

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
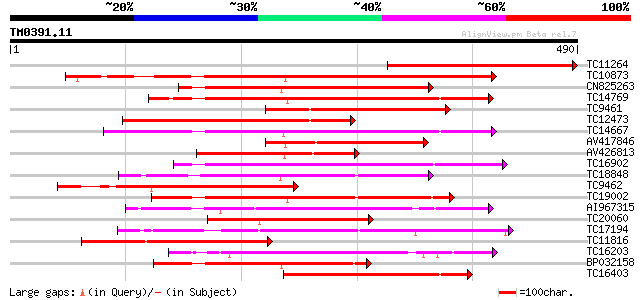
Query= TM0391.11
(490 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC11264 similar to UP|Q84SA6 (Q84SA6) Serine/threonine protein k... 331 2e-91
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 325 1e-89
CN825263 239 7e-64
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 233 4e-62
TC9461 similar to UP|O49840 (O49840) Protein kinase (AT2G02800/T... 231 3e-61
TC12473 similar to UP|Q9M068 (Q9M068) Protein kinase-like protei... 229 1e-60
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 224 2e-59
AV417846 214 3e-56
AV426813 211 2e-55
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 207 4e-54
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 206 7e-54
TC9462 similar to UP|O49840 (O49840) Protein kinase (AT2G02800/T... 202 1e-52
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 193 5e-50
AI967315 189 9e-49
TC20060 similar to UP|Q9LZF8 (Q9LZF8) Protein kinase-like (Prote... 177 3e-45
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 177 3e-45
TC11816 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B ... 175 1e-44
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 172 1e-43
BP032158 170 4e-43
TC16403 homologue to UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein... 164 3e-41
>TC11264 similar to UP|Q84SA6 (Q84SA6) Serine/threonine protein kinase,
partial (24%)
Length = 1156
Score = 331 bits (848), Expect = 2e-91
Identities = 161/164 (98%), Positives = 163/164 (99%)
Frame = +3
Query: 327 KSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSV 386
+ +VYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSV
Sbjct: 6 RHEVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSV 185
Query: 387 KGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPN 446
KGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPN
Sbjct: 186 KGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPN 365
Query: 447 HKNGIRTQLVSLPKKGQPLRILSSPNCPNGSPYSRYSKSPKPVG 490
HKNGIRTQLVSLPKKGQPLRILSSPNCPNGSPYSRYSKSPKPVG
Sbjct: 366 HKNGIRTQLVSLPKKGQPLRILSSPNCPNGSPYSRYSKSPKPVG 497
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 325 bits (833), Expect = 1e-89
Identities = 183/377 (48%), Positives = 238/377 (62%), Gaps = 5/377 (1%)
Frame = +1
Query: 49 SRPKVDSSI--SGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIA 106
SR ++D + SG+S+ NG + E K + T+ SS +SN ++ A
Sbjct: 241 SRVEIDDNATRSGSSSGNGGVNAAAAATDGERK----RETKGKGKGVSSSNGSSNGKTAA 408
Query: 107 STPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKP 166
++ F F L ATRNF+ +L+GEGGFG V+KG +
Sbjct: 409 AS----------------FGFRELADATRNFKEANLIGEGGFGKVYKGRLT--------- 513
Query: 167 GTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRG 226
TG VAVK L+H+G QG +E++ E+ L L H NLV+LIG+C + DQRLLVYE+MP G
Sbjct: 514 -TGEAVAVKQLSHDGRQGFQEFVMEVLMLSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMG 690
Query: 227 SLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYN 283
SLE+HLF PL WS RMK+A+GAA+GL +LH + P+IYRD K++NILLD E+N
