

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.15
(322 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
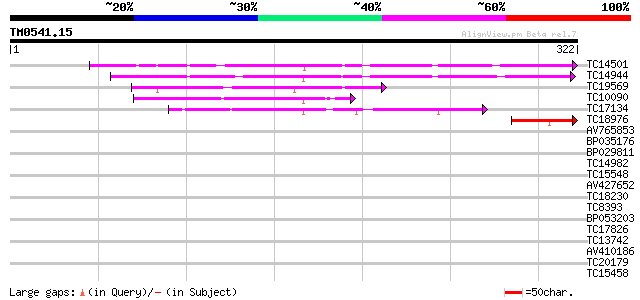
Score E
Sequences producing significant alignments: (bits) Value
TC14501 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA),... 133 4e-32
TC14944 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA),... 132 6e-32
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 98 2e-21
TC10090 weakly similar to UP|Q9ZWP6 (Q9ZWP6) Lectin, partial (43%) 77 4e-15
TC17134 weakly similar to UP|LEC_BOWMI (P42088) Lectin (Agglutin... 46 1e-05
TC18976 similar to UP|LCB2_ROBPS (Q42372) Bark agglutinin I, pol... 43 8e-05
AV765853 38 0.002
BP035176 36 0.008
BP029811 30 0.72
TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%) 29 0.94
TC15548 similar to UP|AAR87327 (AAR87327) Expressed protein, par... 28 2.7
AV427652 28 2.7
TC18230 similar to UP|Q945U3 (Q945U3) Acyl-CoA oxidase , partia... 28 2.7
TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenyla... 27 3.6
BP053203 27 3.6
TC17826 similar to UP|AAR91698 (AAR91698) DNA-binding protein, p... 27 3.6
TC13742 similar to UP|Q9ZR53 (Q9ZR53) Annexin-like protein, part... 27 6.1
AV410186 27 6.1
TC20179 27 6.1
TC15458 26 7.9
>TC14501 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA), partial
(86%)
Length = 1066
Score = 133 bits (335), Expect = 4e-32
Identities = 103/282 (36%), Positives = 151/282 (53%), Gaps = 5/282 (1%)
Frame = +1
Query: 46 KHHQYSQLNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGD 105
+ + YS + + A+ + LLA L F LL I + + ++SF+F F D +I+ GD
Sbjct: 22 QQYTYSSMATPYSNASTQNFLLACLFF-LLAINVNLAA-AISFNFTEFTDDGSLII-QGD 192
Query: 106 ANT-TNGTLQLTKIDQYGNPSPHSSGISAFYGA-VHLSDKKTGRVADFRTEFTFVVNHSQ 163
A +G+L L +PS + A Y V + D TG VA F T F+F++ +
Sbjct: 193 AKIWADGSLALPT-----DPSVGFTTSRALYATPVPIWDSTTGNVASFVTSFSFIIKDFE 357
Query: 164 PHG--DGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPN 221
+ DG FFLA E P+ GG F G+ + + A+N IVAVEFD+F N +D +
Sbjct: 358 DYNPADGLVFFLAPFGTEIPKESTGGRF-GIINGKDAFN----QIVAVEFDTFINPWDSS 522
Query: 222 FPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDA 281
HIGID+N++ S T+PW ++ G+++ I Y S ++ L+V+V + GQ I
Sbjct: 523 PRHIGIDVNSLISLKTVPW---NKVAGSLEKVTIIYDSQTKTLSVLVIHENGQ----IST 681
Query: 282 IKYPIDIESYLSEWVVIGFSATTGEFA-ETHSILSWSFNSTL 322
I ID++ L E V +GFSATT E H I SWSF STL
Sbjct: 682 ISQEIDLKVVLPEEVSVGFSATTTSGGRERHDIYSWSFTSTL 807
>TC14944 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA), partial
(85%)
Length = 1155
Score = 132 bits (333), Expect = 6e-32
Identities = 102/269 (37%), Positives = 139/269 (50%), Gaps = 5/269 (1%)
Frame = +2
Query: 58 TMANITSELLATLIFSLLLITTTVKS-DSLSFSFPTFDPDVRVILLNGDANT-TNGTLQL 115
T+ + S L L LLL+T V S + +SF+F F D +IL GDA T+G L L
Sbjct: 41 TLYSNPSTLKVVLASLLLLLTIKVNSVEGVSFNFTKFTDDGSLIL-QGDAKIWTDGRLAL 217
