

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
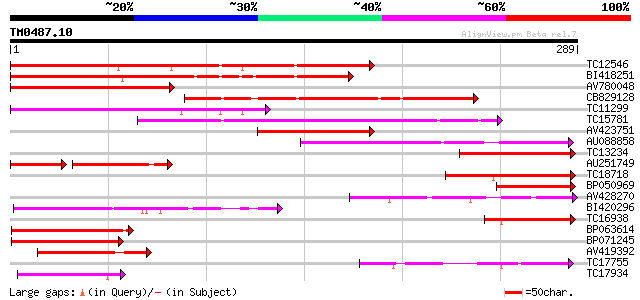
Query= TM0487.10
(289 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat ... 171 1e-43
BI418251 138 1e-33
AV780048 104 2e-23
CB829128 97 2e-21
TC11299 92 8e-20
TC15781 92 1e-19
AV423751 84 4e-17
AU088858 77 3e-15
TC13234 72 1e-13
AU251749 50 2e-13
TC18718 70 3e-13
BP050969 66 8e-12
AV428270 64 2e-11
BI420296 64 3e-11
TC16938 58 2e-09
BP063614 55 1e-08
BP071245 54 2e-08
AV419392 52 9e-08
TC17755 49 1e-06
TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase ... 42 9e-05
>TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat shock
transcription factor HSF30, partial (9%)
Length = 755
Score = 171 bits (433), Expect = 1e-43
Identities = 111/203 (54%), Positives = 135/203 (65%), Gaps = 17/203 (8%)
Frame = +2
Query: 1 MEDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRR------ 54
MED IS+LPD +LC+ILSFL +K+AVATS+LS+RW PLWRSV TL F +AN+
Sbjct: 143 MEDGISTLPDALLCHILSFLTSKEAVATSVLSRRWIPLWRSVPTLHFKDANYHTDIGHAD 322
Query: 55 ---TKGWHSRFVQSVHRVIHTPDFLQPIQK--LRMSFVSRHSEVPSNVNVWVNAAVQRGG 109
K SR VQSV+ VI + DF PI+K LR++ V + P+NV+VWVNA VQR
Sbjct: 323 HDIVKDVRSRHVQSVYAVILSRDFQLPIKKFYLRLNDVCQPFYDPANVSVWVNAVVQR-Q 499
Query: 110 FEHVDICI----YSLSPLNLSSIFSCKTLVVLKLE-GLDLNTTVSSVDLPFLKVLHLQ-V 163
EH+DI + S NLSSIFSC+TLVVLKL GL+L SV P LKVLHLQ
Sbjct: 500 LEHLDISLPYPMLSTPRANLSSIFSCRTLVVLKLRGGLELK-RFPSVHFPCLKVLHLQGA 676
Query: 164 LQLSQRRCLAQLLNGCPVLEDLK 186
L L LA+LL+GCPVLE+LK
Sbjct: 677 LLLHDVPYLAELLSGCPVLENLK 745
>BI418251
Length = 546
Score = 138 bits (348), Expect = 1e-33
Identities = 86/177 (48%), Positives = 112/177 (62%), Gaps = 2/177 (1%)
Frame = +1
Query: 1 MEDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTK--GW 58
M DRIS LPDE+LC ILSFLPT+ AVATS+L KRW LW SV TLDFD+ + + K
Sbjct: 31 MADRISKLPDEVLCQILSFLPTEDAVATSVLCKRWSSLWLSVPTLDFDDYRYLKGKKLKL 210
Query: 59 HSRFVQSVHRVIHTPDFLQPIQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGFEHVDICIY 118
S F+ V+ +I + QPI+ +S +S P +V VW+NAA+QR E++DIC+
Sbjct: 211 QSSFINFVYAIILSRALHQPIKNFTLSVISEECPYP-DVKVWLNAAMQR-QVENLDICL- 381
Query: 119 SLSPLNLSSIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQL 175
SLS L SI SC TLVVLKL + + S VDLP LK LHL+++ + + L L
Sbjct: 382 SLSTLP-CSILSCTTLVVLKLSDVKFH-VFSCVDLPSLKTLHLELVVILNPQSLMDL 546
>AV780048
Length = 501
Score = 104 bits (260), Expect = 2e-23