Sbjct: 691 SLEDHLFELSHDKEPLNWSTRMKVAVGAARGLEYLHCTADPPVIYRDLKSANILLDNEFN 870
Query: 284 AKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTG 343
KLSDFGLAK GP G+ THVSTRVMGTYGY APEY M+G L+ KSD+YSFGVVLLE+LTG
Sbjct: 871 PKLSDFGLAKLGPVGDNTHVSTRVMGTYGYCAPEYAMSGKLTLKSDIYSFGVVLLELLTG 1050
Query: 344 RRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRD 403
RR+ID R GE NLV WARP RR F ++DP L+G F + +A + A CL
Sbjct: 1051RRAIDTSRRPGEQNLVSWARPYFSDRRRFGHMVDPLLQGRFPSRCLHQAIAITAMCLQEQ 1230
Query: 404 PKARPLMSEVVHTLKPL 420
PK RPL++++V L+ L
Sbjct: 1231PKFRPLITDIVVALEYL 1281
>CN825263
Length = 663
Score = 239 bits (610), Expect = 7e-64
Identities = 122/223 (54%), Positives = 156/223 (69%), Gaps = 3/223 (1%)
Frame = +1
Query: 147 GFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKL 206
GFG V+KG + + G VAVKIL + +G +E+LAE+ L L H NLVKL
Sbjct: 1 GFGLVYKGILND----------GRDVAVKILKRDDQRGGREFLAEVEMLSRLHHRNLVKL 150
Query: 207 IGFCIEDDQRLLVYEFMPRGSLENHLF---RRPLPLPWSIRMKIALGAAKGLAFLHEDSQ 263
IG CIE R L+YE +P GS+E+HL + PL W+ RMKIALGAA+GLA+LHEDS
Sbjct: 151 IGICIEKQTRCLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSN 330
Query: 264 RPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGH 323
+I+RDFK+SNILL+ ++ K+SDFGLA+ + H+ST VMGT+GY APEY MTGH
Sbjct: 331 PCVIHRDFKSSNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGH 510
Query: 324 LSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVL 366
L KSDVYS+GVVLLE+LTG + +D +P G+ NLV WARP+L
Sbjct: 511 LLVKSDVYSYGVVLLELLTGTKPVDLSQPPGQENLVTWARPIL 639
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 233 bits (595), Expect = 4e-62
Identities = 129/301 (42%), Positives = 186/301 (60%), Gaps = 3/301 (0%)
Frame = +3
Query: 121 SLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHN 180
S++ FT ++VAT ++ +L+GEGGFG V++G + + G VAVK+ +
Sbjct: 2121 SIQAFTLEYIEVATERYK--TLIGEGGFGSVYRGTLND----------GQEVAVKVRSST 2264
Query: 181 GHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLP-- 238
QG +E+ ELN L + H NLV L+G+C E DQ++LVY FM GSL++ L+ P
Sbjct: 2265 STQGTREFDNELNLLSAIQHENLVPLLGYCNESDQQILVYPFMSNGSLQDRLYGEPAKRK 2444
Query: 239 -LPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPE 297
L W R+ IALGAA+GLA+LH R +I+RD K+SNILLD AK++DFG +K P+
Sbjct: 2445 ILDWPTRLSIALGAARGLAYLHTFPGRSVIHRDIKSSNILLDHSMCAKVADFGFSKYAPQ 2624
Query: 298 GEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHN 357
++VS V GT GY PEY T LS KSDV+SFGVVLLE+++GR ++ KRP E +
Sbjct: 2625 EGDSYVSLEVRGTAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRTEWS 2804
Query: 358 LVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTL 417
LVEWA P + ++ +I+DP ++G + + + ++A QCL RP M +V L
Sbjct: 2805 LVEWATPYIRGSKV-DEIVDPGIKGGYHAEAMWRVVEVALQCLEPFSTYRPSMVAIVREL 2981
Query: 418 K 418
+
Sbjct: 2982 E 2984
>TC9461 similar to UP|O49840 (O49840) Protein kinase (AT2G02800/T20F6.6) ,
partial (37%)
Length = 484
Score = 231 bits (588), Expect = 3e-61
Identities = 112/161 (69%), Positives = 138/161 (85%), Gaps = 1/161 (0%)
Frame = +3
Query: 222 FMPRGSLENHLFRR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDA 280
