

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590b.4
(283 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
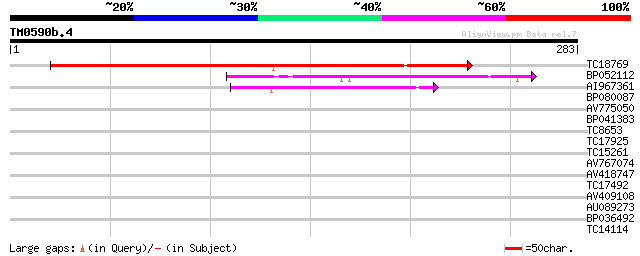
Score E
Sequences producing significant alignments: (bits) Value
TC18769 similar to GB|CAB81029.1|7269936|ATCHRIV72 cyclic nucleo... 207 2e-54
BP052112 75 1e-14
AI967361 74 3e-14
BP080087 39 0.001
AV775050 39 0.001
BP041383 36 0.008
TC8653 similar to UP|AAS46232 (AAS46232) Methionine sulfoxide re... 31 0.27
TC17925 30 0.46
TC15261 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, ... 30 0.61
AV767074 27 3.0
AV418747 27 3.9
TC17492 homologue to UP|R25A_ARATH (Q9SIK2) 40S ribosomal protei... 27 3.9
AV409108 27 5.1
AU089273 26 6.7
BP036492 26 8.8
TC14114 similar to UP|FD62_SOYBN (P48631) Omega-6 fatty acid des... 26 8.8
>TC18769 similar to GB|CAB81029.1|7269936|ATCHRIV72 cyclic nucleotide and
calmodulin-regulated ion channel-like protein
{Arabidopsis thaliana;} , partial (29%)
Length = 646
Score = 207 bits (527), Expect = 2e-54
Identities = 109/213 (51%), Positives = 138/213 (64%), Gaps = 2/213 (0%)
Frame = +2
Query: 21 EWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLPQGLRRDIKYHLCLDLVRQV 80
E WMR RQLP+ LR RVR + + +W A RGVDE ++R LP LRRDI+ HLCLDLVR+V
Sbjct: 11 EEWMRHRQLPEDLRFRVRRFVQYKWLATRGVDEETILRALPADLRRDIQRHLCLDLVRRV 190
Query: 81 PLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLFVVRGHLQSS--QDLRDGVK 138
P F MDD +L+ IC+R+ S + T+G I REGDPV MLF++RG L SS R G
Sbjct: 191 PFFSQMDDQLLDAICERLISSLSTQGTYIVREGDPVTEMLFIIRGRLDSSTTNGGRSGFF 370
Query: 139 SYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQHFRY 198
+ L PG+F G+ELLSW L LP S+ T+ L EAF L AED+++V FR
Sbjct: 371 NSIILRPGDFCGEELLSWALLPKSTVNLPSSTRTVKALSEVEAFALRAEDLKFVANQFR- 547
Query: 199 TFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRY 231
+K++ + R+YS WRTWAA IQ AWRRY
Sbjct: 548 RLRSKKLQHTFRFYSHHWRTWAACFIQAAWRRY 646
>BP052112
Length = 553
Score = 75.1 bits (183), Expect = 1e-14
Identities = 59/167 (35%), Positives = 86/167 (51%), Gaps = 12/167 (7%)
Frame = -2
Query: 109 ITREGDPVKRMLFVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIE---- 164
I +G +++M+FVVRG L+S + DG + L G+ G+EL++W L +
Sbjct: 552 ILSQGSLIEKMVFVVRGKLESIGE--DGTRM--PLSEGDACGEELMTWYLEHSSVSSDGR 385
Query: 165 --RLPP----SSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRT 218
RLP S+ T+ L EAF L A D+ VT F +V+ + RY SP WR+
Sbjct: 384 KVRLPGQRLVSNRTVKCLTNVEAFSLSAADLEEVTILFTRFLRSPQVQGALRYESPYWRS 205
Query: 219 WAAVAIQLAWRRYKHRLSLTSLSFIRPRRPASRC--TSMEEDRLRLY 263
AA IQ+AWR + RLS + S + + S C TS DR+ L+
Sbjct: 204 LAANRIQVAWRYRQKRLSRVN-SSVTTKH*GSSCNVTSDFFDRIMLH 67
>AI967361
Length = 358
Score = 73.9 bits (180), Expect = 3e-14
Identities = 43/106 (40%), Positives = 63/106 (58%), Gaps = 2/106 (1%)
Frame = +2
Query: 111 REGDPVKRMLFVVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPP 168
REGDPV MLF++RG L + + R G + L G+F G+ELL+W L LP
Sbjct: 11 REGDPVDEMLFIMRGKLLTVTTNGGRTGFFNSEFLKAGDFCGEELLTWALDPHSSSNLPI 190
Query: 169 SSATLVTLETTEAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSP 214
S+ T+ TL EAF L+A+D+++V FR K+ ++ + R+YSP
Sbjct: 191 STRTVQTLAEVEAFALKADDLKFVASQFRRLHSKQ-LRHTFRFYSP 325
>BP080087
Length = 382
Score = 38.9 bits (89), Expect = 0.001
Identities = 17/33 (51%), Positives = 23/33 (69%)
Frame = -1
Query: 35 QRVRNYERQRWTAMRGVDECQMIRNLPQGLRRD 67
+RVR ER W A RGV+E ++ NLP+ L+RD
Sbjct: 331 RRVRQAERYNWAATRGVNEEMVMENLPEDLQRD 233
>AV775050
Length = 382
Score = 38.5 bits (88), Expect = 0.001
Identities = 22/52 (42%), Positives = 31/52 (59%)
Frame = -3
Query: 190 RYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSLS 241
RY +Q F + +V+ + RY SP WR+ AA IQ+AWR + RLS + S
Sbjct: 353 RYFSQDF---LRRPQVQGALRYESPYWRSLAANRIQVAWRYRQTRLSRVNSS 207
