

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
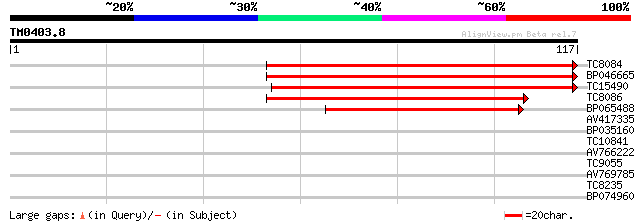
Query= TM0403.8
(117 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 84 5e-18
BP046665 79 2e-16
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 75 4e-15
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 66 2e-12
BP065488 48 4e-07
AV417335 26 2.0
BP035160 25 3.4
TC10841 homologue to UP|Q9AU01 (Q9AU01) Phosphate transporter 1,... 25 4.4
AV766222 24 5.8
TC9055 24 7.6
AV769785 23 9.9
TC8235 similar to UP|Q9LMR0 (Q9LMR0) F7H2.8 protein, partial (27%) 23 9.9
BP074960 23 9.9
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 84.3 bits (207), Expect = 5e-18
Identities = 40/64 (62%), Positives = 48/64 (74%)
Frame = +1
Query: 54 MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQGQSLSHV 113
+I ATV G +G +IPR+ + SDSG PFKF RR F ++LCFAMTIN +QGQSLSHV
Sbjct: 106 VIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*FPISLCFAMTINKSQGQSLSHV 285
Query: 114 SLYL 117
SLYL
Sbjct: 286 SLYL 297
>BP046665
Length = 524
Score = 79.0 bits (193), Expect = 2e-16
Identities = 38/64 (59%), Positives = 46/64 (71%)
Frame = -1
Query: 54 MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQGQSLSHV 113
+I ATV +G +IPR+ + SDSG PFKF RRQF ++LC AMTIN +QGQSLSHV
Sbjct: 386 VIKATVIT*TNIGDDIFIPRMDMVPSDSGYPFKFERRQFPISLCSAMTINKSQGQSLSHV 207
Query: 114 SLYL 117
LYL
Sbjct: 206 GLYL 195
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 74.7 bits (182), Expect = 4e-15
Identities = 36/63 (57%), Positives = 43/63 (68%)
Frame = +2
Query: 55 IVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQGQSLSHVS 114
I TV G +G IPR+ + SDS PFKF RRQ ++LCFAMTIN +QG+SLSHV
Sbjct: 137 IQVTVITGTHIGDDISIPRMDMVPSDSSYPFKFERRQSPISLCFAMTINKSQGRSLSHVG 316
Query: 115 LYL 117
LYL
Sbjct: 317 LYL 325
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 65.9 bits (159), Expect = 2e-12
Identities = 30/54 (55%), Positives = 38/54 (69%)
Frame = +1
Query: 54 MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQG 107
+I ATV G +G +IPR+ + SDSG PFKF RR F ++LCFAMTIN +QG
Sbjct: 214 VIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*FPISLCFAMTINKSQG 375
>BP065488
Length = 439
Score = 48.1 bits (113), Expect = 4e-07
Identities = 22/41 (53%), Positives = 29/41 (70%)
Frame = -1
Query: 66 GKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQ 106
G +IPR+++ S SG P KF R QF ++LCFAMTIN +Q
Sbjct: 379 GDDIFIPRMNMVPSVSGDPLKFERCQFPISLCFAMTINKSQ 257
>AV417335
Length = 428
Score = 25.8 bits (55), Expect = 2.0
Identities = 11/27 (40%), Positives = 14/27 (51%)
Frame = -2
Query: 60 FPGNKLGKTAYIPRISLTSSDSGLPFK 86
FPG GKT P + + +D PFK
Sbjct: 415 FPGTTSGKTLVSPSVVVILNDKDWPFK 335
>BP035160
Length = 482
Score = 25.0 bits (53), Expect = 3.4
Identities = 12/33 (36%), Positives = 17/33 (51%)
Frame = +1
Query: 83 LPFKFSRRQFLVTLCFAMTINNNQGQSLSHVSL 115
LP RR + + CF + + +G LSHV L
Sbjct: 274 LPVP*IRRTLVASHCFPLWLKVEEGHCLSHVLL 372
>TC10841 homologue to UP|Q9AU01 (Q9AU01) Phosphate transporter 1, partial
(59%)
Length = 991
Score = 24.6 bits (52), Expect = 4.4
Identities = 16/41 (39%), Positives = 22/41 (53%), Gaps = 4/41 (9%)
Frame = +3
Query: 62 GNKLG-KTAY---IPRISLTSSDSGLPFKFSRRQFLVTLCF 98
G+KLG K Y + + + S SGL F S + + TLCF
Sbjct: 192 GDKLGRKKVYGMTLMMMVICSIGSGLSFGHSPKSVMATLCF 314
>AV766222
Length = 260
Score = 24.3 bits (51), Expect = 5.8
Identities = 8/17 (47%), Positives = 14/17 (82%)
Frame = -3
Query: 59 VFPGNKLGKTAYIPRIS 75
V GNK+G+T+ +PR++
Sbjct: 258 VVSGNKMGRTSSLPRVA 208
>TC9055
Length = 577
Score = 23.9 bits (50), Expect = 7.6
Identities = 19/62 (30%), Positives = 24/62 (38%), Gaps = 7/62 (11%)
Frame = -1
Query: 56 VATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTL-------CFAMTINNNQGQ 108
+ T FPGNK+G I + + L SRRQ TL C + N Q
Sbjct: 214 ILTHFPGNKMGSLKVIKKAKAIPQNLYL----SRRQLRSTLR**Q*PVCLLTHVKGNNHQ 47
Query: 109 SL 110
L
Sbjct: 46 LL 41
>AV769785
Length = 600
Score = 23.5 bits (49), Expect = 9.9
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -3
Query: 84 PFKFSRRQFLVTLCFAMTINNNQ 106
P+ +R +FLVT + I NNQ
Sbjct: 94 PYTAARTRFLVTFLCFLRIRNNQ 26
>TC8235 similar to UP|Q9LMR0 (Q9LMR0) F7H2.8 protein, partial (27%)
Length = 847
Score = 23.5 bits (49), Expect = 9.9
Identities = 14/28 (50%), Positives = 16/28 (57%)
Frame = -3
Query: 71 IPRISLTSSDSGLPFKFSRRQFLVTLCF 98
IP+ L+S SG S RQFL LCF
Sbjct: 263 IPKFYLSSCHSGTGLAGSGRQFL--LCF 186
>BP074960
Length = 439
Score = 23.5 bits (49), Expect = 9.9
Identities = 12/37 (32%), Positives = 21/37 (56%), Gaps = 2/37 (5%)
Frame = -3
Query: 70 YIPRISLTSSDSGLPFKFSRRQ--FLVTLCFAMTINN 104
+I +SLT +S P+K ++R+ F C + +NN
Sbjct: 266 FILSLSLTRKNSWFPYKGTKRELNFTYIPCLS*VLNN 156
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.353 0.153 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,818,723
Number of Sequences: 28460
Number of extensions: 20049
Number of successful extensions: 190
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 190
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 190
length of query: 117
length of database: 4,897,600
effective HSP length: 78
effective length of query: 39
effective length of database: 2,677,720
effective search space: 104431080
effective search space used: 104431080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0403.8


