

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.5
(359 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
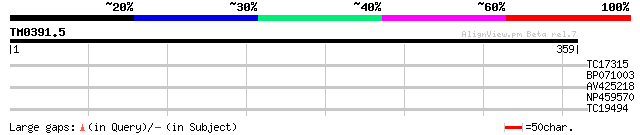
Score E
Sequences producing significant alignments: (bits) Value
TC17315 similar to UP|ATP3_ARATH (Q96250) ATP synthase gamma cha... 30 0.82
BP071003 30 0.82
AV425218 27 5.3
NP459570 hypothetical protein [Lotus japonicus] 26 9.0
TC19494 weakly similar to UP|Q9SND9 (Q9SND9) Anthranilate N-hydr... 26 9.0
>TC17315 similar to UP|ATP3_ARATH (Q96250) ATP synthase gamma chain,
mitochondrial precursor , partial (7%)
Length = 680
Score = 29.6 bits (65), Expect = 0.82
Identities = 16/32 (50%), Positives = 20/32 (62%)
Frame = +2
Query: 152 HELDAAIPARTMPATPPSQGTGESETSQELQS 183
H +DA PAR + PS+GT ETSQ L+S
Sbjct: 347 HCIDAPNPARQSVGSLPSRGT*VIETSQALES 442
>BP071003
Length = 466
Score = 29.6 bits (65), Expect = 0.82
Identities = 16/32 (50%), Positives = 20/32 (62%)
Frame = -1
Query: 152 HELDAAIPARTMPATPPSQGTGESETSQELQS 183
H +DA PAR + PS+GT ETSQ L+S
Sbjct: 220 HCIDAPNPARQSVGSLPSRGT*VIETSQALES 125
>AV425218
Length = 419
Score = 26.9 bits (58), Expect = 5.3
Identities = 15/46 (32%), Positives = 25/46 (53%), Gaps = 1/46 (2%)
Frame = -2
Query: 49 FSIPLLSFQVISSNNVYKMSLKLIYADC-IQKLLAILALTLIIKIS 93
F P + + +N + LKLIY DC I +L +++ L++ IS
Sbjct: 400 FIQPFIVLKAKILHNFLPVPLKLIYYDCIIHNILQMMSYGLVLSIS 263
>NP459570 hypothetical protein [Lotus japonicus]
Length = 972
Score = 26.2 bits (56), Expect = 9.0
Identities = 19/72 (26%), Positives = 29/72 (39%), Gaps = 10/72 (13%)
Frame = +1
Query: 99 KWIITG-LSLSTLPNTLIL---------GIPLMKAMYKDEADALLPQIIFLQSMIWYNLL 148
+W +G L S L +LI +P K D + + P +IF Q LL
Sbjct: 175 RWFFSGHLPFSDLYESLIFLSWGFSIFYMVPRFKKQKNDLSTIIAPSVIFTQGFATSGLL 354
Query: 149 LFLHELDAAIPA 160
+H+ +PA
Sbjct: 355 TEMHQSVILVPA 390
>TC19494 weakly similar to UP|Q9SND9 (Q9SND9) Anthranilate
N-hydroxycinnamoyl/benzoyltransferase-like protein,
partial (4%)
Length = 420
Score = 26.2 bits (56), Expect = 9.0
Identities = 12/27 (44%), Positives = 13/27 (47%)
Frame = +1
Query: 30 WWKLFTPDQCSGINKFVANFSIPLLSF 56
WW L TP QC G +N PL F
Sbjct: 253 WWPL*TPLQCQGSTTSRSNL*CPLERF 333
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.140 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,023,858
Number of Sequences: 28460
Number of extensions: 79469
Number of successful extensions: 612
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 606
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 611
length of query: 359
length of database: 4,897,600
effective HSP length: 91
effective length of query: 268
effective length of database: 2,307,740
effective search space: 618474320
effective search space used: 618474320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0391.5


