

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0545.3
(2087 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
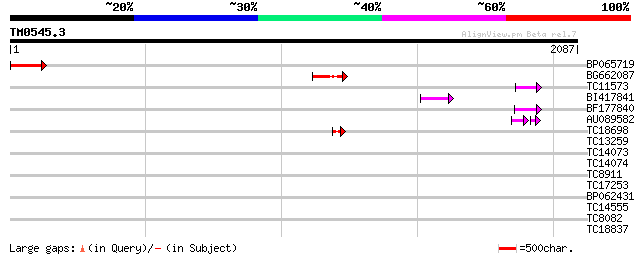
Score E
Sequences producing significant alignments: (bits) Value
BP065719 198 6e-51
BG662087 108 1e-23
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 74 3e-13
BI417841 72 1e-12
BF177840 56 7e-08
AU089582 38 2e-04
TC18698 44 3e-04
TC13259 34 0.27
TC14073 homologue to UP|AAQ08403 (AAQ08403) Methionine synthase,... 32 0.78
TC14074 homologue to UP|AAQ08403 (AAQ08403) Methionine synthase,... 32 1.0
TC8911 homologue to UP|Q8W0Q7 (Q8W0Q7) Methionine synthase prote... 31 1.7
TC17253 similar to UP|Q8SKU2 (Q8SKU2) Tic62 protein precursor, p... 30 3.9
BP062431 30 3.9
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 30 5.1
TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-... 29 8.7
TC18837 weakly similar to UP|Q39856 (Q39856) Epoxide hydrolase ... 29 8.7
>BP065719
Length = 567
Score = 198 bits (504), Expect = 6e-51
Identities = 115/135 (85%), Positives = 117/135 (86%)
Frame = +1
Query: 1 FLEAGRSPNSLSSPGIQESLL*NT*PGTKSKQGT*PSMKT*K*STSRVP*LRTHLPGSPP 60
FLEAGRSPNSLSS GIQES *NT*PGTKSKQ T*PSMK *K*STSRV * + LPGSPP
Sbjct: 163 FLEAGRSPNSLSSQGIQESPR*NT*PGTKSKQET*PSMKI*K*STSRVL*PKMPLPGSPP 342
Query: 61 SLLGQSIHGPSWKEFSMNSSLGESAK*V*RIWLVSKESQQNLLMII*TGLGCLNLDVSLT 120
SLLGQSIHGPSWKEFSMNSSLGESAK*V R WLVSKESQQ+LLMI *TGLGCLNLD S
Sbjct: 343 SLLGQSIHGPSWKEFSMNSSLGESAK*VXRTWLVSKESQQSLLMIT*TGLGCLNLDASPM 522
Query: 121 SLNRS*LS*PLVV*S 135
LN S*LS* LVV*S
Sbjct: 523 CLNTS*LS*XLVV*S 567
>BG662087
Length = 373
Score = 108 bits (269), Expect = 1e-23
Identities = 58/127 (45%), Positives = 81/127 (63%)
Frame = +1
Query: 1116 NGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA*EDVS 1175
+GK R+ +D+ DLN A PKD Y +P + +VD A+ +E LSL+D YSGY+QI + D
Sbjct: 13 SGKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDED 192
Query: 1176 KTAFRCPGALGTYEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVK 1235
KTAF A Y + +PF GLKNAGATYQ +M+ +F D + M+VY+D+++VK
Sbjct: 193 KTAFMT--ARVNYCYQTIPF-----GLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVK 351
Query: 1236 SPSRDGH 1242
S R H
Sbjct: 352 SALRANH 372
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 73.6 bits (179), Expect = 3e-13
Identities = 34/96 (35%), Positives = 53/96 (54%)
Frame = +2
Query: 1861 DQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPKRWHE 1920
+ G F ++ F + GI++ S+ + Q NGQ EAANK+++ IK+ + RW +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 1921 SLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEV 1956
L VLW+Y P + TPF L YG + +LP E+
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEI 295
>BI417841
Length = 617
Score = 71.6 bits (174), Expect = 1e-12
Identities = 44/123 (35%), Positives = 66/123 (52%), Gaps = 2/123 (1%)
Frame = +1
Query: 1512 LYFDGSSHKN--GTGIGMFIMSPQGAPTKFKFRIEGNCSNNEVEYEALISGLEILLALGA 1569
L FDGSS N G G + + G+ + + GN +NN+ EY LI GL+ G
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKVYLREGV-GNQTNNQAEYRGLILGLKHAHEQGY 318
