

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.14
(76 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
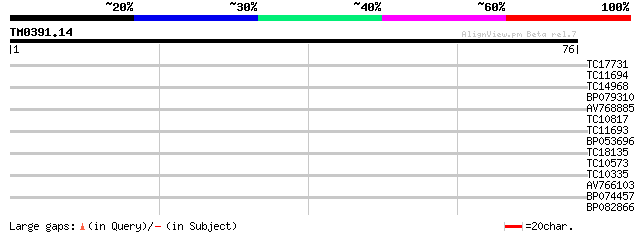
Score E
Sequences producing significant alignments: (bits) Value
TC17731 28 0.41
TC11694 similar to UP|Q8LH29 (Q8LH29) OJ1126_G08.9 protein, part... 27 0.54
TC14968 similar to UP|Q8TTE6 (Q8TTE6) Bacterial extracellular so... 27 0.70
BP079310 27 0.92
AV768885 26 1.2
TC10817 similar to UP|O80717 (O80717) At2g47080 protein, partial... 26 1.2
TC11693 26 1.2
BP053696 25 2.0
TC18135 homologue to UP|Q9MAY4 (Q9MAY4) ABC transporter homolog,... 25 2.0
TC10573 similar to UP|Q9FIK3 (Q9FIK3) Emb|CAB71043.1, partial (10%) 24 4.6
TC10335 24 4.6
AV766103 24 6.0
BP074457 24 6.0
BP082866 23 7.8
>TC17731
Length = 624
Score = 27.7 bits (60), Expect = 0.41
Identities = 9/22 (40%), Positives = 16/22 (71%)
Frame = -1
Query: 54 IYLMDIQIWYAIYSSLKMKLWI 75
+ L +I +WYAI+S+ ++ WI
Sbjct: 597 LILYNITVWYAIFSTQSIEFWI 532
>TC11694 similar to UP|Q8LH29 (Q8LH29) OJ1126_G08.9 protein, partial (13%)
Length = 463
Score = 27.3 bits (59), Expect = 0.54
Identities = 16/62 (25%), Positives = 29/62 (45%)
Frame = +1
Query: 12 IQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLMDIQIWYAIYSSLKM 71
+Q ++ S + ++ + +Q G GL+W+ ++YLMD W+ IY L
Sbjct: 7 LQ*LVLPSLIKPDMIPLAHQTLLLKMGGMVCSEGLVWLTLTILYLMDTLRWF-IYRYLNN 183
Query: 72 KL 73
L
Sbjct: 184DL 189
>TC14968 similar to UP|Q8TTE6 (Q8TTE6) Bacterial extracellular
solute-binding family 3 protein, partial (3%)
Length = 503
Score = 26.9 bits (58), Expect = 0.70
Identities = 7/31 (22%), Positives = 18/31 (57%)
Frame = +1
Query: 32 WHAFIKNGNTVYVGLLWIPAVLIYLMDIQIW 62
WH ++ V+ +++P + YL+ +++W
Sbjct: 229 WHVWVGRNEAVFREGVFLPECIFYLVKLKVW 321
>BP079310
Length = 505
Score = 26.6 bits (57), Expect = 0.92
Identities = 14/45 (31%), Positives = 21/45 (46%)
Frame = +2
Query: 1 IIFFSFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVG 45
IIFFSF Y L + P++A + K + + G + VG
Sbjct: 104 IIFFSFFYHLMLPPLVAINTKNTHTKKITHDVLVIFAFGVLIKVG 238
>AV768885
Length = 266
Score = 26.2 bits (56), Expect = 1.2
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = -2
Query: 2 IFFSFSYFLQIQPIIASSRVV 22
+FF FS F+ I ++ SSR++
Sbjct: 124 VFFIFSIFISIHSVVCSSRIL 62
>TC10817 similar to UP|O80717 (O80717) At2g47080 protein, partial (13%)
Length = 648
Score = 26.2 bits (56), Expect = 1.2
Identities = 8/33 (24%), Positives = 19/33 (57%)
Frame = +3
Query: 44 VGLLWIPAVLIYLMDIQIWYAIYSSLKMKLWIF 76
V L W+ V+I+ + +W+ + L++ +W +
Sbjct: 381 VSLYWLNCVMIFTYVLGMWFLV*LHLRVWIWFY 479
>TC11693
Length = 521
Score = 26.2 bits (56), Expect = 1.2
Identities = 12/32 (37%), Positives = 17/32 (52%)
Frame = -2
Query: 26 KDVEYQWHAFIKNGNTVYVGLLWIPAVLIYLM 57
K E W++FI N VY G+ P L+ L+
