

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0587.12
(583 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
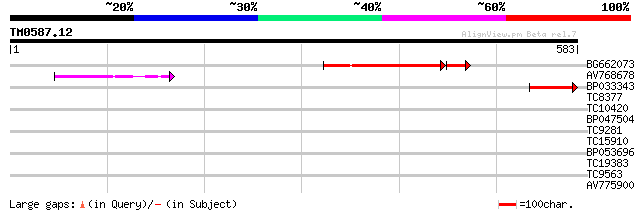
Score E
Sequences producing significant alignments: (bits) Value
BG662073 162 2e-45
AV768678 90 1e-18
BP033343 85 4e-17
TC8377 similar to UP|Q9M3Z7 (Q9M3Z7) Alpha-fucosidase , partial... 29 1.8
TC10420 28 4.1
BP047504 28 5.4
TC9281 similar to UP|O82167 (O82167) T4C15.7 protein (At2g35260/... 28 5.4
TC15910 similar to UP|Q9AR64 (Q9AR64) Urease accessory protein G... 28 5.4
BP053696 27 7.0
TC19383 27 7.0
TC9563 27 7.0
AV775900 27 7.0
>BG662073
Length = 501
Score = 162 bits (409), Expect(2) = 2e-45
Identities = 73/126 (57%), Positives = 90/126 (70%)
Frame = +1
Query: 323 IDAGGSNLPIGGTNDGEFWIDLPCDDDDQREFVKHGDLKGAKLYVHVKPALGGTFTDLAM 382
I G+NLP DG +W+DLP D D+ +E VK GDL+ + YVHVKP LGGTFTDLAM
Sbjct: 37 IAPNGTNLPQDPHTDGAYWLDLPADADN-KERVKKGDLQSFESYVHVKPMLGGTFTDLAM 213
Query: 383 WVFCPFNGPSTLKFGITSIAFSKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVD 442
WVF PFNGP+ K ++ ++GEHVGDWEH TLRI NF G+LW +YF+QHS G WVD
Sbjct: 214 WVFYPFNGPARAKVEFFNVKLGRIGEHVGDWEHVTLRISNFDGQLWKVYFAQHSKGAWVD 393
Query: 443 SYDLEY 448
S +E+
Sbjct: 394 SSQIEF 411
Score = 38.1 bits (87), Expect(2) = 2e-45
Identities = 15/25 (60%), Positives = 19/25 (76%)
Frame = +2
Query: 450 NGNKAIVYSSKSGHASFPHPGTYIQ 474
N N+ +VYSS GHAS+PHPG +Q
Sbjct: 425 NVNRPVVYSSLHGHASYPHPGLILQ 499
>AV768678
Length = 429
Score = 89.7 bits (221), Expect = 1e-18
Identities = 51/124 (41%), Positives = 73/124 (58%), Gaps = 1/124 (0%)
Frame = +3
Query: 47 GQGFASGIINLGELEVCSVTSFELVWNSNVMVEMRKAVAFYKPVGIPDGFHVLSHYCQPS 106
G GFASGII+LG+++V +++F+ VW + F++P +P+GF L Y QP+
Sbjct: 99 GDGFASGIIDLGDVKVSQISTFKKVWTTLEGGPDNAGATFFEPTELPEGFFTLGFYSQPN 278
Query: 107 YNKPLWGFVLVAREVEACSSEKIEIGDQNKLPALRKPLDYTLVWCSNAGSKEI-VTSSAY 165
NKP++G VLVA++ +E AL+KP+DYTLVW SN SK+I Y
Sbjct: 279 -NKPVFGSVLVAKDDSPSDNE-----------ALKKPVDYTLVWSSN--SKKIKQDKDGY 416
Query: 166 FWLP 169
WLP
Sbjct: 417 VWLP 428
>BP033343
Length = 481
Score = 84.7 bits (208), Expect = 4e-17
Identities = 39/49 (79%), Positives = 41/49 (83%)
Frame = -2
Query: 535 ELDKIINALPFRLRISFVNLVKKLPVELYGEEGPTGPKEKNNWIGDERW 583
ELDKIINALP L+ S NL + PVELYGEEGPTGPKEKNNWIGDERW
Sbjct: 480 ELDKIINALPKMLQHSMKNLFNQFPVELYGEEGPTGPKEKNNWIGDERW 334
>TC8377 similar to UP|Q9M3Z7 (Q9M3Z7) Alpha-fucosidase , partial (83%)
Length = 1014
Score = 29.3 bits (64), Expect = 1.8
Identities = 23/71 (32%), Positives = 33/71 (46%)
Frame = +1
Query: 210 QILDVSSEIPEFPFSVWSLRPCDRGMMGKGVSVGTFFCSSCVNMGEQLPVVCLKNLNPVL 269
Q+LDV+ + FP + +RP RG G GV +G S C PV L+ + V
Sbjct: 121 QVLDVNGD-SIFPGGRYYIRPAIRGPPGGGVRLGKTGDSDC-------PVTALQEYSEVK 276
Query: 270 PAMPCLQQIHA 280
+P +I A
Sbjct: 277 NGIPVRFRIAA 309
>TC10420
