

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0546.13
(1118 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
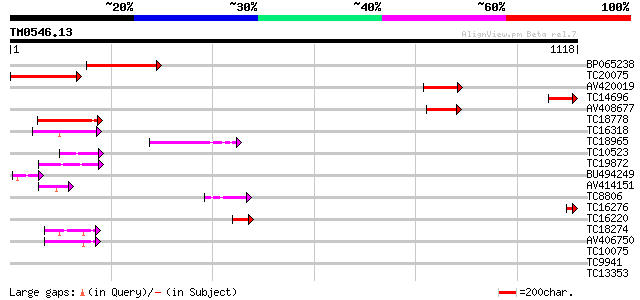
Score E
Sequences producing significant alignments: (bits) Value
BP065238 303 8e-83
TC20075 similar to UP|Q9FG10 (Q9FG10) Ubiquitin carboxyl-termina... 257 8e-69
AV420019 162 2e-40
TC14696 124 8e-29
AV408677 122 4e-28
TC18778 similar to UP|Q8RY18 (Q8RY18) AT5g43560/K9D7_6, partial ... 109 2e-24
TC16318 weakly similar to UP|Q9LUT3 (Q9LUT3) Gb|AAD28294.1, part... 87 2e-17
TC18965 similar to UP|Q9FPS8 (Q9FPS8) Ubiquitin-specific proteas... 67 2e-11
TC10523 64 1e-10
TC19872 similar to UP|Q9LUT3 (Q9LUT3) Gb|AAD28294.1, partial (7%) 59 6e-09
BU494249 53 2e-07
AV414151 53 3e-07
TC8806 similar to GB|AAF21246.1|6648604|AF048705 ubiquitin-speci... 47 1e-05
TC16276 46 3e-05
TC16220 similar to UP|Q9ZSB5 (Q9ZSB5) F3H7.5 protein, partial (6%) 45 8e-05
TC18274 similar to UP|Q8LI88 (Q8LI88) OJ1641_C04.24 protein, par... 43 2e-04
AV406750 43 3e-04
TC10075 similar to UP|SPOP_HUMAN (O43791) Speckle-type POZ prote... 39 0.003
TC9941 39 0.006
TC13353 similar to UP|Q9FPS1 (Q9FPS1) Ubiquitin-specific proteas... 37 0.017
>BP065238
Length = 449
Score = 303 bits (777), Expect = 8e-83
Identities = 144/149 (96%), Positives = 148/149 (98%)
Frame = +3
Query: 151 FTSFMPLGELYDPSRGYLLNDTLVVEAEVLVRRIVDYWTYDSKKETGYVGLKNQGATCYM 210
FTSFMPLGELYDPSRGYL+NDTL++EAEVLVRRIVDYW YDSKKETGYVGLKNQGATCYM
Sbjct: 3 FTSFMPLGELYDPSRGYLVNDTLLIEAEVLVRRIVDYWNYDSKKETGYVGLKNQGATCYM 182
Query: 211 NSLLQTLYHIPYFRKAVYHMPTTENDMPAGSIPLALQSLFYKLQYSDTSVATKELTKSFG 270
NSLLQTLYHIPYFRKAVYHMPTTENDMP+GSIPLALQSLFYKLQYSDTSVATKELTKSFG
Sbjct: 183 NSLLQTLYHIPYFRKAVYHMPTTENDMPSGSIPLALQSLFYKLQYSDTSVATKELTKSFG 362
Query: 271 WDTYDSFMQHDVQELNRVLCEKLEDKMKG 299
WDTYDSFMQHDVQELNRVLCEKLEDKMKG
Sbjct: 363 WDTYDSFMQHDVQELNRVLCEKLEDKMKG 449
>TC20075 similar to UP|Q9FG10 (Q9FG10) Ubiquitin carboxyl-terminal
hydrolase, partial (8%)
Length = 516
Score = 257 bits (656), Expect = 8e-69
Identities = 123/141 (87%), Positives = 130/141 (91%), Gaps = 1/141 (0%)
Frame = +1
Query: 1 MTVMTPAPIDQQEDEEMLVPHTDLPENNHQPMEVVAQPEAAPTVESQPVEEPPQ-SRFTW 59
MTVM APIDQQ+DEE+LVPH DLPENNHQPM+VVAQPE A VESQPV EPP SRFTW
Sbjct: 94 MTVMMSAPIDQQDDEEVLVPHADLPENNHQPMDVVAQPETANAVESQPVAEPPPTSRFTW 273
Query: 60 RIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWSRYAQ 119
RIDNF+R+N KKLYSE+FVVGGYKWRVLIFPKGNNVDYLSMYLDVADS +LPYGWSRYAQ
