

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0404.10
(989 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
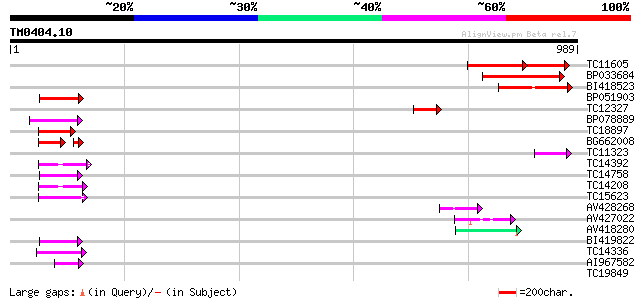
Score E
Sequences producing significant alignments: (bits) Value
TC11605 142 5e-52
BP033684 155 3e-38
BI418523 117 7e-27
BP051903 67 2e-11
TC12327 57 1e-08
BP078889 56 3e-08
TC18897 weakly similar to UP|Q9XEV3 (Q9XEV3) TIA-1 related prote... 55 7e-08
BG662008 49 3e-07
TC11323 51 1e-06
TC14392 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, pa... 48 7e-06
TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding p... 45 4e-05
TC14208 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, pa... 45 4e-05
TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding p... 45 4e-05
AV428268 45 6e-05
AV427022 45 6e-05
AV418280 45 6e-05
BI419822 43 2e-04
TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 43 3e-04
AI967582 42 5e-04
TC19849 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, p... 41 0.001
>TC11605
Length = 546
Score = 142 bits (359), Expect(2) = 5e-52
Identities = 58/105 (55%), Positives = 77/105 (73%)
Frame = +1
Query: 799 WLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIMWEIIPYAMFWSIWRARNDLVF 858
W W +ILAR+ V C P S+ LL EW SLRA SD ++WE++P A+ WSIW ARNDLVF
Sbjct: 1 WRFWNSILARDGVQWCVPGSVGDLLKEWPSLRAKSDRVLWELVPCAVVWSIWLARNDLVF 180
Query: 859 NQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKIRI 903
NQK+F + +WDLHL+RVMWW K+WWK CP++ DF Q + +++
Sbjct: 181 NQKEFIYENVWDLHLMRVMWWTKAWWKGCPYSTTDFYQILKTLKL 315
Score = 80.1 bits (196), Expect(2) = 5e-52
Identities = 40/80 (50%), Positives = 52/80 (65%)
Frame = +3
Query: 897 HFEKIRIKVAAPKVHSADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFS 956
+FE I+ V + WCPP LKFNVDGS+RG+ G+SG GG+L++A S +LGRFS
Sbjct: 294 NFENIKTVGKMRMVRNCVWCPPEANILKFNVDGSARGSPGISGAGGILRNAESAVLGRFS 473
Query: 957 CAVGSTWAYVAERKALLEGL 976
+G AY AE KA+L L
Sbjct: 474 KPLGVLCAYQAEVKAILLAL 533
>BP033684
Length = 557
Score = 155 bits (392), Expect = 3e-38
Identities = 71/144 (49%), Positives = 90/144 (62%)
Frame = -1
Query: 825 EWSSLRAISDPIMWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWW 884
EWSS+ SDP++W +I YA+ WS+W R VF K S+ +WDLHL+ + WWIK+ W
Sbjct: 557 EWSSIVVASDPMLWNLILYALVWSLWLERKQSVFKGKLASSQEVWDLHLMTIFWWIKAVW 378
Query: 885 KECPFTVLDFSQHFEKIRIKVAAPKVHSADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVL 944
KECP+ F + F + K PK W PP G LKFNVDG+S+GN G SG+GGVL
Sbjct: 377 KECPYNFDQFHRDFCMLSFKKLIPKQRLKGWTPPSRGILKFNVDGASQGNPGPSGVGGVL 198
Query: 945 KDANSTMLGRFSCAVGSTWAYVAE 968
D S +LG FS +G WAY AE
Sbjct: 197 CDYRSMVLGFFSINMGHGWAYEAE 126
>BI418523
Length = 496
Score = 117 bits (294), Expect = 7e-27
Identities = 56/131 (42%), Positives = 81/131 (61%), Gaps = 2/131 (1%)
Frame = +1
Query: 853 RNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKIRI--KVAAPKV 910
RN +F K+ + +WD HL+++ WWIKS W+EC + + Q+ IRI K+ P+
Sbjct: 1 RNAYIFRGKEIVSTEVWDSHLLKLSWWIKSMWRECHYDIYQIMQNLGDIRITPKLKPPRC 180
Query: 911 HSADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAVGSTWAYVAERK 970
S W PP G +K NVDG+S+GN G SG+GGV +DAN +LG FS G+ WAY A+ +
Sbjct: 181 WS--WSPPSAGFIKCNVDGASQGNPGPSGVGGVFRDANRKILGYFSLNSGNGWAYEAKVR 354
