

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0587.5
(191 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
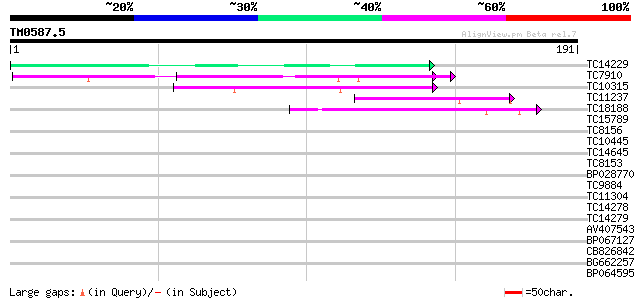
Score E
Sequences producing significant alignments: (bits) Value
TC14229 similar to UP|P93166 (P93166) SCOF-1, partial (38%) 54 2e-08
TC7910 homologue to UP|P93166 (P93166) SCOF-1, partial (52%) 54 2e-08
TC10315 similar to UP|P93166 (P93166) SCOF-1, partial (42%) 52 9e-08
TC11237 similar to UP|Q43614 (Q43614) DNA-binding protein, parti... 42 5e-05
TC18188 similar to UP|ZFP6_ARATH (Q39265) Zinc finger protein 6,... 41 1e-04
TC15789 similar to UP|O22086 (O22086) ZPT2-14, partial (37%) 37 0.002
TC8156 UP|AAR99188 (AAR99188) ATP synthase F0 subunit 8, partial... 35 0.011
TC10445 34 0.018
TC14645 similar to UP|Q41122 (Q41122) Proline-rich protein precu... 33 0.031
TC8153 weakly similar to UP|Q8VAJ3 (Q8VAJ3) Wsv416 (WSSV475), pa... 33 0.041
BP028770 33 0.041
TC9884 weakly similar to GB|AAO63337.1|28950827|BT005273 At5g585... 32 0.054
TC11304 similar to PIR|T04669|T04669 serine O-acetyltransferase ... 32 0.070
TC14278 homologue to UP|O65760 (O65760) Extensin (Fragment), par... 32 0.091
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 32 0.091
AV407543 31 0.12
BP067127 31 0.12
CB826842 31 0.12
BG662257 31 0.12
BP064595 31 0.16
>TC14229 similar to UP|P93166 (P93166) SCOF-1, partial (38%)
Length = 1175
Score = 53.9 bits (128), Expect = 2e-08
Identities = 41/143 (28%), Positives = 55/143 (37%)
Frame = +1
Query: 1 MRNSHSHTQKHVVPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPEREWRGINPPPK 60
+R S + P P +CS C K F S +AL GH H
Sbjct: 304 LRAESSSNSQSQSPPQQQPPLKLTHKCSVCDKAFPSYQALGGHKASH------------- 444
Query: 61 FRRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCS 120
R S + Q P + N NA T + + ++ H+CS
Sbjct: 445 --RKSSSENQTPAAAAV---------------NDNASVSTSTGNGGKM--------HECS 549
Query: 121 ICLRVFSSGQALGGHKRRHWEKG 143
IC + F +GQALGGHKR H+E G
Sbjct: 550 ICHKSFPTGQALGGHKRCHYEGG 618
>TC7910 homologue to UP|P93166 (P93166) SCOF-1, partial (52%)
Length = 1035
Score = 53.9 bits (128), Expect = 2e-08
Identities = 45/145 (31%), Positives = 61/145 (42%), Gaps = 2/145 (1%)
Frame = +3
Query: 2 RNSHSHTQKHVVPSPSSSPSGKWS-QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPK 60
R S + T K + P+ + K S +CS C K F S +AL GH H +
Sbjct: 291 RGSTAVTPKLTLSRPAPVTAEKLSYKCSVCEKTFPSYQALGGHKASHRK----------- 437
Query: 61 FRRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHH-HKC 119
LA A A+ + SS+ + S S+ H+C
Sbjct: 438 ------------------------------LAGAAAEDHSTSSAVTTSSASNGGGKVHQC 527
Query: 120 SICLRVFSSGQALGGHKRRHWEKGG 144
SIC + F +GQALGGHKR H+E GG
Sbjct: 528 SICQKSFPTGQALGGHKRCHYEGGG 602
Score = 46.6 bits (109), Expect = 3e-06
Identities = 30/98 (30%), Positives = 45/98 (45%), Gaps = 4/98 (4%)
Frame = +3
