

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
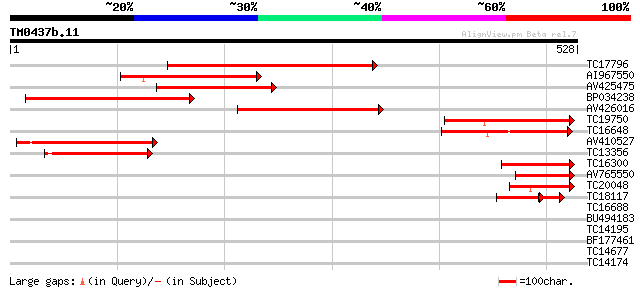
Query= TM0437b.11
(528 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog ... 210 4e-55
AI967550 166 1e-41
AV425475 162 1e-40
BP034238 159 1e-39
AV426016 140 7e-34
TC19750 weakly similar to PIR|E86432|E86432 T5I8.15 protein - Ar... 135 2e-32
TC16648 similar to PIR|T10625|T10625 reticuline oxidase homolog ... 132 1e-31
AV410527 122 2e-28
TC13356 weakly similar to UP|Q9SVG3 (Q9SVG3) Reticuline oxidase-... 100 8e-22
TC16300 similar to UP|Q93Y11 (Q93Y11) Berberine bridge enzyme-li... 98 4e-21
AV765550 80 8e-16
TC20048 weakly similar to PIR|E86432|E86432 T5I8.15 protein - Ar... 74 6e-14
TC18117 weakly similar to UP|Q9AYM8 (Q9AYM8) CPRD2 protein, part... 52 1e-12
TC16688 31 0.57
BU494183 28 3.7
TC14195 UP|Q40215 (Q40215) RAB8A, complete 27 6.3
BF177461 27 6.3
TC14677 similar to PIR|T39903|T39903 serine-rich protein - fissi... 27 6.3
TC14174 UP|PSBC_LOTJA (Q9BBT1) Photosystem II 44 kDa reaction ce... 27 8.3
>TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog F21C20.180
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 594
Score = 210 bits (535), Expect = 4e-55
Identities = 100/196 (51%), Positives = 139/196 (70%), Gaps = 1/196 (0%)
Frame = +1
Query: 148 VQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQ 207
VQ+GATLGELYY I +S VH FPAG CPTVG+GGH SGGG+ + RK+GL+ D+VIDAQ
Sbjct: 7 VQAGATLGELYYGIWQKSKVHGFPAGVCPTVGVGGHISGGGYGNMLRKYGLSVDNVIDAQ 186
Query: 208 IIDVNGKILNRNLMGEDLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTIFDLPKKLDQNV 267
I+DV G+IL+R MGEDLFWAI+GGGG SFGVI S+ VKLV VP VT+F + K LDQN
Sbjct: 187 IVDVQGRILDRKSMGEDLFWAIRGGGGGSFGVILSYTVKLVKVPEIVTVFRVEKTLDQNA 366
Query: 268 SEIFQKWQTIAHKLPGELFLHSVM-GVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSN 326
+++ +WQ +A LF+ ++ +S+ G K+ + ++LG A ++ ++
Sbjct: 367 TDLVLQWQQVAPTTDDRLFMRLLLQPISSKVVKGTKTGRATVIAMFLGGANEVVSILGKE 546
Query: 327 FAELGLRRDDCTEMNW 342
F LGL++++CTE++W
Sbjct: 547 FPVLGLKKENCTELSW 594
>AI967550
Length = 412
Score = 166 bits (419), Expect = 1e-41
Identities = 74/133 (55%), Positives = 102/133 (76%), Gaps = 2/133 (1%)
Frame = +2
Query: 104 LQVRVRSGGHDYEGLSYISN--VPFLIIDLSNFRSISIDIEDESAWVQSGATLGELYYAI 161
+++RVR GGH YEG S +++ F+IID+ N +S+D+E E+AWV+ GATLGE YYAI
Sbjct: 14 VEIRVRGGGHSYEGTSSVADEGTLFVIIDMMNLNHVSVDMETETAWVEGGATLGETYYAI 193