Query: 116 TKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPH--GDGFTFFL 173
G + + V + D TG VA F + F+FVV+ + DG FFL
Sbjct: 218 PSDPLIGKTTSRV----LYATPVPIWDSTTGSVASFVSSFSFVVSDVPGYKISDGLVFFL 385
Query: 174 ASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIR 233
A P G LG+ DE+ YN VAVEFDS++N+ DP F HIGID+N++
Sbjct: 386 APWGTTIPPNSEGKN-LGILDEKNGYN----QFVAVEFDSYNNTRDPTFRHIGIDVNSVM 550
Query: 234 SAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLS 293
S + W ++ GA++ A I Y S ++ L+VVV + GQ TT + ID++ L
Sbjct: 551 SMNLVKW---NRVSGALEKAIIIYDSPTKTLSVVVTHQNGQITT----VAQQIDLKVVLP 709
Query: 294 EWVVIGFSATT-GEFAETHSILSWSFNST 321
+ V +GFSATT E H I SWSF ST
Sbjct: 710 KEVSVGFSATTWNTHRERHDIHSWSFTST 796
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 98.2 bits (243), Expect = 2e-21
Identities = 58/150 (38%), Positives = 81/150 (53%), Gaps = 5/150 (3%)
Frame = +3
Query: 70 LIFSLLLITTTVK---SDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSP 126
L+ LLL+TTT + SLSF+ FD D I GD +TNG++ L K+ +
Sbjct: 48 LLLLLLLLTTTTTFPTTHSLSFNITNFDDDTTAIAYEGDGKSTNGSIDLNKVSYF----- 212
Query: 127 HSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN--HSQPHGDGFTFFLASLDFEFPEGG 184
G + +HL D T + DF T FTF ++ + +GDGF F+LA L ++ P
Sbjct: 213 FRVGRAISAQPLHLYDPTTKTLTDFTTRFTFTISAINKTSYGDGFAFYLAPLGYQIPPNS 392
Query: 185 GGGGFLGLFDEETAYNTSKNSIVAVEFDSF 214
GG F GLF+ T N +N +VAVEFD+F
Sbjct: 393 AGGTF-GLFNATTNNNLPQNHVVAVEFDTF 479
>TC10090 weakly similar to UP|Q9ZWP6 (Q9ZWP6) Lectin, partial (43%)
Length = 461
Score = 77.0 bits (188), Expect = 4e-15
Identities = 50/129 (38%), Positives = 71/129 (54%), Gaps = 3/129 (2%)
Frame = +3
Query: 71 IFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTT-NGTLQLTKIDQYGNPSPHSS 129
+F LLL+ +D LSF+F F P +L GDA G LQLTK++ P S+
Sbjct: 81 LFFLLLVNKVNSTDYLSFTFNQFTPQQPDLLFQGDALVAPTGKLQLTKVEN-DLPVYKST 257
Query: 130 GISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPH--GDGFTFFLASLDFEFPEGGGGG 187
G + + VH+ D KTG VA F T F+F+++ + DG FFLA +D + P+ G
Sbjct: 258 GRALYVAPVHIWDSKTGNVASFITSFSFIIDSPNVNKIADGLAFFLAPVDTQ-PQ--KPG 428
Query: 188 GFLGLFDEE 196
G LG+FD +
Sbjct: 429 GLLGIFDND 455
>TC17134 weakly similar to UP|LEC_BOWMI (P42088) Lectin (Agglutinin) (BMA),
partial (16%)
Length = 741
Score = 45.8 bits (107), Expect = 1e-05
Identities = 55/209 (26%), Positives = 85/209 (40%), Gaps = 28/209 (13%)
Frame = -2
Query: 91 PTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVAD 150
P + DV+ +L + + G LQ+ Q + H +G + + L D T A
Sbjct: 611 PRLEHDVK-LLGSAKLSDQKGALQIPHESQETDLK-HLAGRGLYSFPIRLLDPSTKTPAS 438
Query: 151 FRTEFTFVVNHSQPH----------------------GDGFTFFLASLDFEFPEGGGGGG 188
F T F F +++S G G TF + +F G G
Sbjct: 437 FETTFAFQLHNSTTSVSGGGSGGGGVGGAAAADVGGGGSGLTFMIVQDEFTV---GRSGP 267
Query: 189 FLGLFDE--ETAYNTSKNSIVAVEFDS-FSNSF-DPNFPHIGIDINTIRSAVTIPWSV-- 242
+LG+ ++ E AY VAVEFD+ S F DPN H+G+++ +I S I S
Sbjct: 266 WLGMLNDACENAYKA-----VAVEFDTRMSPEFGDPNDNHVGVNLGSIISTKIINVSEFG 102