Sbjct: 274 RIDNFTRLNTKKLYSEIFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSASLPYGWSRYAQ 453
Query: 120 FSLAVVNQIQNKYTVRKDTQH 140
FSLAVVNQIQNKYTVRKDTQH
Sbjct: 454 FSLAVVNQIQNKYTVRKDTQH 516
>AV420019
Length = 231
Score = 162 bits (411), Expect = 2e-40
Identities = 76/76 (100%), Positives = 76/76 (100%)
Frame = +3
Query: 817 DFCLEMSRLYTYDDVVEKVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDML 876
DFCLEMSRLYTYDDVVEKVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDML
Sbjct: 3 DFCLEMSRLYTYDDVVEKVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDML 182
Query: 877 VHYNQTSDILYYEILD 892
VHYNQTSDILYYEILD
Sbjct: 183 VHYNQTSDILYYEILD 230
>TC14696
Length = 629
Score = 124 bits (311), Expect = 8e-29
Identities = 57/57 (100%), Positives = 57/57 (100%)
Frame = +2
Query: 1062 PEYLQDSDIVSNRFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 1118
PEYLQDSDIVSNRFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN
Sbjct: 2 PEYLQDSDIVSNRFQRRDVYGAWEQYLGLEHTDNAPKRSYAVNQNRHTFEKPVKIYN 172
>AV408677
Length = 209
Score = 122 bits (305), Expect = 4e-28
Identities = 57/69 (82%), Positives = 62/69 (89%)
Frame = +2
Query: 823 SRLYTYDDVVEKVAQQLNLDDPSKIRLTPHNCYSQQPKPQPIKYRGVDHLSDMLVHYNQT 882
SR+ TYD VV +VAQ L L+DPSKIRLT HNCYSQ PKPQPIKYRGV+HLSDMLVHYNQT
Sbjct: 2 SRINTYDYVVTRVAQHLGLNDPSKIRLTSHNCYSQLPKPQPIKYRGVEHLSDMLVHYNQT 181
Query: 883 SDILYYEIL 891
SDILYYE+L
Sbjct: 182 SDILYYEVL 208
>TC18778 similar to UP|Q8RY18 (Q8RY18) AT5g43560/K9D7_6, partial (17%)
Length = 738
Score = 109 bits (273), Expect = 2e-24
Identities = 57/130 (43%), Positives = 91/130 (69%), Gaps = 2/130 (1%)
Frame = +2
Query: 56 RFTWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKGNNV-DYLSMYLDVADSTNLPYGW 114
++TW+I+NFS++N ++L S F VG YKW +LI+P+G +V ++LS++L VA+ L GW
Sbjct: 347 KYTWKIENFSQINKRELRSSAFEVGSYKWYILIYPQGCDVCNHLSLFLCVANHDKLLPGW 526
Query: 115 SRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYL-LNDTL 173
S +AQF++AVVN+ K + DT H+F +E DWG+ FM L ++YD G++ +D L
Sbjct: 527 SHFAQFTIAVVNK-DPKKSKYSDTLHRFWKKEHDWGWKKFMELSKVYD---GFVDSSDNL 694
Query: 174 VVEAEVLVRR 183
+++A+V V R
Sbjct: 695 IIKAQVQVIR 724
>TC16318 weakly similar to UP|Q9LUT3 (Q9LUT3) Gb|AAD28294.1, partial (20%)
Length = 550
Score = 86.7 bits (213), Expect = 2e-17
Identities = 49/147 (33%), Positives = 81/147 (54%), Gaps = 11/147 (7%)
Frame = +3
Query: 46 SQPVEEPPQSRFTWRIDNFSRM---NVKKLYSE-VFVVGGYKWRVLIFPKGNNVD----Y 97
S+ V + P + + ++I+++S + V+K ++ VF GGYKWR++++P GN+ +
Sbjct: 72 SRSVRDLPPANYLFKIESYSVLVDTGVEKYETDHVFHAGGYKWRLILYPSGNHKSNGSGH 251