Query: 1570 KNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNEIGIGHVPRIENQEANEL 1629
+++ +KGDS+LV KQ+ +K + ++A +A L +KF I HVPR N EA+
Sbjct: 319 QHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEADVQ 498
Query: 1630 AQI 1632
A +
Sbjct: 499 ANL 507
>BF177840
Length = 410
Score = 55.8 bits (133), Expect = 7e-08
Identities = 31/100 (31%), Positives = 47/100 (47%)
Frame = +2
Query: 1857 SITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILISLIKKHVGRKPK 1916
SI +D+ T F+ G KLL ST + Q +GQ E NK L +L++ + R K
Sbjct: 14 SIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLK 193
Query: 1917 RWHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLPAEV 1956
W L + +AY T +PF + YG + P ++
Sbjct: 194 MWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>AU089582
Length = 383
Score = 37.7 bits (86), Expect(2) = 2e-04
Identities = 20/63 (31%), Positives = 34/63 (53%)
Frame = +2
Query: 1846 NHIVYRFGLPESITTDQGTVFVGRKVAAFAESWGIKLLTSTPYYAQANGQVEAANKILIS 1905
+ IV G+P SI +D+G F +F + G +L ST ++ Q +GQ E +IL
Sbjct: 38 DEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSERTIQILED 217
Query: 1906 LIK 1908
+++
Sbjct: 218 MLR 226
Score = 25.4 bits (54), Expect(2) = 2e-04
Identities = 12/36 (33%), Positives = 17/36 (46%)
Frame = +3
Query: 1918 WHESLSQVLWAYRNSPTEATGTTPFRLAYGQEAVLP 1953
W + LS + +AY NS + PF YG+ P
Sbjct: 255 WDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSP 362
>TC18698
Length = 808
Score = 43.5 bits (101), Expect = 3e-04
Identities = 23/49 (46%), Positives = 32/49 (64%)
Frame = -2
Query: 1188 YEWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKS 1236
Y + VMP GLKN TYQR+M+ IFH I ++VY++D++VKS
Sbjct: 771 YYYQVMPLGLKNI*-----TTYQRLMDKIFHKQI*KNVEVYVEDMIVKS 640
>TC13259
Length = 506
Score = 33.9 bits (76), Expect = 0.27
Identities = 26/81 (32%), Positives = 40/81 (49%)
Frame = +1
Query: 1553 EYEALISGLEILLALGAKNVVIKGDSELVVKQLTKEYKCVSEHLAKYYVKANNLLAKFNE 1612
E AL GL++ L LG V ++ D+++VV K VS +LA + F
Sbjct: 112 EAMALRWGLQLSLELGFFEVEVEIDNQVVVNSWKGTKKHVS-YLANVIADVVRISQCFRF 288
Query: 1613 IGIGHVPRIENQEANELAQIA 1633
+ +VPR+ + A+ LAQ A
Sbjct: 289 FSLAYVPRLSHLAADYLAQFA 351
>TC14073 homologue to UP|AAQ08403 (AAQ08403) Methionine synthase, complete
Length = 2757
Score = 32.3 bits (72), Expect = 0.78
Identities = 18/74 (24%), Positives = 35/74 (46%)
Frame = +3
Query: 1189 EWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRK 1248
+W V F + N G+++ + + + F+D I + + + D I +++ D LL +
Sbjct: 2049 DWAVHSFRITNVGVQDTTQIHTHMCYSNFNDIIHSIIDMDADVITIENSRSDEKLLSV-- 2222
Query: 1249 SFERMRKYGLKMNP 1262
F KYG + P
Sbjct: 2223 -FREGVKYGAGIGP 2261
>TC14074 homologue to UP|AAQ08403 (AAQ08403) Methionine synthase, partial
(32%)
Length = 1107
Score = 32.0 bits (71), Expect = 1.0
Identities = 18/74 (24%), Positives = 35/74 (46%)
Frame = +2
Query: 1189 EWVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRK 1248
+W V F + N G+++ + + + F+D I + + + D I +++ D LL +
Sbjct: 320 DWAVHAFRITNIGVQDTTQIHTHMCYSNFNDIIHSIIDMDADVITIENSRSDEKLLSV-- 493
Query: 1249 SFERMRKYGLKMNP 1262
F KYG + P
Sbjct: 494 -FREGVKYGAGIGP 532
>TC8911 homologue to UP|Q8W0Q7 (Q8W0Q7) Methionine synthase protein, partial