Query: 243 NDQPQGAIQNAKISYSSVSRDLTVVVHYP 271
G + +A ISY R + + + P
Sbjct: 101 VSLKDGFVHHAWISYDGPQRRMEIHLGLP 15
>TC18976 similar to UP|LCB2_ROBPS (Q42372) Bark agglutinin I, polypeptide B
precursor (RPbAI) (LECRPA2), partial (14%)
Length = 617
Score = 42.7 bits (99), Expect = 8e-05
Identities = 20/40 (50%), Positives = 29/40 (72%), Gaps = 3/40 (7%)
Frame = +1
Query: 286 IDIESYLSEWVVIGFSATTG---EFAETHSILSWSFNSTL 322
+D+++ L E+V +GFSATTG + ET+ +LSWSF S L
Sbjct: 7 VDLKNVLPEYVRVGFSATTGLNPDHVETNDVLSWSFESDL 126
>AV765853
Length = 434
Score = 38.1 bits (87), Expect = 0.002
Identities = 15/36 (41%), Positives = 23/36 (63%)
Frame = +1
Query: 282 IKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWS 317
+ Y +D+ L E + +GFSA+TG A +H +L WS
Sbjct: 43 LSYALDLSPILQESMYVGFSASTGLLASSHYVLGWS 150
>BP035176
Length = 444
Score = 36.2 bits (82), Expect = 0.008
Identities = 14/36 (38%), Positives = 25/36 (68%)
Frame = -3
Query: 286 IDIESYLSEWVVIGFSATTGEFAETHSILSWSFNST 321
+D+ YL++++ +GFS +T ETH + WSF+S+
Sbjct: 355 LDMGQYLNDFMYVGFSGSTQGSTETHRVEWWSFSSS 248
>BP029811
Length = 481
Score = 29.6 bits (65), Expect = 0.72
Identities = 19/52 (36%), Positives = 27/52 (51%)
Frame = -3
Query: 101 LLNGDANTTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFR 152
LL D ++TNGT K + G SP SSG +G HL+ + +V D +
Sbjct: 347 LLVTDLDSTNGTFIDDKRLRPGVISPVSSGSLVTFGDTHLAMFRVSKVGDVK 192
>TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%)
Length = 1036
Score = 29.3 bits (64), Expect = 0.94
Identities = 20/74 (27%), Positives = 37/74 (49%), Gaps = 2/74 (2%)
Frame = +2
Query: 42 LPYSKHHQ-YSQLNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSF-PTFDPDVRV 99
L Y HH +S + + + N+TS + F L + T +++FS+ PT +
Sbjct: 269 LLYRPHHPTFSVTSLKLSYLNLTSSSTLSSKFDLTVAATNPNKKNIAFSYLPT-----SI 433
Query: 100 ILLNGDANTTNGTL 113
+L+GD + +GT+
Sbjct: 434 TILSGDVDVGDGTI 475
>TC15548 similar to UP|AAR87327 (AAR87327) Expressed protein, partial (12%)
Length = 758
Score = 27.7 bits (60), Expect = 2.7
Identities = 12/28 (42%), Positives = 17/28 (59%)
Frame = +1
Query: 190 LGLFDEETAYNTSKNSIVAVEFDSFSNS 217
+G DEET Y+TS+ + V DS+ S
Sbjct: 463 MGRIDEETPYDTSEGGVAFVAPDSYPRS 546
>AV427652
Length = 338
Score = 27.7 bits (60), Expect = 2.7
Identities = 20/58 (34%), Positives = 26/58 (44%), Gaps = 7/58 (12%)
Frame = +3
Query: 46 KHHQYSQ-------LNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPD 96
+HHQ Q +NYQP+ I SE L T + L S SLS + DP+
Sbjct: 33 EHHQQPQPNIDPFLVNYQPSELRIASEFLNTWLPFLTKDLCRRCSQSLSDRIRSIDPE 206
>TC18230 similar to UP|Q945U3 (Q945U3) Acyl-CoA oxidase , partial (53%)
Length = 1222
Score = 27.7 bits (60), Expect = 2.7
Identities = 18/63 (28%), Positives = 28/63 (43%), Gaps = 1/63 (1%)
Frame = +1
Query: 146 GRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKN 204
G+++ + ++ Q G GF L SLD P G G +G+ AYNT N
Sbjct: 436 GKISTHAVVYARLITDGQEQGVHGFIVQLRSLDDHLPLPGITVGDIGMKFGSGAYNTMDN 615