Query: 57 PPPKFRRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEIS----CSS 112
P K +R + + EE +A CL++LA +G T + K +S ++
Sbjct: 183 PWAKRKRSKRSRTDSHHNHASCTEEEYLALCLIMLA----RGSTAVTPKLTLSRPAPVTA 350
Query: 113 SHHHHKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHH 150
+KCS+C + F S QALGGHK H + G H
Sbjct: 351 EKLSYKCSVCEKTFPSYQALGGHKASHRKLAGAAAEDH 464
>TC10315 similar to UP|P93166 (P93166) SCOF-1, partial (42%)
Length = 592
Score = 51.6 bits (122), Expect = 9e-08
Identities = 31/93 (33%), Positives = 46/93 (49%), Gaps = 4/93 (4%)
Frame = +2
Query: 56 NPPPKFRRPSQPQPQPPQT--TVITQEEHEVAACLLLLANANAKGDTCSSSKSEISC--S 111
N P R P++P + ++ + EE +A CL++LA A T + +
Sbjct: 194 NDHPTLRHPAEPWAKRKRSKRSRSCSEEEYLALCLIMLARGGAATTTATPPLQPATAPSG 373
Query: 112 SSHHHHKCSICLRVFSSGQALGGHKRRHWEKGG 144
SS +KCS+C + F S QALGGHK H + G
Sbjct: 374 SSRLSYKCSVCDKAFPSYQALGGHKASHRKHSG 472
Score = 33.9 bits (76), Expect = 0.018
Identities = 29/114 (25%), Positives = 42/114 (36%), Gaps = 3/114 (2%)
Frame = +2
Query: 16 PSSSPSGKWS---QCSECGKKFWSEKALHGHMRCHPEREWRGINPPPKFRRPSQPQPQPP 72
P+++PSG +CS C K F S +AL GH H + G
Sbjct: 353 PATAPSGSSRLSYKCSVCDKAFPSYQALGGHKASHRKHSGGG------------------ 478
Query: 73 QTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVF 126
E+H A +A + +T S K+ H+CSIC + F
Sbjct: 479 ------GEDHSAGATSSAVATTTSSANTGSGGKA----------HECSICHKSF 592
>TC11237 similar to UP|Q43614 (Q43614) DNA-binding protein, partial (20%)
Length = 523
Score = 42.4 bits (98), Expect = 5e-05
Identities = 27/63 (42%), Positives = 31/63 (48%), Gaps = 9/63 (14%)
Frame = +3
Query: 117 HKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHH--------LPPPAPAPAPVLDLNLP- 167
H+CSIC F+SGQALGGH RRH + + P LDLNLP
Sbjct: 54 HECSICGSEFTSGQALGGHMRRHRASANAAAETNSNAITSTTVVARPPRNILQLDLNLPA 233
Query: 168 PPE 170
PPE
Sbjct: 234 PPE 242
Score = 32.0 bits (71), Expect = 0.070
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = +3
Query: 23 KWSQCSECGKKFWSEKALHGHMRCH 47
K +CS CG +F S +AL GHMR H
Sbjct: 48 KIHECSICGSEFTSGQALGGHMRRH 122
>TC18188 similar to UP|ZFP6_ARATH (Q39265) Zinc finger protein 6, partial
(21%)
Length = 594
Score = 41.2 bits (95), Expect = 1e-04
Identities = 29/92 (31%), Positives = 40/92 (42%), Gaps = 7/92 (7%)
Frame = +3
Query: 95 NAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLPPP 154
+ GD S+ S I+ S KC C R F++ QALGGH+ H ++ L L
Sbjct: 246 SGNGDAAGSA-SGINTPSGDRKKKCQYCCREFANSQALGGHQNAHKKERQQLKRAQLQAT 422
Query: 155 APAPA-----PVLDLNLPPPE--SGSDPITVP 179
A P++ PPP S S + VP
Sbjct: 423 RKAAVSFVRNPIISAFAPPPHLLSPSGAMMVP 518
>TC15789 similar to UP|O22086 (O22086) ZPT2-14, partial (37%)
Length = 747
Score = 37.0 bits (84), Expect = 0.002
Identities = 16/28 (57%), Positives = 18/28 (64%)
Frame = +1
Query: 117 HKCSICLRVFSSGQALGGHKRRHWEKGG 144
H CS+C FS GQALGGH R+H G
Sbjct: 382 HVCSVCGLGFSLGQALGGHMRKHRNNEG 465
