

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.11
(269 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
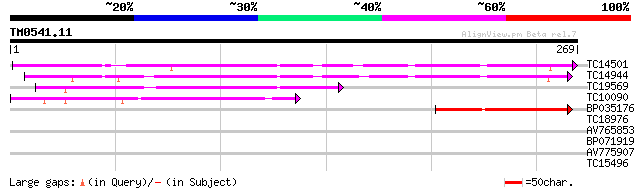
Score E
Sequences producing significant alignments: (bits) Value
TC14501 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA),... 140 2e-34
TC14944 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA),... 137 3e-33
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 91 2e-19
TC10090 weakly similar to UP|Q9ZWP6 (Q9ZWP6) Lectin, partial (43%) 79 1e-15
BP035176 43 6e-05
TC18976 similar to UP|LCB2_ROBPS (Q42372) Bark agglutinin I, pol... 38 0.002
AV765853 37 0.005
BP071919 33 0.051
AV775907 30 0.33
TC15496 homologue to UP|Q9LKL0 (Q9LKL0) 14-3-3 protein, partial ... 27 2.8
>TC14501 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA), partial
(86%)
Length = 1066
Score = 140 bits (354), Expect = 2e-34
Identities = 98/273 (35%), Positives = 142/273 (51%), Gaps = 5/273 (1%)
Frame = +1
Query: 2 ANSFSISLLTTLLPFILLITTVKSDSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKD 61
+N+ + + L L F+L I + + SFNF F D I+ GDA ++
Sbjct: 58 SNASTQNFLLACLFFLLAINVNLAAAISFNFTEFTDDGSLIIQ-GDA-------KIWADG 213
Query: 62 LYGIPSKNSFGQTT----FFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYI 117
+P+ S G TT + V + D+ TG VA F T FSF++ DG F++
Sbjct: 214 SLALPTDPSVGFTTSRALYATPVPIWDSTTGNVASFVTSFSFIIKDFEDYNPADGLVFFL 393
Query: 118 AGLDFDFPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDI 177
A + P++ GG G+ + K+AFN Q+VAVEFD+F N WD S+ H+GID+
Sbjct: 394 APFGTEIPKE-STGGRFGIINGKDAFN----QIVAVEFDTFINPWD----SSPRHIGIDV 546
Query: 178 NSIRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGA 237
NS+ S+ T PW G + K I Y S +K LSV V++ N + + ++ E+DL
Sbjct: 547 NSLISLKTVPWN---KVAGSLEKVTIIYDSQTKTLSVLVIHENGQI---STISQEIDLKV 708
Query: 238 VLSDWVLVGFTAATGSVS-ETHDILSWAFNSFL 269
VL + V VGF+A T S E HDI SW+F S L
Sbjct: 709 VLPEEVSVGFSATTTSGGRERHDIYSWSFTSTL 807
>TC14944 similar to UP|LEC_LOTTE (P19664) Anti-H(O) lectin (LTA), partial
(85%)
Length = 1155
Score = 137 bits (344), Expect = 3e-33
Identities = 102/265 (38%), Positives = 137/265 (51%), Gaps = 5/265 (1%)
Frame = +2
Query: 8 SLLTTLLPFILLITTVKSDSF---SFNFPRFESDTKTILSDGDANT-TGGVLQLTKKDLY 63
S L +L +LL+ T+K +S SFNF +F D IL GDA T G L L L
Sbjct: 59 STLKVVLASLLLLLTIKVNSVEGVSFNFTKFTDDGSLILQ-GDAKIWTDGRLALPSDPLI 235
Query: 64 GIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFD 123
G + + + V + D+ TG VA F + FSFVV+ DG F++A
Sbjct: 236 G----KTTSRVLYATPVPIWDSTTGSVASFVSSFSFVVSDVPGYKISDGLVFFLAPWGTT 403
Query: 124 FPEDPKEGGFLGLFDPKNAFNTSANQVVAVEFDSFTNVWDPSYPSNSPHVGIDINSIRSV 183
P + EG LG+ D KN +N Q VAVEFDS+ N DP++ H+GID+NS+ S+
Sbjct: 404 IPPN-SEGKNLGILDEKNGYN----QFVAVEFDSYNNTRDPTF----RHIGIDVNSVMSM 556
Query: 184 ATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWV 243
W G + KA+I Y S +K LSV V + N + T VA ++DL VL V
Sbjct: 557 NLVKWN---RVSGALEKAIIIYDSPTKTLSVVVTHQNGQI---TTVAQQIDLKVVLPKEV 718
Query: 244 LVGFTAATGSV-SETHDILSWAFNS 267
VGF+A T + E HDI SW+F S
Sbjct: 719 SVGFSATTWNTHRERHDIHSWSFTS 793
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 90.9 bits (224), Expect = 2e-19
Identities = 55/150 (36%), Positives = 75/150 (49%), Gaps = 4/150 (2%)
Frame = +3
Query: 13 LLPFILLITTVKS----DSFSFNFPRFESDTKTILSDGDANTTGGVLQLTKKDLYGIPSK 68
LL +LL+TT + S SFN F+ DT I +GD +T G + L K +
Sbjct: 48 LLLLLLLLTTTTTFPTTHSLSFNITNFDDDTTAIAYEGDGKSTNGSIDLNKVSYFF---- 215
Query: 69 NSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIAGLDFDFPEDP 128
G+ +HL D T + DFTT F+F ++ +GDGF FY+A L + P +
Sbjct: 216 -RVGRAISAQPLHLYDPTTKTLTDFTTRFTFTISAINKTSYGDGFAFYLAPLGYQIPPN- 389
Query: 129 KEGGFLGLFDPKNAFNTSANQVVAVEFDSF 158
