

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0544.8
(1753 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
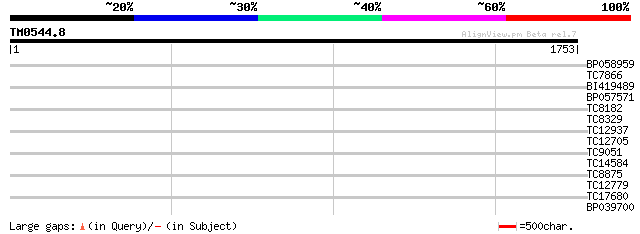
Score E
Sequences producing significant alignments: (bits) Value
BP058959 34 0.23
TC7866 homologue to UP|SAHH_CATRO (P35007) Adenosylhomocysteinas... 33 0.39
BI419489 30 2.5
BP057571 30 2.5
TC8182 weakly similar to UP|Q93VA8 (Q93VA8) At1g76010/T4O12_22, ... 30 2.5
TC8329 similar to UP|BAD06717 (BAD06717) WRKY transcription fact... 29 5.6
TC12937 29 5.6
TC12705 29 7.3
TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 ... 29 7.3
TC14584 similar to UP|Q39193 (Q39193) Protein kinase (Protein ki... 29 7.3
TC8875 29 7.3
TC12779 weakly similar to UP|Q9FMB9 (Q9FMB9) Cysteine-tRNA ligas... 29 7.3
TC17680 similar to UP|O65440 (O65440) CLV1 receptor kinase like ... 28 9.6
BP039700 28 9.6
>BP058959
Length = 435
Score = 33.9 bits (76), Expect = 0.23
Identities = 19/68 (27%), Positives = 40/68 (57%)
Frame = -1
Query: 326 VSLKFWIENMFTLFGLEINMIALLLASFAVLNAISLLYIASLAACVLLHRLLIKKLWPIF 385
V + +W+ +F +I +LL+ ++ + ISLL ++ L +CV+LH LL K +W +
Sbjct: 180 VHILWWVVALF--------VILILLSKYSYV--ISLLILSHLGSCVVLH-LLEKPIWLRY 34
Query: 386 VFLFASVV 393
+ + +++
Sbjct: 33 IIFYPNII 10
>TC7866 homologue to UP|SAHH_CATRO (P35007) Adenosylhomocysteinase
(S-adenosyl-L-homocysteine hydrolase) (AdoHcyase) ,
partial (38%)
Length = 561
Score = 33.1 bits (74), Expect = 0.39
Identities = 24/96 (25%), Positives = 44/96 (45%)
Frame = +2
Query: 1598 YFRVTGSGDVRSLEQSVELVSGDLVLNRGNPEWWSFYDLDISDAHGCGKFPGPMAIIVSE 1657
+ TG+ D+ ++ ++ + ++ N G+ +D +I D HG +PG I +
Sbjct: 251 FVTTTGNKDIIMVDHMRKMKNNAIISNIGH------FDNEI-DMHGLETYPGVKRITIKP 409
Query: 1658 ETPQGIIGDTLSKFSIWGLYITFVLAVGRFIRLQCS 1693
+T + + DT S VLA GR + L C+
Sbjct: 410 QTDRWVFPDTNSGI--------IVLAEGRLMNLGCA 493
>BI419489
Length = 479
Score = 30.4 bits (67), Expect = 2.5
Identities = 17/50 (34%), Positives = 24/50 (48%)
Frame = +1
Query: 891 HIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSY 940
H R RSN + C CCF I + F+ L+ + A LY L + P +
Sbjct: 211 HYNRRHRSNRSLCCCCCFWTILILLFAALAAAIVGAA-LYVLYRPHRPEF 357
>BP057571
Length = 559
Score = 30.4 bits (67), Expect = 2.5
Identities = 22/61 (36%), Positives = 32/61 (52%)
Frame = -3
Query: 889 VRHIWSRMRSNNEVVCYCCFILIYLWNFSLLSVVYLAALFLYALCQNTGPSYIFWVLILI 948
+R I S + EV C+ F +I SLL +Y A L+ LC ++ S+IF LIL
Sbjct: 296 IRAILS*KKGFIEVACFL-F*IICFRILSLL*DIYYIARSLHILCNHSNFSFIFCSLILY 120
Query: 949 Y 949
+
Sbjct: 119 F 117
>TC8182 weakly similar to UP|Q93VA8 (Q93VA8) At1g76010/T4O12_22, partial
(13%)
Length = 611
Score = 30.4 bits (67), Expect = 2.5
Identities = 16/55 (29%), Positives = 26/55 (47%), Gaps = 4/55 (7%)
Frame = -1
Query: 223 WNEPCPLFNMVDIPNETTACTPQNKQAETSTSATIKRLA----RSRSCPTVNSAL 273
W C +F +V +PN + ET+TSAT K A + + PT+ + +
Sbjct: 203 WGRWCIIFTIVIVPNWLGTIASPSSSPETTTSATTKATASATIATSTVPTITTTI 39
>TC8329 similar to UP|BAD06717 (BAD06717) WRKY transcription factor 1,
partial (41%)
Length = 1717
Score = 29.3 bits (64), Expect = 5.6
Identities = 18/50 (36%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Frame = +1
Query: 914 WNFSLLSVVYLAALFLYALCQNT---GPSYIFWVL--ILIYTEVCILLQY 958
+NF LL + + AAL L L NT PS+IF++L +L +V +++++
Sbjct: 1363 YNF-LLFIFFSAALLLNVLTSNTQT*APSFIFFILEIVLCINQVLLMMKF 1509
>TC12937
Length = 526
Score = 29.3 bits (64), Expect = 5.6
Identities = 9/22 (40%), Positives = 17/22 (76%)
Frame = -2
Query: 444 RMLIGYYGVFMFSCFKFRADKS 465
R L+ ++ F+FSCF+F+++ S
Sbjct: 165 RYLVSFFSFFLFSCFRFQSEAS 100