Score = 33.1 bits (74), Expect = 0.031
Identities = 20/63 (31%), Positives = 29/63 (45%)
Frame = +1
Query: 80 EEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRH 139
E ++ CL+L+ + + T + S +C C R FSS QALGGH+ H
Sbjct: 160 ESFDLTNCLMLMLSYPKQKRTINESVE----------FECKTCNRKFSSFQALGGHRASH 309
Query: 140 WEK 142
K
Sbjct: 310 NHK 318
Score = 30.0 bits (66), Expect = 0.27
Identities = 19/55 (34%), Positives = 25/55 (44%), Gaps = 5/55 (9%)
Frame = +1
Query: 2 RNSHSHTQKHV-----VPSPSSSPSGKWSQCSECGKKFWSEKALHGHMRCHPERE 51
R SH+H + + PS + + CS CG F +AL GHMR H E
Sbjct: 298 RASHNHKRVKLEEQAKTPSLWDNNKPRMHVCSVCGLGFSLGQALGGHMRKHRNNE 462
>TC8156 UP|AAR99188 (AAR99188) ATP synthase F0 subunit 8, partial (24%)
Length = 635
Score = 34.7 bits (78), Expect = 0.011
Identities = 22/74 (29%), Positives = 34/74 (45%), Gaps = 11/74 (14%)
Frame = +2
Query: 56 NPPPKFRRPSQPQPQPP-----------QTTVITQEEHEVAACLLLLANANAKGDTCSSS 104
+PPP F + + P P P +TT T HE A LL + N + + +++
Sbjct: 68 SPPPYFSQVAVPTPTPTPCSCFFFPINHETTTFTSFSHEQAVTTLLSMDLNHELLSLAAA 247
Query: 105 KSEISCSSSHHHHK 118
S + SSS HH+
Sbjct: 248 PSCLLLSSSRSHHR 289
>TC10445
Length = 448
Score = 33.9 bits (76), Expect = 0.018
Identities = 25/79 (31%), Positives = 33/79 (41%), Gaps = 1/79 (1%)
Frame = +3
Query: 101 CSSSKSEISCSSSHHHHKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLPPPAPAPAP 160
C S + S+ HHHH G L H+ RH G+ + LPPP P
Sbjct: 126 CESLLRDNKSSNRHHHH--------HRPGHRLKLHRNRHDVDDGEAL---LPPPVPVKDS 272
Query: 161 V-LDLNLPPPESGSDPITV 178
V ++LP SGS T+
Sbjct: 273 VDRQVSLPRLSSGSSLFTL 329
>TC14645 similar to UP|Q41122 (Q41122) Proline-rich protein precursor,
partial (52%)
Length = 1060
Score = 33.1 bits (74), Expect = 0.031
Identities = 13/21 (61%), Positives = 14/21 (65%)
Frame = +2
Query: 148 HHHLPPPAPAPAPVLDLNLPP 168
HHH P PAPAPA + LPP
Sbjct: 392 HHHPPSPAPAPALYTTITLPP 454
Score = 32.3 bits (72), Expect = 0.054
Identities = 15/30 (50%), Positives = 16/30 (53%)
Frame = +2
Query: 148 HHHLPPPAPAPAPVLDLNLPPPESGSDPIT 177
HHH PPAPAPAP+ PP P T
Sbjct: 290 HHHHHPPAPAPAPIKPPVHPPTSVPVKPPT 379
Score = 25.4 bits (54), Expect(2) = 0.56
Identities = 10/16 (62%), Positives = 11/16 (68%), Gaps = 1/16 (6%)
Frame = +1
Query: 147 IHHHLPP-PAPAPAPV 161
+HHH PP PAP PV
Sbjct: 430 VHHHYPPTPAPVHPPV 477
Score = 21.9 bits (45), Expect(2) = 0.56
Identities = 16/59 (27%), Positives = 21/59 (35%)
Frame = +1
Query: 59 PKFRRPSQPQPQPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEISCSSSHHHH 117
P RP+ P P PP + + L + + S SCSS HHH
Sbjct: 265 PTGSRPTPPPPPPPTGSGSCSY*ASSSPSHLCPS*TPHRPPPPPPSSFPCSCSSPVHHH 441
>TC8153 weakly similar to UP|Q8VAJ3 (Q8VAJ3) Wsv416 (WSSV475), partial
(18%)
Length = 476
Score = 32.7 bits (73), Expect = 0.041
Identities = 21/70 (30%), Positives = 33/70 (47%), Gaps = 7/70 (10%)
Frame = +1
Query: 56 NPPPKFRRPSQPQP-------QPPQTTVITQEEHEVAACLLLLANANAKGDTCSSSKSEI 108