GG GLF+ N N VVAVEFD+F
Sbjct: 390 SAGGTFGLFNATTNNNLPQNHVVAVEFDTF 479
>TC10090 weakly similar to UP|Q9ZWP6 (Q9ZWP6) Lectin, partial (43%)
Length = 461
Score = 78.6 bits (192), Expect = 1e-15
Identities = 54/142 (38%), Positives = 75/142 (52%), Gaps = 4/142 (2%)
Frame = +3
Query: 1 MANSFSISLLTTLLP--FILLITTVKS-DSFSFNFPRFESDTKTILSDGDANTTG-GVLQ 56
MA+ S LT+LL F+LL+ V S D SF F +F +L GDA G LQ
Sbjct: 36 MASFQSQKSLTSLLTLFFLLLVNKVNSTDYLSFTFNQFTPQQPDLLFQGDALVAPTGKLQ 215
Query: 57 LTKKDLYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFY 116
LTK + +P S G+ + VH+ D++TG VA F T FSF+++ DG F+
Sbjct: 216 LTKVE-NDLPVYKSTGRALYVAPVHIWDSKTGNVASFITSFSFIIDSPNVNKIADGLAFF 392
Query: 117 IAGLDFDFPEDPKEGGFLGLFD 138
+A +D + K GG LG+FD
Sbjct: 393 LAPVD---TQPQKPGGLLGIFD 449
>BP035176
Length = 444
Score = 42.7 bits (99), Expect = 6e-05
Identities = 22/65 (33%), Positives = 40/65 (60%)
Frame = -3
Query: 203 ISYRSASKVLSVFVVYPNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHDILS 262
I + + + L+V+V Y N K E ++ +D+G L+D++ VGF+ +T +ETH +
Sbjct: 442 IEFDGSIQGLNVWVSYSNLKPK-EPVLSMNLDMGQYLNDFMYVGFSGSTQGSTETHRVEW 266
Query: 263 WAFNS 267
W+F+S
Sbjct: 265 WSFSS 251
>TC18976 similar to UP|LCB2_ROBPS (Q42372) Bark agglutinin I, polypeptide B
precursor (RPbAI) (LECRPA2), partial (14%)
Length = 617
Score = 38.1 bits (87), Expect = 0.002
Identities = 21/40 (52%), Positives = 27/40 (67%), Gaps = 3/40 (7%)
Frame = +1
Query: 233 VDLGAVLSDWVLVGFTAATG---SVSETHDILSWAFNSFL 269
VDL VL ++V VGF+A TG ET+D+LSW+F S L
Sbjct: 7 VDLKNVLPEYVRVGFSATTGLNPDHVETNDVLSWSFESDL 126
>AV765853
Length = 434
Score = 36.6 bits (83), Expect = 0.005
Identities = 15/46 (32%), Positives = 29/46 (62%)
Frame = +1
Query: 219 PNSPVKIETFVAYEVDLGAVLSDWVLVGFTAATGSVSETHDILSWA 264
P P+ ++Y +DL +L + + VGF+A+TG ++ +H +L W+
Sbjct: 28 PKKPI-----LSYALDLSPILQESMYVGFSASTGLLASSHYVLGWS 150
>BP071919
Length = 487
Score = 33.1 bits (74), Expect = 0.051
Identities = 30/117 (25%), Positives = 51/117 (42%), Gaps = 7/117 (5%)
Frame = +1
Query: 9 LLTTLLPFILLITTVKS------DSFSFNFPRFESDTKTILSDGDANT-TGGVLQLTKKD 61
L+ +LL VKS D F F + + + +G A+ + G+L+LT D
Sbjct: 127 LVLVCFAILLLSIRVKSRMHDDDDDEELFFDGFSAASSNMSVNGCASIESNGLLRLTN-D 303
Query: 62 LYGIPSKNSFGQTTFFGAVHLSDNRTGKVADFTTEFSFVVNPKGSEPHGDGFTFYIA 118
G+ G + + L ++ TGK F+T F+F + +P GF F ++
Sbjct: 304 TQGL-----MGDAFYSTPIKLKNSTTGKAFSFSTAFAFAICNGYGKPPYHGFAFSVS 459
>AV775907
Length = 329
Score = 30.4 bits (67), Expect = 0.33
Identities = 13/49 (26%), Positives = 25/49 (50%)
Frame = +2
Query: 187 PWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSPVKIETFVAYEVDL 235
PWP+ ++P +G++ I+YR + V IE F+ ++V +
Sbjct: 104 PWPLKVIPSPXVGESYINYR--------LTIQHELMVHIEAFIQHQVTI 226
>TC15496 homologue to UP|Q9LKL0 (Q9LKL0) 14-3-3 protein, partial (48%)
Length = 677
Score = 27.3 bits (59), Expect = 2.8
Identities = 18/59 (30%), Positives = 31/59 (52%)
Frame = -2
Query: 164 PSYPSNSPHVGIDINSIRSVATAPWPIDLVPQGEIGKALISYRSASKVLSVFVVYPNSP 222
P Y S SP G+D++ I S A++ ++ K +S RS +++V +Y +SP
Sbjct: 385 PHYCSPSPDCGLDVSLISSPASS-------VMSDVHKVRLSRRSC--MINVLSLYDSSP 236
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,174,839
Number of Sequences: 28460
Number of extensions: 49728
Number of successful extensions: 242
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 233
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 234
length of query: 269
length of database: 4,897,600
effective HSP length: 89
effective length of query: 180
effective length of database: 2,364,660
effective search space: 425638800
effective search space used: 425638800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0541.11