Identities = 52/84 (61%), Positives = 65/84 (76%)
Frame = +2
Query: 1 MEDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHS 60
MEDR SSLPD ILC+ILS L TK+AV TSILSKRW PLWRSV TLDFD++N+ ++
Sbjct: 245 MEDRFSSLPDPILCHILSLLTTKEAVTTSILSKRWIPLWRSVPTLDFDDSNWYGGDEVYA 424
Query: 61 RFVQSVHRVIHTPDFLQPIQKLRM 84
R V++V++VI DF QPI+K R+
Sbjct: 425 RLVEAVYKVILARDFNQPIKKFRL 496
>CB829128
Length = 463
Score = 97.4 bits (241), Expect = 2e-21
Identities = 66/151 (43%), Positives = 92/151 (60%), Gaps = 1/151 (0%)
Frame = +3
Query: 90 HSEVPSNVNVWVNAAVQRGGFEHVDICIY-SLSPLNLSSIFSCKTLVVLKLEGLDLNTTV 148
H S++ VW+NAA+QR E+++I + S P SI C TLV+LKL+ + +
Sbjct: 3 HDGPDSDLKVWLNAAIQRQ-VENLEIELTDSQMPC---SILRCTTLVILKLDSVTFDA-F 167
Query: 149 SSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVLEDLKAKAIYFKKYKETGFEYKTLSKL 208
+SV LP LK LHL + L + +CL LL GCP+LEDLK + IYF+ G + KTLSKL
Sbjct: 168 TSVYLPSLKTLHLFEVDLVKAQCLIDLLYGCPILEDLKTQ-IYFRDGSFEG-KVKTLSKL 341
Query: 209 VRADISGTSTDLFRMEVINNVDFLRIHEIYI 239
VRAD+ + ++ NV+FLRI E+ I
Sbjct: 342 VRADVFLDGHFIIPVKAFQNVEFLRIEEVCI 434
>TC11299
Length = 498
Score = 92.4 bits (228), Expect = 8e-20
Identities = 58/140 (41%), Positives = 78/140 (55%), Gaps = 7/140 (5%)
Frame = +1
Query: 1 MEDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHS 60
M DRIS+LPDEI+C+ILSFLPT+ AVAT++LSKRW LWRSV+ LDF
Sbjct: 79 MVDRISALPDEIICHILSFLPTENAVATAVLSKRWTHLWRSVSALDFSSVRMYEPDDHRF 258
Query: 61 RFVQSVHRVIHTPDFLQPIQKLRMSF---VSRHSEVPSNVNVWVNAAVQ--RGGFEHVDI 115
F V+ V+ + PI+K + V+ + +N++ WVN Q GG EH+ +
Sbjct: 259 FFSDIVYSVLLFRNAATPIKKFSLELDDDVAVANLDIANIHKWVNFVTQCRVGGIEHLRL 438
Query: 116 CI--YSLSPLNLSSIFSCKT 133
+ P SI SCKT
Sbjct: 439 HFMDFIQLPQLPISILSCKT 498
>TC15781
Length = 621
Score = 91.7 bits (226), Expect = 1e-19
Identities = 70/189 (37%), Positives = 106/189 (56%), Gaps = 3/189 (1%)
Frame = +2
Query: 66 VHRVIHTPDFLQPIQKLRMSFVSRHSEVPSN-VNVWVNAAVQRGGFEHVDICIYSLSPLN 124
V+ I D QPI + + + S S S+ V+ W+N + R E+++ICI LS
Sbjct: 26 VNATILARDERQPITRF*LLYESTVSLEGSDLVHDWLNTVMPRR-IENLEICIPPLSRGI 202
Query: 125 LSSIFSCKTLVVLKLEGLDLNTTVSS-VDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVLE 183
I+SC+TLVVLKL+G L VSS V+LP LK LHL+ + + +L+ GCP+LE
Sbjct: 203 GLKIYSCRTLVVLKLKGYPLGVYVSSHVELPSLKSLHLEKVHFVDFQSFLKLICGCPLLE 382
Query: 184 DLKAKAI-YFKKYKETGFEYKTLSKLVRADISGTSTDLFRMEVINNVDFLRIHEIYIRIP 242
DL+A + Y + F +++L KLV I+ + F +++I+N +FL I E+ I P