+PPP F + + P P +TT T HE A LL + N + + +++ S +
Sbjct: 175 SPPPYFSQVAVPTPCSCFFFPMNHETTTFTSFSHEQAVTTLLSMDLNHELLSLAAAPSCL 354
Query: 109 SCSSSHHHHK 118
SSS HH+
Sbjct: 355 LLSSSRSHHR 384
>BP028770
Length = 451
Score = 32.7 bits (73), Expect = 0.041
Identities = 14/32 (43%), Positives = 17/32 (52%)
Frame = +3
Query: 148 HHHLPPPAPAPAPVLDLNLPPPESGSDPITVP 179
+HH P P+P+P PP SDP TVP
Sbjct: 168 NHHHPTHLPSPSPASSSTPVPPTEKSDPSTVP 263
>TC9884 weakly similar to GB|AAO63337.1|28950827|BT005273 At5g58530
{Arabidopsis thaliana;}, partial (31%)
Length = 557
Score = 32.3 bits (72), Expect = 0.054
Identities = 15/41 (36%), Positives = 18/41 (43%)
Frame = -1
Query: 135 HKRRHWEKGGDLIHHHLPPPAPAPAPVLDLNLPPPESGSDP 175
H H E+ GD H PP P A + PPP S + P
Sbjct: 173 HNTHHTEQTGDRRKRHTPPDPPPAAL*ESSSAPPPRSSASP 51
>TC11304 similar to PIR|T04669|T04669 serine O-acetyltransferase F8D20.150
- Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (24%)
Length = 775
Score = 32.0 bits (71), Expect = 0.070
Identities = 28/88 (31%), Positives = 35/88 (38%), Gaps = 14/88 (15%)
Frame = +1
Query: 103 SSKSEISCSSSHHH----HKCSICLRVFSSGQALGGHKRRHWEKGGDLIHHHLPPPAPA- 157
S+ ++C S HHH H+ S+ L SG A H RR + HH P A A
Sbjct: 313 STPPRMACVSHHHHHHYRHRNSLALPNMLSGAA---HHRRRLQNDAPEPDHHRPTLASAY 483
Query: 158 ------PAPVLDLNLPPPE---SGSDPI 176
P + P PE G DPI
Sbjct: 484 RLEKVFPVYAMGAADPEPEPDLDGPDPI 567
>TC14278 homologue to UP|O65760 (O65760) Extensin (Fragment), partial (28%)
Length = 619
Score = 31.6 bits (70), Expect = 0.091
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +2
Query: 152 PPPAPAPAPVLDLNLPPPESGSDP 175
PPP+P+PAP PPP + S P
Sbjct: 50 PPPSPSPAPTYIYKSPPPPTKSPP 121
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 31.6 bits (70), Expect = 0.091
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGSDP 175
PPP+P+PAP PPP + S P
Sbjct: 532 PPPSPSPAPTYIYKSPPPPTKSPP 603
Score = 30.0 bits (66), Expect = 0.27
Identities = 12/24 (50%), Positives = 14/24 (58%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGSDP 175
PPP+P+P P PPP S S P
Sbjct: 100 PPPSPSPPPPYYYQSPPPPSHSPP 171
Score = 30.0 bits (66), Expect = 0.27
Identities = 12/24 (50%), Positives = 14/24 (58%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGSDP 175
PPP+P+P P PPP S S P
Sbjct: 196 PPPSPSPPPPYYYKSPPPPSPSPP 267
Score = 28.9 bits (63), Expect = 0.59
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGSDPITVP 179
PPP+P+PAP PPP P+ +P
Sbjct: 436 PPPSPSPAPTYIYKSPPP-----PVKLP 504
Score = 28.5 bits (62), Expect = 0.77
Identities = 11/24 (45%), Positives = 13/24 (53%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGSDP 175
PPP+P P P PPP + S P
Sbjct: 292 PPPSPIPHPPYYYKSPPPPTSSPP 363
Score = 27.7 bits (60), Expect = 1.3
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGS 173
PPP+P+P P PPP S S