>TC12705
Length = 376
Score = 28.9 bits (63), Expect = 7.3
Identities = 15/36 (41%), Positives = 19/36 (52%)
Frame = -3
Query: 1034 YLEDVKCSTYHERVQRLFLPLSNALKLLTRSLCRYW 1069
YLED S +H+R Q L+ S L LL C Y+
Sbjct: 269 YLEDQSLSKHHQR*QELYPHFSIMLTLLLMLFCVYF 162
>TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a),
partial (46%)
Length = 867
Score = 28.9 bits (63), Expect = 7.3
Identities = 28/118 (23%), Positives = 57/118 (47%)
Frame = -1
Query: 1365 AIPFLYELRCVLDWSCTTTSLTMYDWLKLEDIHASLFLVKCDVDLNRASHQQGQKQTKMT 1424
A+ F Y L + S + S+T++ ++ I AS+ + L A + GQ++ + +
Sbjct: 564 ALNFWYPLMTLFTASRKSFSVTVFLRARIAYIPASVHTL-----LISAPVEFGQRRDRSS 400
Query: 1425 KFCNGICLFFVLMCVIWAPMLMYSSGNPTNIANPIKDASARVHIKTLSGRLTLFETTL 1482
K + + F VL+C++ +L+ SG P + + +R + +SG L +T +
Sbjct: 399 KRIS-LSQFIVLVCILNICVLLSKSGRPNSTFRSNRPGRSRAGSR-VSGLLVAIKTLM 232
>TC14584 similar to UP|Q39193 (Q39193) Protein kinase (Protein kinase, 41K)
, partial (10%)
Length = 641
Score = 28.9 bits (63), Expect = 7.3
Identities = 16/65 (24%), Positives = 30/65 (45%)
Frame = +3
Query: 397 YLAIWMRLTFMNLEIEEHVPCRDCWRVSDIYFSYCRKCWLGIIVDDPRMLIGYYGVFMFS 456
++ +W+ L FM+L + H PC+ DI ++ C ++ ++ Y+ S
Sbjct: 471 FVVLWLELCFMSLCLYAHTPCKKF--PVDIIVTFASICVCNLV----SCVVAYF--IFHS 626
Query: 457 CFKFR 461
C FR
Sbjct: 627 CIPFR 641
>TC8875
Length = 579
Score = 28.9 bits (63), Expect = 7.3
Identities = 16/52 (30%), Positives = 26/52 (49%), Gaps = 6/52 (11%)
Frame = +2
Query: 343 INMIALLLASFAVLNAISLLYIASLAACV------LLHRLLIKKLWPIFVFL 388
++ +A LA F + I L+++ + V LL +L+ LW FVFL
Sbjct: 359 VSSVAFALAHFNIQRMIPLIFLGIVMGAVFVRSRNLLPSMLLHSLWNAFVFL 514
>TC12779 weakly similar to UP|Q9FMB9 (Q9FMB9) Cysteine-tRNA ligase, partial
(45%)
Length = 811
Score = 28.9 bits (63), Expect = 7.3
Identities = 19/74 (25%), Positives = 33/74 (43%)
Frame = +1
Query: 522 ITGTLEYDFLHLGYLGFALVFFRMRLKILKQGNNIFRYLRMYNFAVIVLSLAYQSPFLGD 581
+ G YDF HLG+ A+ F + +FRYL+ L Y+ ++ +
Sbjct: 202 VCGVTAYDFSHLGHARAAVSF-----------DILFRYLK---------HLGYEVTYVRN 321
Query: 582 FSEIKSGFIEYINE 595
F+++ I+ NE
Sbjct: 322 FTDVDDKIIKRANE 363
>TC17680 similar to UP|O65440 (O65440) CLV1 receptor kinase like protein,
partial (16%)
Length = 685
Score = 28.5 bits (62), Expect = 9.6
Identities = 29/109 (26%), Positives = 45/109 (40%), Gaps = 1/109 (0%)
Frame = +1
Query: 477 IHSQWKSASVLSNLSFETKGYWTFLDHLRLYGYCHLLDFVLSLILI-TGTLEYDFLHLGY 535
IH KS ++L N FE H+ DF L+ L TGT + G
Sbjct: 25 IHRDVKSNNILLNSEFEA----------------HVADFGLAKFLHDTGTSQCMSSIAGS 156
Query: 536 LGFALVFFRMRLKILKQGNNIFRYLRMYNFAVIVLSLAYQSPFLGDFSE 584
G+ + LK+ ++ + +Y+F V++L L +GDF E
Sbjct: 157 YGYIAPEYAYTLKVDEKSD-------VYSFGVVLLELLTGRRPVGDFGE 282
>BP039700
Length = 506
Score = 28.5 bits (62), Expect = 9.6
Identities = 17/56 (30%), Positives = 28/56 (49%), Gaps = 8/56 (14%)
Frame = +3
Query: 1671 FSIWGLYITFVLAVGRFIRLQCS---DLRMRIPFENL-----PSCDRLMALCEDIY 1718
F + LY+ L RF+ C ++ +R+ F+ L P C RL+ L +D+Y
Sbjct: 252 FRLCSLYLVLTLFSKRFLVPDCQQ**EVHLRMLFQLLTLSLTPHCKRLLILMKDLY 419
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,388,435
Number of Sequences: 28460
Number of extensions: 494972
Number of successful extensions: 3706
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 3640
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3705
length of query: 1753
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1650
effective length of database: 1,966,220
effective search space: 3244263000
effective search space used: 3244263000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0544.8