Sbjct: 383 DLEAGTVSYLDTSFDQEFRFRSLPKLVSVVINDYNF-RFPLKIISNAEFLSIEEVCI-YP 556
Query: 243 HDDQFTDMF 251
D F +F
Sbjct: 557 EDYVFDFIF 583
>AV423751
Length = 254
Score = 83.6 bits (205), Expect = 4e-17
Identities = 42/60 (70%), Positives = 47/60 (78%)
Frame = +2
Query: 127 SIFSCKTLVVLKLEGLDLNTTVSSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVLEDLK 186
S+FSC TLVVLKLEGL+L SSVDLPFLKVLHLQ L L CLA++L+GC LEDLK
Sbjct: 74 SMFSCTTLVVLKLEGLELKANFSSVDLPFLKVLHLQDLLLQDEGCLAEILSGCLALEDLK 253
>AU088858
Length = 765
Score = 77.4 bits (189), Expect = 3e-15
Identities = 53/140 (37%), Positives = 76/140 (53%), Gaps = 1/140 (0%)
Frame = +3
Query: 149 SSVDLPFLKVLHLQVLQLSQRRCLAQLLNGCPVLEDLKAKAI-YFKKYKETGFEYKTLSK 207
S V LP LK L L L + + + +LL+GCP+LE + + Y + K+L+K
Sbjct: 9 SFVHLPSLKTLQLYCLIFPEPQYVMELLSGCPILEYMHLSRMRYADCSSPSNTNVKSLTK 188
Query: 208 LVRADISGTSTDLFRMEVINNVDFLRIHEIYIRIPHDDQFTDMFHNLTHIELGYYDYNTN 267
LV ADIS ++ ++VI NV+FLRI + IP +F NLTH+EL N
Sbjct: 189 LVSADISFIGFEI-PLKVICNVEFLRIDKYPYEIP-------VFPNLTHLELMMSGIGMN 344
Query: 268 WIEVVEFLNYCPKLQVLVIN 287
W V+ L CPKLQ L+++
Sbjct: 345 WHLVLAMLKKCPKLQKLLLD 404
>TC13234
Length = 609
Score = 71.6 bits (174), Expect = 1e-13
Identities = 34/60 (56%), Positives = 44/60 (72%), Gaps = 1/60 (1%)
Frame = +3
Query: 230 DFLRIHEIYIRI-PHDDQFTDMFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQVLVINQ 288
+FLRI +I R+ P+ + DMFHNLT +EL Y + +W++VVEFL YCPKLQVLVI Q
Sbjct: 3 NFLRIRDIDFRLLPYGGEDIDMFHNLTRVELVYTGHTRDWLDVVEFLKYCPKLQVLVIKQ 182
>AU251749
Length = 362
Score = 49.7 bits (117), Expect(2) = 2e-13
Identities = 25/51 (49%), Positives = 31/51 (60%)
Frame = +1
Query: 33 KRWKPLWRSVTTLDFDEANFRRTKGWHSRFVQSVHRVIHTPDFLQPIQKLR 83
KRWKPLWRSV T D D+ F R + ++R VQ V+ + D PI KLR
Sbjct: 214 KRWKPLWRSVPTRDLDDGRFWRDREAYARLVQGVNALFR--DLHHPIPKLR 360
Score = 41.6 bits (96), Expect(2) = 2e-13
Identities = 21/29 (72%), Positives = 24/29 (82%)
Frame = +2
Query: 1 MEDRISSLPDEILCYILSFLPTKKAVATS 29
M D IS+LPD IL +ILSFLPTK+ VATS
Sbjct: 131 MVDWISALPDTILNFILSFLPTKQVVATS 217
>TC18718
Length = 518
Score = 70.5 bits (171), Expect = 3e-13
Identities = 38/72 (52%), Positives = 46/72 (63%), Gaps = 6/72 (8%)
Frame = +1
Query: 223 MEVINNVDFLRIHEIYIRIPHDD------QFTDMFHNLTHIELGYYDYNTNWIEVVEFLN 276
ME + NV+FLRI+E+ HDD FHNLTHIEL Y ++W+ VVE L
Sbjct: 10 MEAVKNVEFLRIYEMQRLRRHDDGEGAGPYGFPKFHNLTHIELEYNCRISDWLTVVELLK 189
Query: 277 YCPKLQVLVINQ 288
+CPKLQVLVINQ