Query: 162 ANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHVIDAQIIDVNGKILNRNLM 221
+ S+VH F AGS PTVG+GGH GGG + RK+GLAAD+V+DA ++D NG++L++ M
Sbjct: 194 SQASSVHGFSAGSSPTVGVGGHIGGGGVGLMSRKYGLAADNVVDALLVDANGQLLDKETM 373
Query: 222 GEDLFWAIKGGGG 234
GED+FWAI+GG G
Sbjct: 374 GEDVFWAIRGGRG 412
>AV425475
Length = 344
Score = 162 bits (410), Expect = 1e-40
Identities = 68/112 (60%), Positives = 93/112 (82%)
Frame = +3
Query: 137 ISIDIEDESAWVQSGATLGELYYAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKH 196
I++DIED SAW+Q+GAT+GE+YY I+ +S VH FPAG C ++G+GGH GG + ++ RK+
Sbjct: 9 INVDIEDNSAWIQAGATIGEVYYRISEKSAVHGFPAGLCTSLGVGGHIIGGAYGSMMRKY 188
Query: 197 GLAADHVIDAQIIDVNGKILNRNLMGEDLFWAIKGGGGSSFGVITSWKVKLV 248
GL AD+V+DA+I+D NG+IL+R MGEDLFWAI+GGGG SFG++ WK+KLV
Sbjct: 189 GLGADNVLDARIVDANGRILDRKAMGEDLFWAIRGGGGGSFGILLWWKIKLV 344
>BP034238
Length = 518
Score = 159 bits (401), Expect = 1e-39
Identities = 77/158 (48%), Positives = 104/158 (65%)
Frame = -1
Query: 15 IFISIFLETSISFDIEKPFLHCFSSIVRNSNPTEEIVLTQNSSSYESLLQSSIRNTRFLD 74
+ +S+ S + F+ C + S+P E + T N+ S+ S+LQ+ IRN RF
Sbjct: 515 MLLSLVCAASATNSAHNTFVQCLVNHSEPSHPIAEAIFTPNTPSFSSVLQAYIRNLRFNT 336
Query: 75 SSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSNF 134
S+ KP LI+TP +SHVQAAI C K LQ++ RSGGHDYEG+SY++ PF I+D+ N
Sbjct: 335 STTRKPFLILTPLHVSHVQAAIICGQKHNLQMKTRSGGHDYEGVSYVAEDPFFILDMFNL 156
Query: 135 RSISIDIEDESAWVQSGATLGELYYAIANQSNVHAFPA 172
RSI +DI E+AWVQ+GATLGE+YY IA +S H FPA
Sbjct: 155 RSIEVDIATETAWVQAGATLGEVYYRIAEKSRKHGFPA 42
>AV426016
Length = 421
Score = 140 bits (352), Expect = 7e-34
Identities = 62/136 (45%), Positives = 94/136 (68%)
Frame = +1
Query: 213 GKILNRNLMGEDLFWAIKGGGGSSFGVITSWKVKLVHVPPKVTIFDLPKKLDQNVSEIFQ 272
G++L+R MGEDLFWAI GGGG+SFGV+ S+K+KLV VP VT+F + + L+QN ++I
Sbjct: 10 GRLLDRKSMGEDLFWAIAGGGGASFGVVLSYKIKLVQVPETVTVFQVQRTLEQNATDIIY 189
Query: 273 KWQTIAHKLPGELFLHSVMGVSNSPKHGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGL 332
WQ +A +LF+ ++ V + G K+V +F L+LG ++ L+ LM F+++GL
Sbjct: 190 NWQHVAPTTSNDLFIRLILEVVKGAQEGTKTVRATFIALFLGDSKTLVSLMSETFSQIGL 369
Query: 333 RRDDCTEMNWIQSVLY 348
R+ DCTE W++SVL+
Sbjct: 370 RQSDCTETTWLRSVLF 417
>TC19750 weakly similar to PIR|E86432|E86432 T5I8.15 protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (21%)
Length = 579
Score = 135 bits (340), Expect = 2e-32
Identities = 62/124 (50%), Positives = 91/124 (73%), Gaps = 3/124 (2%)
Frame = +1
