

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
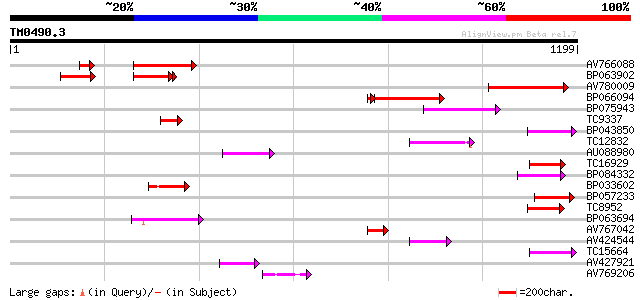
Query= TM0490.3
(1199 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV766088 206 1e-53
BP063902 170 3e-43
AV780009 166 2e-41
BP066094 121 4e-28
BP075943 100 2e-21
TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B pre... 92 6e-19
BP043850 87 1e-17
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 85 6e-17
AU088980 84 2e-16
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 83 3e-16
BP084332 79 6e-15
BP033602 77 1e-14
BP057233 76 3e-14
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 73 2e-13
BP063694 72 7e-13
AV767042 70 2e-12
AV424544 68 7e-12
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 67 1e-11
AV427921 65 8e-11
AV769206 59 5e-09
>AV766088
Length = 501
Score = 206 bits (525), Expect = 1e-53
Identities = 96/132 (72%), Positives = 112/132 (84%)
Frame = -3
Query: 263 KTGMPLLRCNSLGDLYPVTRSSPFAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRS 322
+TG+PL+RCNS GDLYPVT S PFAG+A S+WH+RLGHP+SSAL +LR+NK I E
Sbjct: 397 QTGIPLMRCNSPGDLYPVTLSFPFAGIAQSLWHSRLGHPSSSALRYLRSNKFISYELLNY 218
Query: 323 SSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFL 382
S VC+SCV GKHVRLPF SS +T+ PFDILHSDLWTSPVLS+AGHR+YVLFLDD TDFL
Sbjct: 217 SPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTDFL 38
Query: 383 WTFPISKKSQVY 394
WTFP+S KSQV+
Sbjct: 37 WTFPLSNKSQVF 2
Score = 55.8 bits (133), Expect = 4e-08
Identities = 25/33 (75%), Positives = 28/33 (84%)
Frame = -2
Query: 147 PTDIAAAMHTMSLTPPDTPWYMDTGASSHTAAS 179
PTD+A AMHTM+L PPD+ YMDTGASSH AAS
Sbjct: 497 PTDLATAMHTMTLAPPDST*YMDTGASSHVAAS 399
>BP063902
Length = 494
Score = 170 bits (430), Expect(2) = 3e-43
Identities = 80/83 (96%), Positives = 83/83 (99%)
Frame = -3
Query: 263 KTGMPLLRCNSLGDLYPVTRSSPFAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRS 322
KTGMPLLRCNSLGDLYPVTRSSPFAGLASSVWH+RLGHPASSALNHLRNNKLIFCEPSRS
Sbjct: 273 KTGMPLLRCNSLGDLYPVTRSSPFAGLASSVWHHRLGHPASSALNHLRNNKLIFCEPSRS 94
Query: 323 SSVCDSCVLGKHVRLPFSSSAII 345
SSVCDSCVLGKHVRLPFSSSA++
Sbjct: 93 SSVCDSCVLGKHVRLPFSSSALL 25
Score = 152 bits (383), Expect = 4e-37
Identities = 70/74 (94%), Positives = 70/74 (94%)
Frame = -2
Query: 107 WGMPPCPYPTSQWSRPMGVPQQPGVLGPRPQAYTATASPAPTDIAAAMHTMSLTPPDTPW 166
W PCPYPTSQ SRPMGVPQQPGVLGPRPQAYTATASPAPTDIAAAMHTMSLTPPDTPW
Sbjct: 493 WACLPCPYPTSQXSRPMGVPQQPGVLGPRPQAYTATASPAPTDIAAAMHTMSLTPPDTPW 314
Query: 167 YMDTGASSHTAASQ 180
YMDTGASSHTAASQ
Sbjct: 313 YMDTGASSHTAASQ 272
Score = 23.5 bits (49), Expect(2) = 3e-43
Identities = 10/10 (100%), Positives = 10/10 (100%)
Frame = -2
Query: 344 IITLRPFDIL 353
IITLRPFDIL
Sbjct: 31 IITLRPFDIL 2
>AV780009
Length = 529
Score = 166 bits (420), Expect = 2e-41
Identities = 82/169 (48%), Positives = 108/169 (63%)
Frame = -1
Query: 1012 ALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWS 1071
A RVLRYV+G GL L +Y+D+DW GCPDTRRS +GY +FLG +LISW
Sbjct: 526 AATRVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISWR 347
Query: 1072 SKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNP 1131
+K+Q T+SRSS+EAEYR +A V E W+ L L + + CDN SA++++ NP
Sbjct: 346 TKKQTTVSRSSSEAEYRALAATVCEVQWLSYLFQFLKLNVPLPVPLFCDNQSALHIAHNP 167
Query: 1132 VHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLF 1180