Query: 971 ALLEGLLLCGK 981
++L L+ K
Sbjct: 355 SILNALVFIQK 387
>BP051903
Length = 342
Score = 66.6 bits (161), Expect = 2e-11
Identities = 33/77 (42%), Positives = 50/77 (64%)
Frame = +2
Query: 53 VDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRMNG 112
VDG+ D+T + Q++ FA FGR++ VFVQ ++ R +FGFVRF+ A A+ +NG
Sbjct: 68 VDGIPDRTDYYQIRGLFAQFGRVVNVFVQRQRKIGRLFRFGFVRFASLEVASKAIDFLNG 247
Query: 113 SILNGATINVSLARLPQ 129
L A ++VS+AR P+
Sbjct: 248 FRLGDAALSVSMARFPR 298
>TC12327
Length = 568
Score = 57.4 bits (137), Expect = 1e-08
Identities = 25/48 (52%), Positives = 35/48 (72%)
Frame = +1
Query: 705 ETKMDVHRNKVLLGWATSMDMGLEIVPAVGSAGGLVTLWKTSLFQVSQ 752
E+K+D +R K ++ WA S+ M E VPA+G AGGLV+LW+ S F+V Q
Sbjct: 1 ESKLDENRKKTIVSWAKSIGMEFEFVPAIGVAGGLVSLWRKSTFKVVQ 144
>BP078889
Length = 474
Score = 55.8 bits (133), Expect = 3e-08
Identities = 32/92 (34%), Positives = 49/92 (52%)
Frame = +1
Query: 35 GNNRRADGIHRRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGF 94
G R +G RR + FVDG+ + + ++ FA FGRI +FVQ + RR +FGF
Sbjct: 196 GGLNRNNGGERRRKPTPFVDGIEEAIEYHHLRGLFALFGRISNLFVQRKHKFGRRFRFGF 375
Query: 95 VRFSFDSDAEVAMRRMNGSILNGATINVSLAR 126
VRF + A ++G + G ++V+ AR
Sbjct: 376 VRFFSEEQAAAVAFVLDGLRVGGTALSVAPAR 471
>TC18897 weakly similar to UP|Q9XEV3 (Q9XEV3) TIA-1 related protein, partial
(5%)
Length = 481
Score = 54.7 bits (130), Expect = 7e-08
Identities = 23/66 (34%), Positives = 41/66 (61%)
Frame = +1
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
+LF+DG+++ T + ++ F G +L VF+Q +R +R +FGFVR+S A +A+
Sbjct: 280 TLFIDGISEPTNYYHIRGLFGQIGSLLNVFLQKQRRIRRNFRFGFVRYSTRDVASMAINL 459
Query: 110 MNGSIL 115
NG ++
Sbjct: 460 FNGVLM 477
>BG662008
Length = 457
Score = 48.5 bits (114), Expect(2) = 3e-07
Identities = 22/48 (45%), Positives = 32/48 (65%)
Frame = +3
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRF 97
+LFVDG++D + Q++ F+ G +L VFVQ +R R +FGFVRF
Sbjct: 102 TLFVDGISDFVDYSQIRGLFSHIGSLLNVFVQRQRRIGRAFRFGFVRF 245
Score = 23.9 bits (50), Expect(2) = 3e-07
Identities = 8/19 (42%), Positives = 16/19 (84%)
Frame = +2
Query: 111 NGSILNGATINVSLARLPQ 129
NG+++ G+ ++V+LAR P+
Sbjct: 269 NGALVGGSFLSVALARFPR 325
>TC11323
Length = 657
Score = 50.8 bits (120), Expect = 1e-06
Identities = 27/65 (41%), Positives = 35/65 (53%)
Frame = +2
Query: 915 WCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAVGSTWAYVAERKALLE 974
W P G K N DGSS GN G +G GG+L+D +S + FS + A+LE
Sbjct: 5 WLKPVHGRFKLNFDGSSLGNRGNAGGGGLLRDGSSNFIFGFSIFLAVAQIMKLSLCAILE 184
Query: 975 GLLLC 979
GLL+C
Sbjct: 185 GLLVC 199
>TC14392 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (77%)
Length = 1576
Score = 48.1 bits (113), Expect = 7e-06
Identities = 29/93 (31%), Positives = 49/93 (52%)
Frame = +3
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
++FV L T ++ F +G ++ V + KR GFV+F+ S AE A+R
Sbjct: 945 TIFVGNLDPNVTDDHLRQVFGLYGDLVHVKIPQGKR------CGFVQFADRSCAEEALRV 1106
Query: 110 MNGSILNGATINVSLARLPQDQHTRPRPTDNQW 142
+NG++L G + +S R P ++ T+ P +QW
Sbjct: 1107 LNGTLLGGQNVRLSWGRSPSNKQTQSDP--SQW 1199
Score = 29.3 bits (64), Expect = 3.2
Identities = 19/77 (24%), Positives = 34/77 (43%), Gaps = 1/77 (1%)
Frame = +3