Query: 406 PYGGRMSEISESETPFPHRNKSIFGIQYLVNWDKN--EETKMHVEWMRRLYAYMKPYVSK 463
PYGGRM+EI + +PFPHR +++ IQY NW ++ E ++ R+L+ YM P+VS
Sbjct: 1 PYGGRMAEIPSTASPFPHRAGNLWKIQYQANWMQSGKEVADHYINLTRKLHDYMTPFVSM 180
Query: 464 NPRGAYLNYRDLDIGVN-RGNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQS 522
NPR A+ NY+DLD+G++ +G SY + + +G++YFK NF+RL +VK++VDP NFFR+EQS
Sbjct: 181 NPREAFFNYKDLDLGIHHQGKKSYSKGRVYGVEYFKDNFDRLVQVKSKVDPGNFFRNEQS 360
Query: 523 IPPL 526
IP L
Sbjct: 361 IPTL 372
>TC16648 similar to PIR|T10625|T10625 reticuline oxidase homolog F21C20.180
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(19%)
Length = 483
Score = 132 bits (332), Expect = 1e-31
Identities = 66/125 (52%), Positives = 84/125 (66%), Gaps = 3/125 (2%)
Frame = +3
Query: 403 ILTPYGGRMSEISESETPFPHRNKSIFGIQYLVNWDKNEET--KMHVEWMRRLYAYMKPY 460
+ PYGG+M+EI TPFPHR ++F +QY VNW + + + +Y+YM P+
Sbjct: 21 VFNPYGGKMNEIPSDATPFPHRTGNLFKMQYSVNWHDPSPALAQNYTNQAKIMYSYMTPF 200
Query: 461 VSKNPRGAYLNYRDLDIGVNR-GNTSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRH 519
VSK R AYLNYRDLDIGVN SY+E + +G KYF +NFERL +VK VDP NFFR+
Sbjct: 201 VSKT-RSAYLNYRDLDIGVNNFDQRSYQEGEVYGTKYFGNNFERLVKVKTAVDPVNFFRN 377
Query: 520 EQSIP 524
EQSIP
Sbjct: 378 EQSIP 392
>AV410527
Length = 429
Score = 122 bits (305), Expect = 2e-28
Identities = 61/131 (46%), Positives = 87/131 (65%)
Frame = +1
Query: 7 KLALLSITIFISIFLETSISFDIEKPFLHCFSSIVRNSNPTEEIVLTQNSSSYESLLQSS 66
+L+L I + +S+ + S S K FLHC S S+P + T ++SS+ S+LQ+
Sbjct: 37 QLSLFPIVVLLSL-ISLSNSAPHTKTFLHCLESHSEPSHPITSAIFTPSNSSFSSVLQAY 213
Query: 67 IRNTRFLDSSVLKPNLIVTPQELSHVQAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPF 126
IRN RF S+ KP LI+T +SHVQAA+ C K LQ+++RSGGHDYEG+SY+++VPF
Sbjct: 214 IRNLRFNTSTTRKPYLIITALHVSHVQAAVICGQKHNLQMKIRSGGHDYEGVSYVADVPF 393
Query: 127 LIIDLSNFRSI 137
I+D+ N RSI
Sbjct: 394 FILDMFNLRSI 426
>TC13356 weakly similar to UP|Q9SVG3 (Q9SVG3) Reticuline oxidase-like
protein, partial (16%)
Length = 433
Score = 100 bits (248), Expect = 8e-22
Identities = 49/101 (48%), Positives = 71/101 (69%)
Frame = +3
Query: 33 FLHCFSSIVRNSNPTEEIVLTQNSSSYESLLQSSIRNTRFLDSSVLKPNLIVTPQELSHV 92
FL C + +++ T +V + + S+ ++LQ+ IRN RF S+ KP +IVTP + SHV
Sbjct: 135 FLKC---LTQHTTSTTNLVFSPTNPSFSTVLQNYIRNARFNTSATKKPLIIVTPLQESHV 305
Query: 93 QAAITCSVKQGLQVRVRSGGHDYEGLSYISNVPFLIIDLSN 133
QAA+ C+ +Q+++RSGGHDYEG+SYISN PF+IIDL N
Sbjct: 306 QAAVICAKTIKVQLKIRSGGHDYEGISYISNEPFIIIDLFN 428