H+RTKHIE+D H VR K+ G +L + + HQ+ADIFTK P L F
Sbjct: 166 TFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTKPCPHLCF 20
>BP066094
Length = 532
Score = 121 bits (304), Expect(2) = 4e-28
Identities = 59/147 (40%), Positives = 93/147 (63%)
Frame = +2
Query: 772 KPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKS 831
K IR ++S +++ + +HQ++V++AFL+G + E +Y+H P G D + D++ +LKKS
Sbjct: 89 KTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKKS 268
Query: 832 LYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHD 891
LYGLKQAPRAWY+R + ++ D +LF DI + +YVDDII +++
Sbjct: 269 LYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANPS 448
Query: 892 LRKSFMALLASEFAMKDLGPLSYFLGI 918
L K F ++ +EF M+ +G L YFLGI
Sbjct: 449 LCKEFSEMMQAEFEMRMMGELKYFLGI 529
Score = 21.6 bits (44), Expect(2) = 4e-28
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +3
Query: 756 SQIAGVDCDETFSPVVK 772
SQ G+D ETF+PV +
Sbjct: 42 SQQ*GIDYTETFAPVAR 92
>BP075943
Length = 547
Score = 100 bits (248), Expect = 2e-21
Identities = 60/164 (36%), Positives = 88/164 (53%), Gaps = 1/164 (0%)
Frame = -3
Query: 876 ILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYA 935
+LLYVDD++L + + + L +F +KDLG YFLG+ + R G+ L+Q YA
Sbjct: 521 LLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQRKYA 342
Query: 936 TEIIARAGMASCNPSATPVDT-KQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYA 994
++I+ +G P T Q L +++GTP D YR + G L YL TRPDI++A
Sbjct: 341 LQLISDSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTTRPDITFA 162
Query: 995 VQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETL 1038
V Q+ + AP H L L+Y+ G+ GL YP+ TL
Sbjct: 161 VNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGL-FYPASSSTL 33
>TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B precursor
(Pectin methylesterase) (PE) , partial (54%)
Length = 1145
Score = 91.7 bits (226), Expect = 6e-19
Identities = 43/46 (93%), Positives = 44/46 (95%)
Frame = +3
Query: 319 PSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLS 364
PSRS+SVCDSCVLGKHVRLPFSSS ITLRPFDILHSDLWTSPVLS
Sbjct: 1008 PSRSNSVCDSCVLGKHVRLPFSSSETITLRPFDILHSDLWTSPVLS 1145
>BP043850
Length = 515
Score = 87.4 bits (215), Expect = 1e-17
Identities = 50/104 (48%), Positives = 62/104 (59%)
Frame = -1
Query: 1095 SESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRG 1154
SE W+R LL EL F SQ T +H DN SAI ++ NPV+H+ T+HIE+D H VRE R
Sbjct: 515 SEIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRR 336
Query: 1155 QARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPPASTAG 1198
+ HV + QIADI TK L R + S L + E PAS G
Sbjct: 335 VITLPHVSTSVQIADILTKSLTRQRHNFLVSKLMLVESPASI*G 204
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 85.1 bits (209), Expect = 6e-17
Identities = 53/148 (35%), Positives = 78/148 (51%), Gaps = 10/148 (6%)
Frame = +2
Query: 846 FADYVSSIGFRHSTSDHSLFIFR-QGSDIAYILLYVDDIILVASSHDLRKSFMALLASEF 904
F ++ S+G+ +SDH + R +D +LLYVDD+++V + D + A LA EF
Sbjct: 2 FDSFIMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREF 181
Query: 905 AMKDLGPLSYFLGIAVTRHAGG--LFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSS 962
MKDLGP + LG+ + R ++LSQ Y +++ R M CNP +TP+ KLSS
Sbjct: 182 DMKDLGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSS 361
