

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590c.10
(993 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
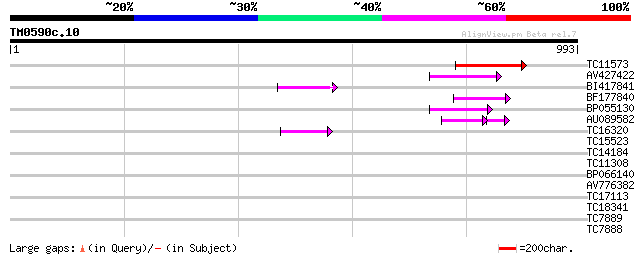
Score E
Sequences producing significant alignments: (bits) Value
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 172 2e-43
AV427422 70 2e-12
BI417841 69 4e-12
BF177840 66 2e-11
BP055130 62 6e-10
AU089582 52 1e-08
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 51 1e-06
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 36 0.026
TC14184 similar to UP|Q93WZ6 (Q93WZ6) Abscisic stress ripening-l... 33 0.29
TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoa... 31 1.1
BP066140 31 1.1
AV776382 30 1.9
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 30 2.5
TC18341 similar to UP|Q25936 (Q25936) Major merozoite surface an... 28 5.5
TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 28 7.2
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 28 9.3
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 172 bits (436), Expect = 2e-43
Identities = 80/124 (64%), Positives = 100/124 (80%)
Frame = +2
Query: 781 DNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLD 840
+NG QF+S QT++FC MGIQMRF+SV+HPQTNGQ E+AN+VIL+G++RRL EA+G W+D
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 841 ELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEIDNFTWRTRPGFEEENQANMAVELD 900
ELP VLWSYNT QS+ +ETPF +TYG D MLPVEI NF+W T+ EEEN+ NMAV +
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEISNFSW*TKARHEEENRMNMAVGWE 367
Query: 901 LLSE 904
L E
Sbjct: 368 KLPE 379
>AV427422
Length = 417
Score = 70.1 bits (170), Expect = 2e-12
Identities = 39/127 (30%), Positives = 70/127 (54%), Gaps = 1/127 (0%)
Frame = +1
Query: 736 ILVAVDYFTKWIEAEPLAKITSAKIV-NFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREF 794
+LV VD +K+ PL +AK++ + + + +V G+P +IVSD F S+ +E
Sbjct: 37 VLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMSNFWKEL 216
Query: 795 CKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAVLWSYNTTEQ 854
K G +++ ++ HP+++GQ E NR + LR +A+ +W +P + YNT+
Sbjct: 217 FKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYWYNTSYH 396
Query: 855 STTRETP 861
+T +TP
Sbjct: 397 VSTGQTP 417
>BI417841
Length = 617
Score = 68.9 bits (167), Expect = 4e-12
Identities = 38/105 (36%), Positives = 57/105 (54%)
Frame = +1
Query: 469 TNNQAEYEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLM 528
TNNQAEY LI GLK A E + + ++ DSQLV QV+G+++ ++PN+ + L
Sbjct: 253 TNNQAEYRGLILGLKHAHEQGYQHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELK 432
Query: 529 TLFQEVVVEYVPRAENQRADALAKLASTRKPGNNKSVIQETLAYP 573
+ FQ + +VPR N AD A L G+ ++E +P
Sbjct: 433 SKFQSFDINHVPRQYNSEADVQANLGVNLPAGH----VEEYCEFP 555
>BF177840
Length = 410
Score = 66.2 bits (160), Expect = 2e-11
Identities = 33/100 (33%), Positives = 56/100 (56%)
Frame = +2
Query: 777 AIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKG 836
+IVSD T+F S R ++G ++ +++ HPQT+GQ E N+ + LR L
Sbjct: 14 SIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLK 193
Query: 837 AWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPVEI 876
W LP + ++YN STT+ +PF + YG + + P+++
Sbjct: 194 MWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>BP055130
Length = 567
Score = 61.6 bits (148), Expect = 6e-10
Identities = 39/111 (35%), Positives = 59/111 (53%), Gaps = 1/111 (0%)
Frame = +2
Query: 736 ILVAVDYFTKWIEAEP-LAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREF 794
I+V VD +K+ P A TS+++ + IV G+PRAIVSD F+S+ + F
Sbjct: 191 IIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAFWKHF 370
Query: 795 CKEMGIQMRFASVEHPQTNGQVESANRVILRGLRRRLAEAKGAWLDELPAV 845
K G + +S HPQT+GQ E+ N+ + LR + E W+ L A+
Sbjct: 371 FKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLWVSYLLAL 523
>AU089582
Length = 383
Score = 52.0 bits (123), Expect(2) = 1e-08
Identities = 25/80 (31%), Positives = 45/80 (56%)
Frame = +2
Query: 757 SAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSSSQTREFCKEMGIQMRFASVEHPQTNGQV 816
+++ Y IV G+P +I+SD G QF+S R F +G +++ ++ HPQT+GQ
Sbjct: 11 ASQYAKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQS 190
Query: 817 ESANRVILRGLRRRLAEAKG 836
E +++ LR + + +G
Sbjct: 191 ERTIQILEDMLRACVXDLRG 250