Query: 98 LSMYLDVADSTNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQ---HQFNARESDWGFTSF 154
+S+YL +AD+ +LP GW F L V +Q N Y + +F+ + +WGF
Sbjct: 252 VSLYLAIADTDDLPEGWEVNVNFKLFVFDQKNNNYLTIQAADGAVRKFHEMKKEWGFDQM 431
Query: 155 MPLGELYDPSRGYLLNDTLVVEAEVLV 181
+ L L D GY + DT V AEV V
Sbjct: 432 IELEALLDSCNGYNVKDTCVFGAEVFV 512
>TC18965 similar to UP|Q9FPS8 (Q9FPS8) Ubiquitin-specific protease 16,
partial (17%)
Length = 557
Score = 66.6 bits (161), Expect = 2e-11
Identities = 51/187 (27%), Positives = 90/187 (47%), Gaps = 7/187 (3%)
Frame = +1
Query: 277 FMQHDVQELNRVLCEKLEDKMKGTVVEGT--IQKLFEGHHMNYIECINVDYKSTRKESFY 334
F++H + + + K G++ E T + F G+ + I+C+ KS R E
Sbjct: 19 FLRHAIDTMQSACLVEAGIKASGSLEEDTTLMGLTFGGYLRSKIKCMKCGGKSERHERMM 198
Query: 335 DLQLDVKG-CHDVYASFDKYVEVEPLEGDNKYHAEQ-YGLQDAKKGVLFIDFPPVLQLQL 392
DL ++++G + + ++ E L+G+NKYH ++ + AKK + + P VL + L
Sbjct: 199 DLTVEIEGEITTLVEALRRFTSTEALDGENKYHCDRCKSYEKAKKKLTISEAPNVLTIAL 378
Query: 393 KRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSVLVH---SG 449
KRF+ K+N +FP LDL ++S +D++ +Y L+ V+VH
Sbjct: 379 KRFQ----SGKFGKLNKPIQFPEILDL----APFVSGTSDKS--PIYRLYGVVVHLDIMN 528
Query: 450 GVHGGHY 456
GHY
Sbjct: 529 AAFSGHY 549
>TC10523
Length = 501
Score = 64.3 bits (155), Expect = 1e-10
Identities = 32/90 (35%), Positives = 52/90 (57%), Gaps = 2/90 (2%)
Frame = +2
Query: 98 LSMYLDVADSTNLPYGWSRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESD--WGFTSFM 155
LS+YL++ D P + +F L +++Q+ +KY K H F+A S WG+ F+
Sbjct: 5 LSVYLELTDCEKFPKNRKLFGKFKLGLLDQLHDKY-YEKTASHWFDAPSSQVWWGYKKFV 181
Query: 156 PLGELYDPSRGYLLNDTLVVEAEVLVRRIV 185
PL +L+ ++GYL +D +VV E+L IV
Sbjct: 182 PLSDLHKAAKGYLKDDAIVVNVEILTVSIV 271
>TC19872 similar to UP|Q9LUT3 (Q9LUT3) Gb|AAD28294.1, partial (7%)
Length = 542
Score = 58.5 bits (140), Expect = 6e-09
Identities = 35/131 (26%), Positives = 71/131 (53%), Gaps = 3/131 (2%)
Frame = +2
Query: 58 TWRIDNFSRMNVKKLYSEVFVVGGYKWRVLIFPKG---NNVDYLSMYLDVADSTNLPYGW 114
+W+ + FS+ +++K SE F G YKW+++++P G + +S++L V D + LP
Sbjct: 20 SWKFNKFSQ-SMEKHESESFFGGDYKWKLVLYPNGIVEGKGNSMSLFL-VLDVSTLPPNT 193
Query: 115 SRYAQFSLAVVNQIQNKYTVRKDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLV 174
L +Q++ ++ ++K + +F+ S WG + L ++ D G LL D+ +
Sbjct: 194 KLVVDCILRAKDQLREQHAIQKFCR-KFSESASVWGSRRLVALAKVRDLKNGLLLGDSCI 370
Query: 175 VEAEVLVRRIV 185
+EAE + ++
Sbjct: 371 LEAEFTILGLI 403
>BU494249
Length = 365
Score = 53.1 bits (126), Expect = 2e-07
Identities = 33/74 (44%), Positives = 40/74 (53%), Gaps = 13/74 (17%)