>TC16300 similar to UP|Q93Y11 (Q93Y11) Berberine bridge enzyme-like protein,
partial (13%)
Length = 550
Score = 97.8 bits (242), Expect = 4e-21
Identities = 44/69 (63%), Positives = 53/69 (76%), Gaps = 1/69 (1%)
Frame = +1
Query: 459 PYVSKNPRGAYLNYRDLDIGVNRGN-TSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFF 517
PYVS PRGAY+NYRDLD+G+N N TSY +A WG +Y+K NF RL ++K +VD N F
Sbjct: 4 PYVSSFPRGAYVNYRDLDLGINSKNSTSYIQASAWGYRYYKDNFNRLVKIKTRVDLENVF 183
Query: 518 RHEQSIPPL 526
RHEQSIPPL
Sbjct: 184 RHEQSIPPL 210
>AV765550
Length = 369
Score = 80.1 bits (196), Expect = 8e-16
Identities = 36/56 (64%), Positives = 42/56 (74%), Gaps = 1/56 (1%)
Frame = -1
Query: 472 YRDLDIGVNRGN-TSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQSIPPL 526
YRDLD+G N+ N TSY +A WG YFK NF RL ++K +VDP N FRHEQSIPPL
Sbjct: 369 YRDLDLGTNKKNSTSYIQATAWGYMYFKDNFNRLVKIKTKVDPENVFRHEQSIPPL 202
>TC20048 weakly similar to PIR|E86432|E86432 T5I8.15 protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (9%)
Length = 530
Score = 73.9 bits (180), Expect = 6e-14
Identities = 34/63 (53%), Positives = 44/63 (68%), Gaps = 2/63 (3%)
Frame = +1
Query: 466 RGAYLNYRDLDIGVNRGN--TSYEEAKCWGLKYFKSNFERLARVKAQVDPSNFFRHEQSI 523
R AYLNY+DLD+G N +SY E +G++Y+ NF RL ++K +VDP NFFR EQSI
Sbjct: 1 RQAYLNYKDLDLGTNHHGFLSSYSEGSVYGVQYYMDNFNRLVQIKTKVDPGNFFRSEQSI 180
Query: 524 PPL 526
P L
Sbjct: 181 PVL 189
>TC18117 weakly similar to UP|Q9AYM8 (Q9AYM8) CPRD2 protein, partial (7%)
Length = 445
Score = 51.6 bits (122), Expect(2) = 1e-12
Identities = 24/46 (52%), Positives = 33/46 (71%), Gaps = 1/46 (2%)
Frame = +1
Query: 454 YAYMKPYVSKNPRGAYLNYRDLDIGVN-RGNTSYEEAKCWGLKYFK 498
Y+Y PY+SK PR AY+NYRDLDIG+N + TS+ +A +G F+
Sbjct: 1 YSYTAPYLSKYPREAYVNYRDLDIGMNQKHGTSFSQASLFGF*VFQ 138
Score = 37.7 bits (86), Expect(2) = 1e-12
Identities = 16/24 (66%), Positives = 19/24 (78%)
Frame = +2
Query: 493 GLKYFKSNFERLARVKAQVDPSNF 516
G KYFK NF RL VK++VDPS+F
Sbjct: 122 GSKYFKGNFNRLVMVKSRVDPSHF 193
>TC16688
Length = 694
Score = 30.8 bits (68), Expect = 0.57
Identities = 23/84 (27%), Positives = 38/84 (44%), Gaps = 15/84 (17%)
Frame = +3
Query: 299 HGGKSVIVSFTGLYLGIAENLLPLMQSNFAELGLRRD----DCTEM-----------NWI 343
H G ++ V LYL + L+ L++ +A + ++ D + T+M NW
Sbjct: 366 HQGWAMFVDCDFLYLADIKELIDLIEDKYAIMCVQHDYAPKETTKMDGAVQTVYPRKNWS 545
Query: 344 QSVLYLAGYPINASLNVLLQRNQT 367
VLY G+P N+ L +QT
Sbjct: 546 SMVLYNCGHPKNSVLTPDTVNSQT 617
>BU494183
Length = 460