Query: 50 SLFVDGLTDQTTHLQVKNAFAS-FGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMR 108
++FV L T + F + + + G V + R +GFVRF+ +S+ AM
Sbjct: 618 TIFVGDLAADVTDYHLTEVFRTRYNSVKGAKVVIDRLTSRTKGYGFVRFADESEQVRAMT 797
Query: 109 RMNGSILNGATINVSLA 125
M G + + + + A
Sbjct: 798 EMQGVVCSTRPMRIGPA 848
>TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding protein 2,
partial (93%)
Length = 725
Score = 45.4 bits (106), Expect = 4e-05
Identities = 27/75 (36%), Positives = 40/75 (53%)
Frame = +1
Query: 52 FVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRMN 111
FV GL T ++ AF+S+G I+ V N + R FGFV F+ + A+ MN
Sbjct: 91 FVGGLAWTTDSHTLEQAFSSYGEIIDSKVVNDRETGRSRGFGFVTFTSEEAMRSAIEGMN 270
Query: 112 GSILNGATINVSLAR 126
G+ L+G I V+ A+
Sbjct: 271 GNELDGRNITVNEAQ 315
>TC14208 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (41%)
Length = 837
Score = 45.4 bits (106), Expect = 4e-05
Identities = 25/86 (29%), Positives = 45/86 (52%)
Frame = +2
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
++FV L ++ F+ +G ++ V + KR GFV+F+ S AE A+R
Sbjct: 194 TIFVGNLDPNVNDDHLRQVFSPYGELVHVKIPAGKR------CGFVQFADRSGAEEALRV 355
Query: 110 MNGSILNGATINVSLARLPQDQHTRP 135
+NG++L G + +S R P ++ +P
Sbjct: 356 LNGTLLGGQNVRLSWGRSPSNKQAQP 433
>TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 683
Score = 45.4 bits (106), Expect = 4e-05
Identities = 30/86 (34%), Positives = 42/86 (47%)
Frame = +1
Query: 51 LFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRM 110
LF+ GL+ +++AF+SFG + V + R FGFV F + A A+ M
Sbjct: 190 LFIGGLSYGVDDKSLEDAFSSFGTVAEARVIVDRDTGRSRGFGFVSFDSEESASSALSSM 369
Query: 111 NGSILNGATINVSLARLPQDQHTRPR 136
+G LNG I VS A D+ PR
Sbjct: 370 DGQDLNGRNIRVSYA---NDRPAGPR 438
>AV428268
Length = 429
Score = 45.1 bits (105), Expect = 6e-05
Identities = 24/76 (31%), Positives = 40/76 (52%)
Frame = +3
Query: 750 VSQQLMECNIAVKASLIRRGIPIENGGLCDICGLIGESESHLMLHCPKIWLLWTTILARE 809
+S +++ + K +L RRG+ I N +C +C L ES HL+ CP + +W+ I
Sbjct: 114 LSWKVLINRVQTKVNLNRRGVVISN--VCPLCSLDEESTDHLLFSCPIV*RIWSKISE*F 287
Query: 810 EVYCCFPSSIQGLLLE 825
V+ FP+ G L+
Sbjct: 288 GVFSVFPNDSHGHFLQ 335
>AV427022
Length = 429
Score = 45.1 bits (105), Expect = 6e-05
Identities = 31/116 (26%), Positives = 52/116 (44%), Gaps = 10/116 (8%)
Frame = +1
Query: 777 LCDICGLIGESESHLMLHCPKIWLL------W----TTILAREEVYCCFPSSIQGLLLEW 826
+C +C L ES SHL+ C L+ W T ++ +V+ SSI W
Sbjct: 37 ICPLCCLAEESGSHLLFSCSNSMLI*YECHAWLGVSTAQVSDPKVHLLQFSSIG-----W 201
Query: 827 SSLRAISDPIMWEIIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKS 882
S + + + +W ++ WSIW RN +VFN + D + + + W++S
Sbjct: 202 SKAQKLGESAIW----MSVLWSIWCLRNMVVFNGGELDKDQLLEQIQVTAWRWLRS 357
>AV418280
Length = 398
Score = 45.1 bits (105), Expect = 6e-05
Identities = 30/126 (23%), Positives = 46/126 (35%), Gaps = 11/126 (8%)
Frame = -2
Query: 778 CDICGLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIM 837
C C + E HL C +W + + P+S ++S + +
Sbjct: 388 CPFCTTVEECSGHLFFTCVFSMGVWQALHRWLGISVALPASTLANFAQFSITARNKNQRL 209
Query: 838 WEI-IPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKS----------WWKE 886
E+ I A WS+W RN ++F Y+ DL R W+K+ WK
Sbjct: 208 GELAIWIATVWSLWIQRNSIIFRNNALDHSYLLDLIQSRSWHWLKAKFCGFTYSLYEWKS 29