>BP041383
Length = 476
Score = 35.8 bits (81), Expect = 0.008
Identities = 14/19 (73%), Positives = 14/19 (73%)
Frame = -2
Query: 216 WRTWAAVAIQLAWRRYKHR 234
WRTWAA IQ AWRRY R
Sbjct: 475 WRTWAACFIQAAWRRYSKR 419
>TC8653 similar to UP|AAS46232 (AAS46232) Methionine sulfoxide reductase A,
partial (71%)
Length = 1175
Score = 30.8 bits (68), Expect = 0.27
Identities = 12/42 (28%), Positives = 24/42 (56%)
Frame = -1
Query: 4 SATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRW 45
+AT +K + + +M +EWWMRK ++ R +++ +W
Sbjct: 221 AATLDRKSSGRGKMGKIEWWMRKNKMKTRARDWDWDWDWAKW 96
>TC17925
Length = 546
Score = 30.0 bits (66), Expect = 0.46
Identities = 17/50 (34%), Positives = 26/50 (52%)
Frame = +2
Query: 225 QLAWRRYKHRLSLTSLSFIRPRRPASRCTSMEEDRLRLYTALLTSPKPND 274
Q+ WR + H L+LTSL+ RP+ P+S C R + +P+D
Sbjct: 137 QVQWRLHTH-LNLTSLNHQRPQVPSSTCHQCPPTRPETHLQRKGFGRPHD 283
>TC15261 weakly similar to UP|Q9ZQ55 (Q9ZQ55) At2g22570 protein, partial
(75%)
Length = 1031
Score = 29.6 bits (65), Expect = 0.61
Identities = 14/44 (31%), Positives = 24/44 (53%)
Frame = +1
Query: 109 ITREGDPVKRMLFVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDE 152
+ +E PVK+ + G +++ L D V +CT+G GN + E
Sbjct: 55 LVKEEIPVKQQPLRLSGDIKTGLVLVDVVNGFCTVGAGNMAPTE 186
>AV767074
Length = 481
Score = 27.3 bits (59), Expect = 3.0
Identities = 14/51 (27%), Positives = 26/51 (50%)
Frame = -1
Query: 197 RYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSLSFIRPRR 247
+++ M ++K + +Y + A+ + WR+ KH+L LS R RR
Sbjct: 268 KWSEMWRQLKNISHHYEFQYPPKKEDAVNMLWRQVKHKLPGVRLSVHRSRR 116
>AV418747
Length = 386
Score = 26.9 bits (58), Expect = 3.9
Identities = 11/26 (42%), Positives = 12/26 (45%)
Frame = -2
Query: 21 EWWMRKRQLPQGLRQRVRNYERQRWT 46
EWW R G R V YE RW+
Sbjct: 82 EWWWRSHARAYGRR*GVDKYEAARWS 5
>TC17492 homologue to UP|R25A_ARATH (Q9SIK2) 40S ribosomal protein S25-1,
complete
Length = 559
Score = 26.9 bits (58), Expect = 3.9
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +3
Query: 148 FSGDELLSWCLRRPFIERLPPSSATLVTLET 178
FS +L+SW RR + LPPS L T
Sbjct: 24 FSFPQLISWLQRRIRLPHLPPSPPNLAMAST 116
>AV409108
Length = 359
Score = 26.6 bits (57), Expect = 5.1
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = +1
Query: 228 WRRYKHRLSLTSLSFIRP 245
WRRY H S+TSL F P
Sbjct: 40 WRRYLHHHSITSLPFHLP 93
>AU089273
Length = 420
Score = 26.2 bits (56), Expect = 6.7
Identities = 11/21 (52%), Positives = 18/21 (85%)
Frame = -3
Query: 87 DDLVLENICDRVKSLVFTKGE 107
D++VLE++C+ +K L FTKG+
Sbjct: 139 DNMVLESVCNEMK-LYFTKGK 80
>BP036492
Length = 552
Score = 25.8 bits (55), Expect = 8.8
Identities = 12/41 (29%), Positives = 17/41 (41%)
Frame = -3
Query: 155 SWCLRRPFIERLPPSSATLVTLETTEAFGLEAEDVRYVTQH 195
SWC I +P S+ATL GL+ + + H
Sbjct: 280 SWCFAVNIISEIPASAATLAHWAVLRPVGLKIAGLSWPVPH 158
>TC14114 similar to UP|FD62_SOYBN (P48631) Omega-6 fatty acid desaturase,
endoplasmic reticulum isozyme 2 , complete
Length = 1634
Score = 25.8 bits (55), Expect = 8.8
Identities = 14/43 (32%), Positives = 22/43 (50%)
Frame = -1
Query: 29 LPQGLRQRVRNYERQRWTAMRGVDECQMIRNLPQGLRRDIKYH 71
LP+ L+ + + R R + +RNLP GL R ++YH
Sbjct: 680 LPETLKVKYKGQPRVRVIVI--------VRNLPSGLLRYLEYH 576
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,153,914
Number of Sequences: 28460
Number of extensions: 72193
Number of successful extensions: 468
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 462
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 464
length of query: 283
length of database: 4,897,600
effective HSP length: 89
effective length of query: 194
effective length of database: 2,364,660
effective search space: 458744040
effective search space used: 458744040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0590b.4