(58%)
Length = 1786
Score = 31.2 bits (69), Expect = 1.7
Identities = 18/73 (24%), Positives = 33/73 (44%)
Frame = +1
Query: 1190 WVVMPFGLKNAGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDGHLLHLRKS 1249
W V F + N G+ + + + + F+D I + + + D I +++ D LL +
Sbjct: 997 WAVHSFRITNCGVNDTTQIHTHMCYSNFNDIIHSIIDMDADVITIENSRSDEKLLSV--- 1167
Query: 1250 FERMRKYGLKMNP 1262
F KYG + P
Sbjct: 1168 FREGVKYGAGIGP 1206
>TC17253 similar to UP|Q8SKU2 (Q8SKU2) Tic62 protein precursor, partial (9%)
Length = 793
Score = 30.0 bits (66), Expect = 3.9
Identities = 31/117 (26%), Positives = 56/117 (47%), Gaps = 2/117 (1%)
Frame = -2
Query: 858 IPLRD*TLTKLTNLMP*VDGTTMKKRMPLLSGKTMLKMINSSLARISAYAAENRIRTALE 917
+P D T+ + L P VD T++ T+L + ++S+ + ++ T++E
Sbjct: 318 LPTVDSTVVEAATLSP-VDSTSV-------GAATLLPVDSTSVEAATLSPVDS---TSVE 172
Query: 918 AARIAKEMAIDEANVSILPVETTSQEEATTNK-DNTCIEENEQS-VDECEFEDLRNL 972
AA ++ + ++LPV++TS E T + D+T +E S VD E R L
Sbjct: 171 AATLSPVDSTSVGAATLLPVDSTSVEAETLSPVDSTSVEAATLSPVDSTSVEAARGL 1
>BP062431
Length = 482
Score = 30.0 bits (66), Expect = 3.9
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = -3
Query: 1541 FRIEGNCSNNEVEYEALISGLEIL 1564
F+ G CSNNE E A++ GL +L
Sbjct: 405 FQFFGRCSNNEAELRAILEGLLVL 334
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 29.6 bits (65), Expect = 5.1
Identities = 14/26 (53%), Positives = 16/26 (60%)
Frame = -3
Query: 150 CLCSQIE*GTSNY*RPKSLRKGPRLL 175
CL Q+E G + Y* P L GPRLL
Sbjct: 751 CLDQQVEEGRAEY*APNHLLHGPRLL 674
>TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(54%)
Length = 1913
Score = 28.9 bits (63), Expect = 8.7
Identities = 15/46 (32%), Positives = 23/46 (49%), Gaps = 9/46 (19%)
Frame = +1
Query: 1969 IPSEVYWNMMLD---------EMVNLDEERLLALDVLTRQKDRIAK 2005
+P E+YW MML E++ D +D T+ KD+I+K
Sbjct: 97 LPPEIYWKMMLPSTPMPKAVIELLRGDVSIETCVDTYTQSKDKISK 234
>TC18837 weakly similar to UP|Q39856 (Q39856) Epoxide hydrolase , partial
(89%)
Length = 1100
Score = 28.9 bits (63), Expect = 8.7
Identities = 31/111 (27%), Positives = 49/111 (43%), Gaps = 4/111 (3%)
Frame = +2
Query: 485 GLSQHLQGQNHTPSGRKSTWARGLLTSTIPVCHGTIPMAPGTTTLLNK*PELSSDVS*GK 544
G SQH +N + W + + V T + P L K* + * K
Sbjct: 704 GSSQHHGLENRSKCRPSLLWETWTVHIIV*VLRSTYIVVP-----LRK*YHTCKNWL*WK 868
Query: 545 EKL----KEKEVASSVHYGT*SQKTTLAMNLVWRRLLIN*ITPSKREQLCR 591
E L ++ + S++ Y T S+ + NLV+ L++N*IT S + +CR
Sbjct: 869 EWLTL*IRKDQKRSALIYMTSSRNS----NLVFNNLMVN*ITSSIKHNMCR 1009
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.333 0.143 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,312,584
Number of Sequences: 28460
Number of extensions: 493138
Number of successful extensions: 3661
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 3585
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3657
length of query: 2087
length of database: 4,897,600
effective HSP length: 104
effective length of query: 1983
effective length of database: 1,937,760
effective search space: 3842578080
effective search space used: 3842578080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0545.3