Length = 871
Score = 28.1 bits (61), Expect = 4.1
Identities = 15/49 (30%), Positives = 25/49 (50%)
Frame = -1
Query: 403 FSKVGEHVGDWEHFTLRICNFSGELWSIYFSQHSGGKWVDSYDLEYING 451
F + H E+ T + N++ I+F Q + G + + DL+YING
Sbjct: 307 FERQATHKNPLENTTFSLNNYN----PIFFCQSAYGTFFRNQDLQYING 173
>BP047504
Length = 486
Score = 27.7 bits (60), Expect = 5.4
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = -1
Query: 8 EIKCQFHSHSPSTVFPSFSTTPMASRSVYSVIVHVCV 44
EI+C H+ S T F FS ++ S+ S +V +C+
Sbjct: 228 EIRCHHHTLSLYTHFKYFSNIVFSTWSLSSCVVSICM 118
>TC9281 similar to UP|O82167 (O82167) T4C15.7 protein (At2g35260/T4C15.7),
partial (31%)
Length = 618
Score = 27.7 bits (60), Expect = 5.4
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = -2
Query: 110 PLWGFVLVAREVEACSSEKIEIGDQN 135
P WGF + E S+ K+E G+QN
Sbjct: 605 PTWGFYFILLENTTYSNSKLENGEQN 528
>TC15910 similar to UP|Q9AR64 (Q9AR64) Urease accessory protein G, partial
(88%)
Length = 1061
Score = 27.7 bits (60), Expect = 5.4
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = +2
Query: 8 EIKCQFHSHSPSTVFPSFSTTPMASRSVYSVIVHVCV 44
EI+C H+ S T F FS ++ S+ S +V +C+
Sbjct: 884 EIRCHHHTLSLYTHFKYFSNIVFSTWSLSSCVVSICM 994
>BP053696
Length = 509
Score = 27.3 bits (59), Expect = 7.0
Identities = 11/17 (64%), Positives = 12/17 (69%)
Frame = -2
Query: 284 HYGPTVYFHPEEVYLPS 300
H GPTVYF EE Y+ S
Sbjct: 91 HIGPTVYFLTEEAYVAS 41
>TC19383
Length = 558
Score = 27.3 bits (59), Expect = 7.0
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +3
Query: 128 KIEIGDQNKLPALRKPLDYTLVWC 151
+IE Q KLP LR P D + +WC
Sbjct: 357 QIEPFFQ*KLPPLRYPFDQSTIWC 428
>TC9563
Length = 757
Score = 27.3 bits (59), Expect = 7.0
Identities = 16/54 (29%), Positives = 30/54 (54%)
Frame = -1
Query: 448 YINGNKAIVYSSKSGHASFPHPGTYIQGSSKLGIGIRNDACSSNLYVDSSIQYE 501
+IN K +V + + + PHP I+ + K I ++ AC+ Y+ S++QY+
Sbjct: 589 FINSTKHLVL*NITSK*TIPHPP*VIRQNFKKLISRKHYACTD--YIISAVQYQ 434
>AV775900
Length = 523
Score = 27.3 bits (59), Expect = 7.0
Identities = 17/50 (34%), Positives = 25/50 (50%), Gaps = 3/50 (6%)
Frame = -2
Query: 14 HSHSPSTVFPSFSTTP--MASRSVYSVIVHVCVITGQGFASGIINL-GEL 60
H HS + PSF T P A + ++ +C+I F+S + L GEL
Sbjct: 258 HGHSDLLLLPSFVTQPASQAGEGILLLLAVLCLIRVNRFSSERVKLVGEL 109
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,504,174
Number of Sequences: 28460
Number of extensions: 199712
Number of successful extensions: 885
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 880
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 881
length of query: 583
length of database: 4,897,600
effective HSP length: 95
effective length of query: 488
effective length of database: 2,193,900
effective search space: 1070623200
effective search space used: 1070623200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0587.12