Frame = +1
Query: 6 PAPIDQQE-------------DEEMLVPHTDLPENNHQPMEVVAQPEAAPTVESQPVEEP 52
P P+DQQ+ DEEM VP +D+PE QPM+ AQPE + V+E
Sbjct: 148 PPPLDQQQQQQQEEEEEQQPLDEEMEVPPSDVPEGP-QPMD--AQPETTNLFGAPIVDES 318
Query: 53 PQSRFTWRIDNFSR 66
P RFTW I NFSR
Sbjct: 319 PSGRFTWTIRNFSR 360
>AV414151
Length = 289
Score = 52.8 bits (125), Expect = 3e-07
Identities = 28/77 (36%), Positives = 46/77 (59%), Gaps = 7/77 (9%)
Frame = +1
Query: 57 FTWRIDNFS---RMNVKKLYSEVFVVGGYKWRVLIFP----KGNNVDYLSMYLDVADSTN 109
+T+RI++FS + +V+K SE F GGYKW + I+P KG ++S+YL + D+++
Sbjct: 52 YTFRIESFSMLSKFSVEKCASEEFEAGGYKWSLSIYPTGNTKGEGEGHVSIYLVLMDTSS 231
Query: 110 LPYGWSRYAQFSLAVVN 126
LP W A + + N
Sbjct: 232 LPVDWEVKAIVNFSAYN 282
>TC8806 similar to GB|AAF21246.1|6648604|AF048705 ubiquitin-specific
protease {Arabidopsis thaliana;}, partial (12%)
Length = 751
Score = 47.4 bits (111), Expect = 1e-05
Identities = 30/93 (32%), Positives = 46/93 (49%)
Frame = +2
Query: 385 PPVLQLQLKRFEYDFMRDTMVKINDRYEFPLELDLDRDDGKYLSPDADRNVRNLYTLHSV 444
P VL + LKRF Y R K+ P+ D D Y++ + D + LY L+++
Sbjct: 23 PEVLFIHLKRFSYS--RSMKHKLETFVNLPIH---DFDLTSYIANENDSRPQ-LYELYAL 184
Query: 445 LVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVT 477
H G + GHY A I+ ++WY FDD ++
Sbjct: 185 TNHYGTLGSGHYTAHIKLLDENRWYNFDDTHIS 283
>TC16276
Length = 566
Score = 46.2 bits (108), Expect = 3e-05
Identities = 20/20 (100%), Positives = 20/20 (100%)
Frame = +3
Query: 1099 RSYAVNQNRHTFEKPVKIYN 1118
RSYAVNQNRHTFEKPVKIYN
Sbjct: 3 RSYAVNQNRHTFEKPVKIYN 62
>TC16220 similar to UP|Q9ZSB5 (Q9ZSB5) F3H7.5 protein, partial (6%)
Length = 757
Score = 44.7 bits (104), Expect = 8e-05
Identities = 18/42 (42%), Positives = 26/42 (61%)
Frame = +2
Query: 439 YTLHSVLVHSGGVHGGHYYAFIRPTLSDQWYKFDDERVTKED 480
Y L+++ H GG+ GGHY A+ + ++WY FDD VT D
Sbjct: 20 YNLYAISNHYGGLGGGHYTAYAKLIDENKWYHFDDSHVTPAD 145
>TC18274 similar to UP|Q8LI88 (Q8LI88) OJ1641_C04.24 protein, partial (40%)
Length = 622
Score = 43.1 bits (100), Expect = 2e-04
Identities = 33/122 (27%), Positives = 54/122 (44%), Gaps = 12/122 (9%)
Frame = +1
Query: 70 KKLYSEVFVVGGYKWRVLIFPKGNNVD----YLSMYLDVADSTNLPYGWSRYAQFSLAVV 125
K + SE F VGGY+W + +P G N + Y+S+++ +A G A F L ++
Sbjct: 208 KHIASETFTVGGYQWAIYFYPDGKNPEDNSAYVSVFIALASE-----GTDVRALFELTLL 372
Query: 126 NQIQN-KYTVRKDTQHQFNA-------RESDWGFTSFMPLGELYDPSRGYLLNDTLVVEA 177
+Q N K+ V + R S WG+ F +L + +L +D L +
Sbjct: 373 DQSPNAKHKVHSHFDRSLESGPYTLKYRGSMWGYKRFFKRAQL--EASTFLKDDCLKINC 546
Query: 178 EV 179
V
Sbjct: 547 TV 552