Score = 28.1 bits (61), Expect = 3.7
Identities = 9/26 (34%), Positives = 17/26 (64%)
Frame = +2
Query: 254 VTIFDLPKKLDQNVSEIFQKWQTIAH 279
+ +FDL + QN++E++ WQ + H
Sbjct: 224 MNVFDLVEWKRQNITEVYHNWQKLNH 301
>TC14195 UP|Q40215 (Q40215) RAB8A, complete
Length = 1171
Score = 27.3 bits (59), Expect = 6.3
Identities = 19/70 (27%), Positives = 34/70 (48%), Gaps = 7/70 (10%)
Frame = -1
Query: 5 YTKLALLSITIFIS---IFLETSISFDIEKPFLH----CFSSIVRNSNPTEEIVLTQNSS 57
Y+ L + I++ +++ S+ F+I P L+ CF S + N T + +
Sbjct: 619 YSPLTFIHISLVSHKYLVYIVRSMLFNIANPILNIVE*CFISNIINQQDTHGSTVIGGRN 440
Query: 58 SYESLLQSSI 67
S +SLL SS+
Sbjct: 439 SSKSLLTSSV 410
>BF177461
Length = 439
Score = 27.3 bits (59), Expect = 6.3
Identities = 15/44 (34%), Positives = 21/44 (47%), Gaps = 4/44 (9%)
Frame = +1
Query: 95 AITCSVKQGLQVRVRSGG----HDYEGLSYISNVPFLIIDLSNF 134
A+TC + R GG HD+ L Y S+ FL+ SN+
Sbjct: 25 ALTCKRMKVTMSRKTPGGLVYVHDWNNLQYASSAAFLLAVYSNY 156
>TC14677 similar to PIR|T39903|T39903 serine-rich protein - fission yeast
(Schizosaccharomyces pombe)
{Schizosaccharomyces pombe;}, partial (3%)
Length = 616
Score = 27.3 bits (59), Expect = 6.3
Identities = 23/81 (28%), Positives = 38/81 (46%), Gaps = 4/81 (4%)
Frame = +2
Query: 7 KLALLSITIFISIFLETSISFDIEKPFLHCFSSIVRNSNPTEEIVLTQNSS----SYESL 62
+L LS+TIF SIF K F C + ++S+ T +S+ S+ S
Sbjct: 185 ELPTLSLTIFTSIF----------KNFNFCLITSSKSSSSTRVTESPSSSTYVKISFASA 334
Query: 63 LQSSIRNTRFLDSSVLKPNLI 83
+S I + L ++L+ NL+
Sbjct: 335 AKSKIEHILALSPAMLRSNLL 397
>TC14174 UP|PSBC_LOTJA (Q9BBT1) Photosystem II 44 kDa reaction center
protein (P6 protein) (CP43), complete
Length = 3406
Score = 26.9 bits (58), Expect = 8.3
Identities = 14/45 (31%), Positives = 22/45 (48%)
Frame = +3
Query: 159 YAIANQSNVHAFPAGSCPTVGIGGHFSGGGFSTIFRKHGLAADHV 203
+ S + FP C +GG F+G F T + HGLA+ ++
Sbjct: 585 FVFVGWSGLLLFP---CAYFALGGWFTGTTFVTSWYTHGLASSYL 710
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,265,257
Number of Sequences: 28460
Number of extensions: 126868
Number of successful extensions: 622
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 605
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 612
length of query: 528
length of database: 4,897,600
effective HSP length: 94
effective length of query: 434
effective length of database: 2,222,360
effective search space: 964504240
effective search space used: 964504240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0437b.11