Score = 24.6 bits (52), Expect(2) = 1e-08
Identities = 11/40 (27%), Positives = 20/40 (49%)
Frame = +3
Query: 835 KGAWLDELPAVLWSYNTTEQSTTRETPFRMTYGVDAMLPV 874
+G+W L + ++YN + +S+ PF YG P+
Sbjct: 246 EGSWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSPI 365
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 50.8 bits (120), Expect = 1e-06
Identities = 27/91 (29%), Positives = 45/91 (48%)
Frame = +2
Query: 475 YEALIAGLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEV 534
Y LI GLK A + + + ++ DS LV NQV+G +++K+ N+ + L F
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 535 VVEYVPRAENQRADALAKLASTRKPGNNKSV 565
+ ++PR N AD A + + G + V
Sbjct: 182 KINHIPREYNSEADVQANFGISLRAGQVEEV 274
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 36.2 bits (82), Expect = 0.026
Identities = 18/37 (48%), Positives = 24/37 (64%)
Frame = -2
Query: 52 RLLLFHHHHLLHRLHR*DLWSAHQRIHLLRNNSRR*H 88
R+LL +HH LHR HR + + H RIH L ++ RR H
Sbjct: 728 RILLLRYHHSLHRHHRYNGYHHHHRIH-LHDDHRRIH 621
>TC14184 similar to UP|Q93WZ6 (Q93WZ6) Abscisic stress ripening-like
protein, partial (57%)
Length = 1141
Score = 32.7 bits (73), Expect = 0.29
Identities = 15/42 (35%), Positives = 21/42 (49%), Gaps = 6/42 (14%)
Frame = -2
Query: 54 LLFHHH------HLLHRLHR*DLWSAHQRIHLLRNNSRR*HR 89
+L HHH HL H+ H LWS H + H N ++ H+
Sbjct: 477 ILHHHHR*NIRHHLHHQCHSLHLWSIHHQRHQCHNRHQKAHQ 352
>TC11308 similar to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (4%)
Length = 756
Score = 30.8 bits (68), Expect = 1.1
Identities = 14/42 (33%), Positives = 27/42 (63%)
Frame = -2
Query: 747 IEAEPLAKITSAKIVNFYWKRIVCRFGIPRAIVSDNGTQFSS 788
++ L ++++AK V F R+V RF +PR ++ D+G + +S
Sbjct: 509 VQGTSLLQVSTAKTVEFSDDRVV-RFPLPRDVIGDDGAETTS 387
>BP066140
Length = 320
Score = 30.8 bits (68), Expect = 1.1
Identities = 26/73 (35%), Positives = 34/73 (45%)
Frame = -2
Query: 481 GLKLAREVKIRSLLIRTDSQLVENQVKGTFQVKDPNLIKYLERVRYLMTLFQEVVVEYVP 540
GL LA E IR LL +D + V + G + YL R L+ E ++
Sbjct: 316 GLSLAWERGIRKLLCISDCEDVVKALTGLGSFHLDSWAAYLAEARSLIHPEWECLLLPHH 137
Query: 541 RAENQRADALAKL 553
R N+ ADALAKL
Sbjct: 136 RERNKPADALAKL 98
>AV776382
Length = 462
Score = 30.0 bits (66), Expect = 1.9
Identities = 17/43 (39%), Positives = 22/43 (50%)
Frame = -1
Query: 36 LIVWTWILRYERQEYSRLLLFHHHHLLHRLHR*DLWSAHQRIH 78
LI W+LR+ +Q S L + H HL LW A+QR H
Sbjct: 420 LIYLMWLLRF*KQNPSLLPFWFHQHLSSL----SLWVAYQRKH 304
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 29.6 bits (65), Expect = 2.5
Identities = 14/25 (56%), Positives = 15/25 (60%)
Frame = +2
Query: 55 LFHHHHLLHRLHR*DLWSAHQRIHL 79
L HHHHLLH H L S H +HL
Sbjct: 743 LHHHHHLLH--HHFSLLSLHCSLHL 811
>TC18341 similar to UP|Q25936 (Q25936) Major merozoite surface antigen
(Fragment), partial (5%)
Length = 497
Score = 28.5 bits (62), Expect = 5.5
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = +1
Query: 342 ASSRGRGRGSRYSNRPRNSCIH 363
++ RG G+G R+SN P NS I+
Sbjct: 184 SNRRGSGKGERWSNEPTNSTIN 249
>TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (66%)
Length = 1158
Score = 28.1 bits (61), Expect = 7.2
Identities = 15/34 (44%), Positives = 17/34 (49%), Gaps = 4/34 (11%)
Frame = +2
Query: 48 QEYSRLLLFHH----HHLLHRLHR*DLWSAHQRI 77
Q Y LL HH H LLH LH+ W HQ +
Sbjct: 368 QLYHHHLLLHHLHQPHRLLHHLHQHLPWLHHQAL 469
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 27.7 bits (60), Expect = 9.3
Identities = 15/32 (46%), Positives = 16/32 (49%), Gaps = 4/32 (12%)
Frame = +1
Query: 48 QEYSRLLLFHH----HHLLHRLHR*DLWSAHQ 75
Q Y LL HH H LLH LH+ W HQ
Sbjct: 814 QLYHHHLLLHHLHQPHRLLHHLHQHLPWLHHQ 909
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.336 0.143 0.468
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,043,398
Number of Sequences: 28460
Number of extensions: 266248
Number of successful extensions: 2715
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2478
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2642
length of query: 993
length of database: 4,897,600
effective HSP length: 99
effective length of query: 894
effective length of database: 2,080,060
effective search space: 1859573640
effective search space used: 1859573640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0590c.10