>AV406750
Length = 426
Score = 42.7 bits (99), Expect = 3e-04
Identities = 33/118 (27%), Positives = 51/118 (42%), Gaps = 8/118 (6%)
Frame = +3
Query: 70 KKLYSEVFVVGGYKWRVLIFPKGNNVDYLSMYLDVADSTNLPYGWSRYAQFSLAVVNQI- 128
K + S+VF VGGY+W + +P G N + S Y+ V + G A F L +++Q
Sbjct: 72 KHIASDVFTVGGYQWAIYFYPDGKNPEDNSAYVSVFIALE-SEGTDVRALFELTLMDQSG 248
Query: 129 QNKYTVRKDTQHQFNA-------RESDWGFTSFMPLGELYDPSRGYLLNDTLVVEAEV 179
Q K+ V + + S WG+ F L + +L ND L + V
Sbjct: 249 QGKHKVHSHFDRSLESGPYTLKYKRSMWGYKRFFRRSLL--ENSDFLRNDCLKINCTV 416
>TC10075 similar to UP|SPOP_HUMAN (O43791) Speckle-type POZ protein, partial
(5%)
Length = 1051
Score = 39.3 bits (90), Expect = 0.003
Identities = 33/118 (27%), Positives = 51/118 (42%), Gaps = 12/118 (10%)
Frame = +1
Query: 74 SEVFVVGGYKWRVLIFPKGNNVD----YLSMYLDVADSTNLPYGWSRYAQFSLAVVNQI- 128
S+ F VGGY W + +P G N + Y+S+++ +A G A F L +++Q
Sbjct: 337 SDTFSVGGYDWAIYFYPDGKNPEDNSVYVSVFIALASD-----GVDVRALFKLTLMDQTN 501
Query: 129 QNKYTVR-------KDTQHQFNARESDWGFTSFMPLGELYDPSRGYLLNDTLVVEAEV 179
+ + V + + R S WG+ F L S YL ND LV+ V
Sbjct: 502 KGNHKVHSHFDRPLESGPYSLKYRGSMWGYKRFFRRSLL--ESSEYLKNDCLVMHCTV 669
>TC9941
Length = 557
Score = 38.5 bits (88), Expect = 0.006
Identities = 25/97 (25%), Positives = 45/97 (45%), Gaps = 8/97 (8%)
Frame = +2
Query: 43 TVESQPVEEPPQSRFTWRIDNFSRMNV----KKLYSEVFVVGGYKWRVLIFPKG----NN 94
T S + E + ++I +S K + S++F VGGY W + +P G +N
Sbjct: 260 TTSSTSITETVRGSHQFKITGYSLSKGIGIGKYIASDIFTVGGYDWAIYFYPDGKSPEDN 439
Query: 95 VDYLSMYLDVADSTNLPYGWSRYAQFSLAVVNQIQNK 131
Y+S+++ +A G A F L +++Q N+
Sbjct: 440 ASYVSLFIALASD-----GTDVRALFELTLLDQSGNE 535
>TC13353 similar to UP|Q9FPS1 (Q9FPS1) Ubiquitin-specific protease 26,
partial (11%)
Length = 532
Score = 37.0 bits (84), Expect = 0.017
Identities = 16/29 (55%), Positives = 20/29 (68%)
Frame = +2
Query: 200 GLKNQGATCYMNSLLQTLYHIPYFRKAVY 228
GL N GATCY NS+LQ LY FR+ ++
Sbjct: 443 GLTNLGATCYANSILQCLYMNKAFREGIF 529
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,046,799
Number of Sequences: 28460
Number of extensions: 237884
Number of successful extensions: 1287
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 1215
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1266
length of query: 1118
length of database: 4,897,600
effective HSP length: 100
effective length of query: 1018
effective length of database: 2,051,600
effective search space: 2088528800
effective search space used: 2088528800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0546.13