Sbjct: 388 PPPSPSPPPPYHYASPPPPSPS 453
Score = 26.9 bits (58), Expect = 2.3
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPES 171
PPP+P+P P PPP S
Sbjct: 244 PPPSPSPPPPYYYKSPPPPS 303
Score = 26.6 bits (57), Expect = 2.9
Identities = 11/24 (45%), Positives = 13/24 (53%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGSDP 175
PPP+ +P P PPP S S P
Sbjct: 148 PPPSHSPPPPYYYKSPPPPSPSPP 219
Score = 26.2 bits (56), Expect = 3.8
Identities = 11/24 (45%), Positives = 12/24 (49%)
Frame = +1
Query: 152 PPPAPAPAPVLDLNLPPPESGSDP 175
PPP +P P PPP S S P
Sbjct: 340 PPPTSSPPPPYHYVSPPPPSPSPP 411
>AV407543
Length = 227
Score = 31.2 bits (69), Expect = 0.12
Identities = 19/46 (41%), Positives = 20/46 (43%), Gaps = 6/46 (13%)
Frame = +2
Query: 152 PPPAPAPAPVLDLNLPP------PESGSDPITVPLDFAALDLSLRL 191
PPP PAP L N PP P + S P T LD A L L
Sbjct: 74 PPPQQVPAPPLTANSPPISVPPFPSTISSPSTTSLDAALRGFELPL 211
Score = 27.3 bits (59), Expect = 1.7
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = +2
Query: 152 PPPAPAPAPVLDLNLPPPESGSDPITVP 179
PP P P P + PP + S PI+VP
Sbjct: 53 PP*PPPPPPPQQVPAPPLTANSPPISVP 136
>BP067127
Length = 346
Score = 31.2 bits (69), Expect = 0.12
Identities = 14/42 (33%), Positives = 21/42 (49%), Gaps = 2/42 (4%)
Frame = -3
Query: 36 SEKALHGHMRCHPEREW--RGINPPPKFRRPSQPQPQPPQTT 75
S+ ++H H+RC R PPP+ R ++ P PP T
Sbjct: 302 SDPSIHPHLRCRRSSTTTARASPPPPEMRTNTKAHPLPPSAT 177
>CB826842
Length = 637
Score = 31.2 bits (69), Expect = 0.12
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = +1
Query: 117 HKCSICLRVFSSGQALGGHKRRH 139
+ C+ C R FS+ QALGGH H
Sbjct: 127 YSCTFCKRGFSNAQALGGHMNIH 195
>BG662257
Length = 336
Score = 31.2 bits (69), Expect = 0.12
Identities = 19/59 (32%), Positives = 25/59 (42%), Gaps = 3/59 (5%)
Frame = +3
Query: 94 ANAKGDTCSSSKSEISCSSSHHHHKCSICLRVFSSGQAL---GGHKRRHWEKGGDLIHH 149
A A G C S E ++ HKC+ C RV SG ++ H+ G IHH
Sbjct: 150 AEAAGGNCKSHSKESKNITTR*SHKCT*C-RV*ESGSGCTRPSEGRQNHYSSGSQAIHH 323
>BP064595
Length = 474
Score = 30.8 bits (68), Expect = 0.16
Identities = 17/44 (38%), Positives = 24/44 (53%)
Frame = +3
Query: 137 RRHWEKGGDLIHHHLPPPAPAPAPVLDLNLPPPESGSDPITVPL 180
+RHW L+ H P P +P+L L+ PP S S P+T+ L
Sbjct: 255 QRHWSS---LLQPH--PSLPHLSPLLHLSSPPSPSPSSPLTLSL 371
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,470,470
Number of Sequences: 28460
Number of extensions: 178575
Number of successful extensions: 3201
Number of sequences better than 10.0: 331
Number of HSP's better than 10.0 without gapping: 2459
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3049
length of query: 191
length of database: 4,897,600
effective HSP length: 85
effective length of query: 106
effective length of database: 2,478,500
effective search space: 262721000
effective search space used: 262721000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0587.5