Query: 963 SSGTPCEDASL-------YRSLDGALQY 983
S P +A Y S G+L Y
Sbjct: 362 SM-IPSSEAERMEMSRVPYASAVGSLMY 442
>AU088980
Length = 360
Score = 83.6 bits (205), Expect = 2e-16
Identities = 44/110 (40%), Positives = 59/110 (53%)
Frame = +2
Query: 450 NGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHR 509
N ERK R I N+ R L+ HS +P + W A++ A YL+N LP L+ P Q L +
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 510 DPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDL 559
P+ L+VFG LC+ + KL PR+ CVFLG+ GY DL
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMDL 331
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 82.8 bits (203), Expect = 3e-16
Identities = 37/77 (48%), Positives = 52/77 (67%)
Frame = +1
Query: 1099 WIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARV 1158
W+ LL +L P Q LV+CDN SA +++ NPV H+RTKHIE+D H VRE++ +G +
Sbjct: 4 WLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIHL 183
Query: 1159 LHVPSRHQIADIFTKGL 1175
L + S +ADI+TK L
Sbjct: 184 LPISSSEPLADIYTKAL 234
>BP084332
Length = 368
Score = 78.6 bits (192), Expect = 6e-15
Identities = 42/101 (41%), Positives = 61/101 (59%)
Frame = +2
Query: 1075 QPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHH 1134
Q T++ S+AEAEY A ++ W+++ L + L ++CDN +AI LS NP+ H
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILH 208
Query: 1135 QRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGL 1175
R KHIE+ HF+R+ V +G + V + HQ ADIFTK L
Sbjct: 209 SRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPL 331
>BP033602
Length = 533
Score = 77.4 bits (189), Expect = 1e-14
Identities = 39/88 (44%), Positives = 54/88 (61%), Gaps = 1/88 (1%)
Frame = +3
Query: 293 VWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDI 352
+WHNRLGHP L HL + +F + SS C+SCVL K+ R P+ S RPF +
Sbjct: 276 LWHNRLGHPNFQYLRHLFPD--LFKNVNCSSLECESCVLAKNQRAPYYSQPYHASRPFYL 449
Query: 353 LHSDLW-TSPVLSTAGHRYYVLFLDDHT 379
+HSD+W S + + G R++V F+DDHT
Sbjct: 450 IHSDVWGPSKITTQFGKRWFVTFIDDHT 533
>BP057233
Length = 473
Score = 76.3 bits (186), Expect = 3e-14
Identities = 35/84 (41%), Positives = 53/84 (62%)
Frame = -2
Query: 1110 PLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIAD 1169
P+ + ++ CDN+SA L+ NPV H R+KHIE+D+H++R++V++ + V +VP+ QIAD
Sbjct: 466 PMPRKPILWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQIAD 287
Query: 1170 IFTKGLPRLLFDDFRSSLSVGEPP 1193
TK L F R L V P
Sbjct: 286 CLTKPLSHTRFSQLRDKLGVIHSP 215
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 73.2 bits (178), Expect = 2e-13
Identities = 34/78 (43%), Positives = 51/78 (64%)
Frame = +3
Query: 1096 ESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQ 1155
E+ W+ L +L +++CDN SA++L+ N V H+RT++IE+D H V KV+ G
Sbjct: 21 EALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFGI 200
Query: 1156 ARVLHVPSRHQIADIFTK 1173
+LHVPS Q+AD+FTK
Sbjct: 201 LHLLHVPSSDQVADVFTK 254
>BP063694
Length = 511
Score = 71.6 bits (174), Expect = 7e-13
Identities = 47/163 (28%), Positives = 72/163 (43%), Gaps = 10/163 (6%)
Frame = -2
Query: 257 FSVSDFKTGMPLLRCNSLGDLYPV--------TRSSPFAGLASSVWHNRLGHPASSALNH 308
F + + KTG L R LY + T SSP + +WH+RLGH +
Sbjct: 492 FVIQNRKTGAVLGRXRCDQGLYVLNQDSQALLTTSSPLPRASFELWHSRLGHVNFDIIKQ 313