Sbjct: 190 HCPKLQVLVINQ 225
>BP050969
Length = 557
Score = 65.9 bits (159), Expect = 8e-12
Identities = 27/40 (67%), Positives = 34/40 (84%)
Frame = -3
Query: 249 DMFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQVLVINQ 288
DMFHNLTH+E+ Y ++N +W EVVEFL +CPKLQ LVI+Q
Sbjct: 306 DMFHNLTHVEIVYANFNEDWFEVVEFLTHCPKLQTLVISQ 187
Score = 33.1 bits (74), Expect = 0.057
Identities = 20/37 (54%), Positives = 25/37 (67%)
Frame = -1
Query: 201 EYKTLSKLVRADISGTSTDLFRMEVINNVDFLRIHEI 237
++KTL KLVRA IS + EVINNV FLRI ++
Sbjct: 539 KFKTLPKLVRAHISEGYPIV---EVINNVKFLRIDQV 438
>AV428270
Length = 428
Score = 64.3 bits (155), Expect = 2e-11
Identities = 46/122 (37%), Positives = 66/122 (53%), Gaps = 6/122 (4%)
Frame = +1
Query: 174 QLLNGCPVLEDLKAKAIYF----KKYKETGFEYKTLSKLVRADISGTSTDLFRMEVINNV 229
+ L GCP+LE+LKA+ I+ Y+ L KL+RADI + L ++ NV
Sbjct: 1 EFLAGCPILENLKARRIWITGESSFYRGGELADLGLPKLIRADI--VRSRLPMLQPFYNV 174
Query: 230 DFLR--IHEIYIRIPHDDQFTDMFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQVLVIN 287
FLR I E++ P+ FHNLTH+EL YN+ + +VE L +CP LQ LV
Sbjct: 175 AFLRVDIEEVHYAFPY-------FHNLTHLELVIELYNS--LVLVEMLKHCPNLQSLVFE 327
Query: 288 QV 289
++
Sbjct: 328 RL 333
>BI420296
Length = 592
Score = 63.9 bits (154), Expect = 3e-11
Identities = 51/149 (34%), Positives = 75/149 (50%), Gaps = 12/149 (8%)
Frame = +2
Query: 3 DRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRF 62
DR+S LPD +L ++L F+ TK+AV T +LS RWK LW+ VTTL + ++F T S F
Sbjct: 194 DRLSDLPDLVLLHVLKFMSTKQAVQTCVLSTRWKDLWKGVTTLALNSSDF-ATAPRFSEF 370
Query: 63 VQSV--HR----VIHTPDF-----LQP-IQKLRMSFVSRHSEVPSNVNVWVNAAVQRGGF 110
+ V HR +H D ++P + MS+ H ++ + N ++ G
Sbjct: 371 LSCVLSHRNDSVSLHNLDLRRKGCVEPELLNRVMSYAVSHD--VQSLTIEFNLYLKLG-- 538
Query: 111 EHVDICIYSLSPLNLSSIFSCKTLVVLKL 139
+ L P IFSC+TL LKL
Sbjct: 539 -------FKLHP----CIFSCRTLTYLKL 592
>TC16938
Length = 511
Score = 58.2 bits (139), Expect = 2e-09
Identities = 28/48 (58%), Positives = 34/48 (70%), Gaps = 2/48 (4%)
Frame = +2
Query: 243 HDDQFTD--MFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQVLVINQ 288
HD + D MFHNLTH++L Y D N +W +VVE + CPKLQVLVI Q
Sbjct: 14 HDTEPYDFPMFHNLTHMQLDYIDCNIDWSDVVELIKNCPKLQVLVIYQ 157
>BP063614
Length = 508
Score = 55.5 bits (132), Expect = 1e-08
Identities = 30/62 (48%), Positives = 41/62 (65%)
Frame = +2
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSR 61
+DR+S LPD IL +ILSF+ K AV T ILS RWK LW+ + +L +NF K + ++
