14/164 (8%)
Frame = +3
Query: 203 GKGKSVFKPTKEQLYPRRDD-----------YEQRRPWQSKSHRQREETDMVMNTDVSDM 251
G+GK KPT + + +D ++Q+ K + E T + + +
Sbjct: 87 GRGKFKAKPTGRRQFSTPEDMLAGTSTRPRTFKQKEVEDEKEVSEEEATSGDESEEETTK 266
Query: 252 LRGASDANLVDEPEAPK---YQPRDANPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAGY 308
+G ++ P K + RD + +K E R + + QR ++ +R
Sbjct: 267 TKGTQGVIEIENPNLVKPKSLKARDVDLEKTTELSRREREEIEK----QRAHERYMRLQE 434
Query: 309 QGNRQGQWRNGGDQNKAHKREEERADTKGKKKQESAAIATKGAD 352
QG + ++ D + ++RA+ K+ +E AA K ++
Sbjct: 435 QGKTE---QSKKDLERLALIRQQRAEAAKKRDEEKAAKEQKKSE 557
>TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) precursor,
partial (51%)
Length = 1566
Score = 28.5 bits (62), Expect = 2.3
Identities = 18/81 (22%), Positives = 36/81 (44%), Gaps = 2/81 (2%)
Frame = +1
Query: 162 DEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGD--GKQQRPGKGKSVFKPTKEQLYPR 219
DE+ E R + P R + + ED+ + G ++ PG+ + F+ +E+
Sbjct: 532 DEDQEQHGRHSRPSGRRTRRPSREEDEDEDEDEEERQGGRRGPGERQRHFEREEEEEREE 711
Query: 220 RDDYEQRRPWQSKSHRQREET 240
R + + WQ + + EE+
Sbjct: 712 RGPRGRGKHWQREEEEEEEES 774
>BI419411
Length = 408
Score = 28.5 bits (62), Expect = 2.3
Identities = 19/65 (29%), Positives = 32/65 (49%)
Frame = -3
Query: 140 KLTRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKRVKKDRSGEDKGDGKQ 199
K R A ++ + RAR +L + +R+R + G T+P RV R G+D +
Sbjct: 277 KAVRLEALAVDDRRARLVVLLLGDPHGLERRQRR--QNGATNPHRVLALRRGDDLDLHRS 104
Query: 200 QRPGK 204
+R G+
Sbjct: 103 RRQGR 89
>BP059356
Length = 436
Score = 28.1 bits (61), Expect = 3.0
Identities = 28/92 (30%), Positives = 40/92 (43%), Gaps = 9/92 (9%)
Frame = +1
Query: 321 DQNKAHKREEERADTKGKKKQESAAIAT----KGADDTFARHS----GPPVGTINTIAGG 372
D+N +K+E A+ KK E+ + + + D F PP G AGG
Sbjct: 184 DKNPNNKKE---AEANFKKISEAYDVLSDPQKRAVYDQFGEEGLNGQVPPPG-----AGG 339
Query: 373 FGGGGDTHAARKR-HVRAVNSVHEVAFGFVHP 403
F GGGD A R + R+ + + FGF P
Sbjct: 340 FSGGGDGGPASFRFNPRSADDILSEFFGFSSP 435
>TC7927 similar to UP|Q93YR3 (Q93YR3) HSP associated protein like, partial
(56%)
Length = 1116
Score = 27.7 bits (60), Expect = 3.9
Identities = 16/59 (27%), Positives = 28/59 (47%), Gaps = 5/59 (8%)
Frame = +1
Query: 324 KAHKREEERADTKGKKKQESAAIAT-----KGADDTFARHSGPPVGTINTIAGGFGGGG 377
+ HK EE+ + ++++ + A A K + +R+ G G + GGF GGG
Sbjct: 298 RLHKEREEKRTERERQRRRAEAQAAYEKAKKQEQSSSSRNPGGMPGEFPGMPGGFPGGG 474
>TC11979 similar to UP|Q9LVH4 (Q9LVH4) GTPase activating protein-like,
partial (94%)
Length = 711
Score = 27.7 bits (60), Expect = 3.9
Identities = 16/56 (28%), Positives = 24/56 (42%)
Frame = +3
Query: 224 EQRRPWQSKSHRQREETDMVMNTDVSDMLRGASDANLVDEPEAPKYQPRDANPKKW 279
E R W S+ HR +E T T ++L G L P++ P+ A + W
Sbjct: 498 EDERNWPSQRHRDQENT-----TKQGELLGGRELYLL*QRQNYPRHDPQIATCRMW 650
>TC15229 weakly similar to UP|O60585 (O60585) Ser/Arg-related nuclear matrix
protein, partial (4%)
Length = 965
Score = 26.9 bits (58), Expect = 6.7
Identities = 29/97 (29%), Positives = 40/97 (40%), Gaps = 3/97 (3%)
Frame = +2
Query: 135 GGLNSKLTRKPARSMGEMRARASTYILDEEDEAFKRKRAKLEKGDTSPKR-VKKDRSGED 193
GG + K PAR+ G R ++S + E + + G P R + K R GE
Sbjct: 179 GGDSPKPDLVPARAQG--RRKSSLRAVPGEAVTAESGCRRSAGGANRPSRGL*KVRCGEG 352
Query: 194 KGDGKQQRPGKGKS--VFKPTKEQLYPRRDDYEQRRP 228
GDG PG G V +P + D+ E RP
Sbjct: 353 FGDGGAPLPGSGGLC*VRRPRADGADAGEDNIENIRP 463
>BP057647
Length = 567
Score = 26.6 bits (57), Expect = 8.8
Identities = 18/61 (29%), Positives = 28/61 (45%)
Frame = -3
Query: 270 QPRDANPKKWCEFHRSAGHDTDDCWTLQREIDKLIRAGYQGNRQGQWRNGGDQNKAHKRE 329
+P +++P E A + WT++REI +R G+ W GGD + A E
Sbjct: 250 RPGESSPATSREKKSGAAESGEAQWTVEREIS--LRKTESGS---WWSEGGDLSTAAVEE 86
Query: 330 E 330
E
Sbjct: 85 E 83
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,156,268
Number of Sequences: 28460
Number of extensions: 72262
Number of successful extensions: 545
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 541
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 544
length of query: 441
length of database: 4,897,600
effective HSP length: 93
effective length of query: 348
effective length of database: 2,250,820
effective search space: 783285360
effective search space used: 783285360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0590c.5