Query: 309 LRNNKLIFCEPSRSSSVC-DSCVLGKHVRLPFSSSAIITLRPFDILHSDL-WTSPVLSTA 366
L + + +C SC + K RL F + D++H DL SPV S
Sbjct: 312 LHKHGCLDVSSILPKPICCTSCQMAKSKRLVFHDNNKRASAVLDLIHCDLRGPSPVASID 133
Query: 367 GHRYYVLFLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQFS 409
G Y+V+F+DD + F W +P+ +KS + ++ +FS
Sbjct: 132 GFSYFVIFVDDFSRFTWFYPLKRKSDFSDVLLRFKVFMENRFS 4
>AV767042
Length = 444
Score = 70.5 bits (171), Expect = 2e-12
Identities = 33/44 (75%), Positives = 39/44 (88%)
Frame = -3
Query: 757 QIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFL 800
Q AGVDC++TFSPVVKPA IRTV +IALSRSWPI QL+VQN ++
Sbjct: 349 QTAGVDCNKTFSPVVKPAPIRTVFTIALSRSWPIPQLNVQNLWI 218
>AV424544
Length = 276
Score = 68.2 bits (165), Expect = 7e-12
Identities = 36/90 (40%), Positives = 52/90 (57%)
Frame = +3
Query: 845 RFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEF 904
+ + Y+ +G+ S DHSLF + + IL+YVDD+IL + + + L +F
Sbjct: 6 KLSSYLHILGYIQSAHDHSLFTKFRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQF 185
Query: 905 AMKDLGPLSYFLGIAVTRHAGGLFLSQSTY 934
+KDLG L YFLG+ V R + GLFLSQ Y
Sbjct: 186 RIKDLGTLKYFLGLEVARSSCGLFLSQRKY 275
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 67.4 bits (163), Expect = 1e-11
Identities = 37/100 (37%), Positives = 55/100 (55%)
Frame = +1
Query: 1099 WIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARV 1158
W+ +L +L P +L++CD+ SA +++ N V H+RTKH+++D H VREK+ +
Sbjct: 13 WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAKLFHL 192
Query: 1159 LHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPPASTAG 1198
L + S Q ADI TK L F S L V +S G
Sbjct: 193 LPISSVDQTADILTKPLESGPFSHLVSKLGVLNIYSSACG 312
>AV427921
Length = 284
Score = 64.7 bits (156), Expect = 8e-11
Identities = 33/84 (39%), Positives = 47/84 (55%)
Frame = +1
Query: 445 HTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQ 504
+T QNG +ERK RTI NM+R+LL S +P SF A+ + ++LN P +QN P +
Sbjct: 19 YTPQQNGVSERKNRTILNMVRSLLTMSGLPKSFLPEAVMWSLHILNRSPTLVVQNMMPEE 198
Query: 505 LLYHRDPSYSHLRVFGCLCFPLFP 528
R P+ H R+ CL + P
Sbjct: 199 AWSGRQPAVDHFRISRCLAYAHVP 270
>AV769206
Length = 536
Score = 58.9 bits (141), Expect = 5e-09
Identities = 43/109 (39%), Positives = 53/109 (48%), Gaps = 5/109 (4%)
Frame = -3
Query: 534 KLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETRFPFADL---SLTPAPSY 590
KL S CVF+GY +NH+GYKC D S R I IS+ V+F E RFP+ L T + S
Sbjct: 510 KLSLHSKECVFIGYSINHKGYKCLDQSGR-IYISKDVLFHEHRFPYPSLFPDETTSSTSA 334
Query: 591 ECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSPTDS--SPPLSALVSPTA 637
E F P + H PP SS S S L + +PTA
Sbjct: 333 EFFPLSTIPIVSH---------TTPPALASSGFSSLGSATLPSSFNPTA 214
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.134 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,167,202
Number of Sequences: 28460
Number of extensions: 571145
Number of successful extensions: 11322
Number of sequences better than 10.0: 850
Number of HSP's better than 10.0 without gapping: 5741
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8785
length of query: 1199
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1098
effective length of database: 2,023,140
effective search space: 2221407720
effective search space used: 2221407720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0490.3