Sbjct: 433 KTTELTWNSFITNNILVYAGVQHFPFNLLKLI 338
>BP053696
Length = 509
Score = 25.4 bits (54), Expect = 2.0
Identities = 11/32 (34%), Positives = 19/32 (59%)
Frame = -1
Query: 24 ELKDVEYQWHAFIKNGNTVYVGLLWIPAVLIY 55
+LK+V + H ++ NG+TV L W + I+
Sbjct: 467 KLKEVVDRLHCYVXNGDTVCGCLFWFGFLYIH 372
>TC18135 homologue to UP|Q9MAY4 (Q9MAY4) ABC transporter homolog, partial
(50%)
Length = 968
Score = 25.4 bits (54), Expect = 2.0
Identities = 13/44 (29%), Positives = 22/44 (49%)
Frame = -1
Query: 5 SFSYFLQIQPIIASSRVVLELKDVEYQWHAFIKNGNTVYVGLLW 48
SFS+FLQ QP + ++ + + V F G + + +LW
Sbjct: 578 SFSFFLQSQPFLITAHDSFQCRHV----RGFNLPGKVINIHVLW 459
>TC10573 similar to UP|Q9FIK3 (Q9FIK3) Emb|CAB71043.1, partial (10%)
Length = 605
Score = 24.3 bits (51), Expect = 4.6
Identities = 15/59 (25%), Positives = 29/59 (48%), Gaps = 10/59 (16%)
Frame = +3
Query: 6 FSYFLQIQPIIASSRVVLELKDVEYQ----------WHAFIKNGNTVYVGLLWIPAVLI 54
F + I+ +ASS+VV+ L +++ +++F Y+G+LWI + I
Sbjct: 201 FLIHMIIEVFLASSKVVMHLLGLKFLMCRASVL*KLYNSFFVYVFLFYIGILWISTLKI 377
>TC10335
Length = 602
Score = 24.3 bits (51), Expect = 4.6
Identities = 14/66 (21%), Positives = 31/66 (46%), Gaps = 7/66 (10%)
Frame = +1
Query: 3 FFSFSYFLQIQPIIASSRVVLELKDVE-------YQWHAFIKNGNTVYVGLLWIPAVLIY 55
+FSF F+ + + S + L++ Y W + + +YVGLL++ + ++
Sbjct: 304 WFSFFLFIFLVGCLVRSSPLFGLRETSRPFMGIFYTWLHMHTDFHCLYVGLLYVGLLSLF 483
Query: 56 LMDIQI 61
L + +
Sbjct: 484 LREYML 501
>AV766103
Length = 582
Score = 23.9 bits (50), Expect = 6.0
Identities = 10/23 (43%), Positives = 13/23 (56%)
Frame = -3
Query: 32 WHAFIKNGNTVYVGLLWIPAVLI 54
WH +KNGN + IPA +I
Sbjct: 487 WHRLLKNGNHLMTTPRSIPASII 419
>BP074457
Length = 527
Score = 23.9 bits (50), Expect = 6.0
Identities = 14/39 (35%), Positives = 21/39 (52%), Gaps = 2/39 (5%)
Frame = +3
Query: 7 SYFLQIQPIIASSRVVLELKDVEYQW--HAFIKNGNTVY 43
S+ L I+ IASS ++ + ++EYQ F GN Y
Sbjct: 327 SHHLFIRTSIASSHILTKASELEYQTLISEFFPPGNASY 443
>BP082866
Length = 470
Score = 23.5 bits (49), Expect = 7.8
Identities = 10/20 (50%), Positives = 13/20 (65%), Gaps = 2/20 (10%)
Frame = +2
Query: 32 WHAFIKNGNTVYVG--LLWI 49
WH NG+T+ +G LLWI
Sbjct: 356 WHKEESNGSTILLGMLLLWI 415
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.333 0.146 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,660,606
Number of Sequences: 28460
Number of extensions: 19904
Number of successful extensions: 167
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 167
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 167
length of query: 76
length of database: 4,897,600
effective HSP length: 52
effective length of query: 24
effective length of database: 3,417,680
effective search space: 82024320
effective search space used: 82024320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0391.14