Sbjct: 164 KDRLSDLPDCILLHILSFVKAKAAVQTCILSTRWKDLWKRLPSLILLSSNFWTYKSF-TK 340
Query: 62 FV 63
FV
Sbjct: 341 FV 346
>BP071245
Length = 552
Score = 54.3 bits (129), Expect = 2e-08
Identities = 26/57 (45%), Positives = 37/57 (64%)
Frame = +1
Query: 2 EDRISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGW 58
+DR+S LPD +L +ILSF+ K AV T LS RWK LW+ + +L ++F KG+
Sbjct: 379 KDRLSDLPDCVLLHILSFVNAKYAVQTCTLSTRWKDLWKRLPSLILHSSDFSTFKGF 549
>AV419392
Length = 318
Score = 52.4 bits (124), Expect = 9e-08
Identities = 25/58 (43%), Positives = 38/58 (65%)
Frame = +3
Query: 15 YILSFLPTKKAVATSILSKRWKPLWRSVTTLDFDEANFRRTKGWHSRFVQSVHRVIHT 72
+ILS++ TK A+ S+LSKRW+ LWR++ L+FD A+F +S F V V+H+
Sbjct: 3 HILSYVETKDAMQCSVLSKRWRSLWRALPVLNFDSASFTH----YSSFANFVLHVLHS 164
>TC17755
Length = 776
Score = 48.9 bits (115), Expect = 1e-06
Identities = 38/117 (32%), Positives = 55/117 (46%), Gaps = 8/117 (6%)
Frame = +2
Query: 179 CPVLEDLKAKAIYFKK-----YKETGFEYKTLSKLVRADISGTSTDLFRMEVINNVDFLR 233
CP LE + IY+ YK +K+LSKLVRADI D+F
Sbjct: 2 CPNLEYMHLWVIYYDVGCIPCYKN----FKSLSKLVRADIHAIGFDIF------------ 133
Query: 234 IHEIYIRIPHDDQFTD---MFHNLTHIELGYYDYNTNWIEVVEFLNYCPKLQVLVIN 287
+ P D++ D +F NLTH EL ++ +W V+ L +CP LQ +V++
Sbjct: 134 -----VESPEFDKYLDDIPLFPNLTHAEL-MLGFDMSWDLVLAILKHCPMLQNVVLD 286
>TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (11%)
Length = 564
Score = 42.4 bits (98), Expect = 9e-05
Identities = 26/64 (40%), Positives = 36/64 (55%), Gaps = 9/64 (14%)
Frame = -1
Query: 5 ISSLPDEILCYILSFLPTKKAVATSILSKRWKPLWRSVT-TLDFD--------EANFRRT 55
I+ LPD I ILS LP +A TSILS++W+ +W + TL FD + N +R
Sbjct: 564 INQLPDGIPGAILSKLPINEAAKTSILSRKWRYMWTFYSGTLKFDGSPVMKDMKKNHKRV 385
Query: 56 KGWH 59
+G H
Sbjct: 384 RGRH 373
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.139 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,333,164
Number of Sequences: 28460
Number of extensions: 72887
Number of successful extensions: 637
Number of sequences better than 10.0: 70
Number of HSP's better than 10.0 without gapping: 618
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 621
length of query: 289
length of database: 4,897,600
effective HSP length: 89
effective length of query: 200
effective length of database: 2,364,660
effective search space: 472932000
effective search space used: 472932000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0487.10