Query: 887 CPFTVL 892
CP L
Sbjct: 28 CPLECL 11
>BI419822
Length = 380
Score = 43.1 bits (100), Expect = 2e-04
Identities = 26/75 (34%), Positives = 40/75 (52%)
Frame = +2
Query: 52 FVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRMN 111
FV GL T + ++ AF+ +G I+ V N + R FGFV F+ + + A+ MN
Sbjct: 86 FVGGLAWATDNEALEKAFSPYGEIVESKVINDRETGRSRGFGFVTFASEQAMKDAIEAMN 265
Query: 112 GSILNGATINVSLAR 126
G L+G I V+ A+
Sbjct: 266 GQNLDGRNITVNEAQ 310
>TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(85%)
Length = 1917
Score = 42.7 bits (99), Expect = 3e-04
Identities = 27/87 (31%), Positives = 42/87 (48%)
Frame = +2
Query: 48 GKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAM 107
G +L+V L D ++K F+S+G I V R GFV FS +A A+
Sbjct: 671 GANLYVKNLDDSIADEKLKELFSSYGTITSCKVMRDPNGISRGS-GFVAFSTPEEASRAL 847
Query: 108 RRMNGSILNGATINVSLARLPQDQHTR 134
MNG ++ + V+LA+ +D+ R
Sbjct: 848 LEMNGKMVVSKPLYVTLAQRKEDRRAR 928
Score = 40.8 bits (94), Expect = 0.001
Identities = 23/86 (26%), Positives = 42/86 (48%)
Frame = +2
Query: 37 NRRADGIHRRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVR 96
+ R I + ++F+ L H + + F++FG IL V Q + +GFV+
Sbjct: 56 SHRDPSIRKSGAGNIFIKNLDKAIDHKALHDTFSTFGNILSCKVATDSSGQSKG-YGFVQ 232
Query: 97 FSFDSDAEVAMRRMNGSILNGATINV 122
F + A+ A+ ++NG +LN + V
Sbjct: 233 FETEEAAQNAIEKLNGMLLNDKQVYV 310
Score = 39.3 bits (90), Expect = 0.003
Identities = 24/64 (37%), Positives = 35/64 (54%), Gaps = 1/64 (1%)
Frame = +2
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSK-FGFVRFSFDSDAEVAMR 108
++FV L++ TT ++KN F FG I V ++ +SK FGFV F DA A+
Sbjct: 368 NVFVKNLSESTTDDELKNVFGEFGTITSAVV--MRDGDGKSKCFGFVNFENADDAARAVE 541
Query: 109 RMNG 112
+NG
Sbjct: 542 SLNG 553
>AI967582
Length = 356
Score = 42.0 bits (97), Expect = 5e-04
Identities = 22/51 (43%), Positives = 30/51 (58%)
Frame = +2
Query: 78 VFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRRMNGSILNGATINVSLARLP 128
VFVQ RR FGFV F SDA A+ R+NG + GA ++V+++ P
Sbjct: 20 VFVQRKHEVGRRFHFGFVHFLSWSDAXTAVARLNGMKIGGAHLSVTVSTFP 172
>TC19849 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, partial
(30%)
Length = 603
Score = 40.8 bits (94), Expect = 0.001
Identities = 23/87 (26%), Positives = 43/87 (48%)
Frame = +2
Query: 48 GKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAM 107
G +L++ L D +++ F+ FG I + R GFV FS +A A+
Sbjct: 233 GANLYLKNLDDNVDDEKLRELFSEFGTITSCKIMRDPHGVSRGS-GFVAFSTPEEATRAL 409
Query: 108 RRMNGSILNGATINVSLARLPQDQHTR 134
MNG +++G + V+LA+ +++ +
Sbjct: 410 GEMNGKMVDGKPLYVALAQKKEERRAK 490
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.134 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,808,928
Number of Sequences: 28460
Number of extensions: 297204
Number of successful extensions: 2213
Number of sequences better than 10.0: 113
Number of HSP's better than 10.0 without gapping: 2108
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2201
length of query: 989
length of database: 4,897,600
effective HSP length: 99
effective length of query: 890
effective length of database: 2,080,060
effective search space: 1851253400
effective search space used: 1851253400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0404.10