Query: 205 SIV 207
++
Sbjct: 616 GVL 624
>TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenylated Rab
receptor 2 {Arabidopsis thaliana;}, partial (78%)
Length = 1012
Score = 27.3 bits (59), Expect = 3.6
Identities = 23/91 (25%), Positives = 41/91 (44%)
Frame = +3
Query: 25 LFSSTCFKIQDTIYSSILPYSKHHQYSQLNYQPTMANITSELLATLIFSLLLITTTVKSD 84
L + F ++ + L K+ Y ++NY ++ I + L T FSL+ +T + S
Sbjct: 228 LTDRSAFSKPESFSDATLRVRKNFSYFRINYYAVVSLILAVSLLTNPFSLITLTGLLASW 407
Query: 85 SLSFSFPTFDPDVRVILLNGDANTTNGTLQL 115
+ + F P R ++L G T TL +
Sbjct: 408 TFLY---LFRPADRPLVLFGRTFTDFETLAI 491
>BP053203
Length = 530
Score = 27.3 bits (59), Expect = 3.6
Identities = 10/33 (30%), Positives = 18/33 (54%)
Frame = +1
Query: 7 HVLLLVIYEVNSEPCLTCLFSSTCFKIQDTIYS 39
H ++ + PC + +S+TCF+I I+S
Sbjct: 265 HSPFIIYFIFTMGPCFSLSYSNTCFRILSQIFS 363
>TC17826 similar to UP|AAR91698 (AAR91698) DNA-binding protein, partial
(14%)
Length = 613
Score = 27.3 bits (59), Expect = 3.6
Identities = 11/27 (40%), Positives = 15/27 (54%)
Frame = -1
Query: 195 EETAYNTSKNSIVAVEFDSFSNSFDPN 221
E YN + N + ++FDS SN PN
Sbjct: 388 ESNTYNNTTNLKIILQFDSESNKIHPN 308
>TC13742 similar to UP|Q9ZR53 (Q9ZR53) Annexin-like protein, partial (53%)
Length = 612
Score = 26.6 bits (57), Expect = 6.1
Identities = 9/22 (40%), Positives = 13/22 (58%)
Frame = +1
Query: 24 CLFSSTCFKIQDTIYSSILPYS 45
C+F C IQ+ +Y+ PYS
Sbjct: 40 CIFLLVCVLIQENVYAQCAPYS 105
>AV410186
Length = 433
Score = 26.6 bits (57), Expect = 6.1
Identities = 14/62 (22%), Positives = 31/62 (49%), Gaps = 11/62 (17%)
Frame = +2
Query: 241 SVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAI-----------KYPIDIE 289
S + + + +++NA S+ + R ++ H+P NTT++ I +PI ++
Sbjct: 83 SSSPETEPSLRNAH*SHLHLPRTFPLISHHPSAPNTTHLATITESSSFLCPLVAFPIKLQ 262
Query: 290 SY 291
S+
Sbjct: 263 SF 268
>TC20179
Length = 511
Score = 26.6 bits (57), Expect = 6.1
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = -3
Query: 251 QNAKISYSSVSRDLTVVVHYPPGQNTTN 278
+ +K S ++S+ L VH+ PG N+TN
Sbjct: 215 KTSKSSGEALSKLLNSTVHFSPGSNSTN 132
>TC15458
Length = 525
Score = 26.2 bits (56), Expect = 7.9
Identities = 14/48 (29%), Positives = 23/48 (47%)
Frame = -1
Query: 27 SSTCFKIQDTIYSSILPYSKHHQYSQLNYQPTMANITSELLATLIFSL 74
+S C + + +Y PY++ + Q NY+ T E+LA I L
Sbjct: 372 TSGCSRNKWMLYKESKPYNQQLPF*QYNYESFTKK*TKEILANAILVL 229
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,917,016
Number of Sequences: 28460
Number of extensions: 81500
Number of successful extensions: 635
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 611
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 623
length of query: 322
length of database: 4,897,600
effective HSP length: 90
effective length of query: 232
effective length of database: 2,336,200
effective search space: 541998400
effective search space used: 541998400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0541.15