FMP+GSLENHLFRR P PL WS+RMK+A+GAA+GL+FLH +++ +IYRDFK SNILLDA
Sbjct: 3 FMPKGSLENHLFRRGPQPLSWSVRMKVAIGAARGLSFLH-NAKSQVIYRDFKASNILLDA 179
Query: 281 EYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEM 340
E+NAKLSDFGLAK GP G++THVST+VMGT GYAAPEYV TG L++KSDVYSFGVVLLE+
Sbjct: 180 EFNAKLSDFGLAKAGPTGDRTHVSTQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLEL 359
Query: 341 LTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLE 381
L+GRR++DK + NLV+WARP LG +R F+I+D +LE
Sbjct: 360 LSGRRAVDKTIAGVDQNLVDWARPYLGDKRRLFRIMDSKLE 482
>TC12473 similar to UP|Q9M068 (Q9M068) Protein kinase-like protein, partial
(42%)
Length = 946
Score = 229 bits (583), Expect = 1e-60
Identities = 114/203 (56%), Positives = 147/203 (72%), Gaps = 1/203 (0%)
Frame = +3
Query: 98 TTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIE 157
T S S+ + K S E + L+ F+F LK AT++F+ ++L+GEGGFG V+KGW++
Sbjct: 339 TESESGSVVNNVKGGSGEFLEVAKLKVFSFGDLKSATKSFKADALIGEGGFGKVYKGWLD 518
Query: 158 ENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRL 217
E +P KPG+G+ VAVK LN QG EW +E+N+LG + HPNLVKL+G+C +D++ L
Sbjct: 519 EKKLSPTKPGSGIMVAVKKLNPESMQGFHEWQSEINFLGRISHPNLVKLLGYCRDDEEFL 698
Query: 218 LVYEFMPRGSLENHLFRRPL-PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNI 276
LVYEFMPRGSLENHLFRR L W+ R+KIA+GAA+GLAFLH + +IYRDFK SNI
Sbjct: 699 LVYEFMPRGSLENHLFRRNTESLSWNTRLKIAIGAARGLAFLH-SLXKIVIYRDFKASNI 875
Query: 277 LLDAEYNAKLSDFGLAKDGPEGE 299
LLD YNAK+S FGL K GP GE
Sbjct: 876 LLDGNYNAKISXFGLTKFGPSGE 944
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 224 bits (571), Expect = 2e-59
Identities = 135/349 (38%), Positives = 204/349 (57%), Gaps = 10/349 (2%)
Frame = +2
Query: 82 FEKNTEETEAPPESSTTTSNEESIASTP-KLFSEELKVASSLRKFTFNGLKVATRNFRPE 140
+ N + P + +++ S AS P K +++ + + + LK T NF +
Sbjct: 191 YPSNENDHLKSPRNYGDGNSKGSKASAPVKHETQKAPPPIEVPALSLDELKEKTDNFGSK 370
Query: 141 SLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGH-QGHKEWLAELNYLGDLL 199
+L+GEG +G V+ + + G VAVK L+ + + + E+L +++ + L
Sbjct: 371 ALIGEGSYGRVYYATLND----------GNAVAVKKLDVSSEPETNNEFLTQVSMVSRLK 520
Query: 200 HPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRR-------PLP-LPWSIRMKIALGA 251
+ N V+L G+C+E + R+L YEF GSL + L R P P L W R++IA+ A
Sbjct: 521 NDNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAVDA 700
Query: 252 AKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTY 311
A+GL +LHE Q II+RD ++SN+L+ +Y AK++DF L+ P+ STRV+GT+
Sbjct: 701 ARGLEYLHEKVQPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTRVLGTF 880
Query: 312 GYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRM 371
GY APEY MTG L+ KSDVYSFGVVLLE+LTGR+ +D P G+ +LV WA P L ++
Sbjct: 881 GYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVTWATPRLSEDKV 1060
Query: 372 FFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPL 420
Q +DP+L+G + KG K A +AA C+ + + RP MS VV L+PL
Sbjct: 1061-KQCVDPKLKGEYPPKGVAKLAAVAALCVQYEAEFRPNMSIVVKALQPL 1204
>AV417846
Length = 429
Score = 214 bits (545), Expect = 3e-56
Identities = 105/144 (72%), Positives = 124/144 (85%), Gaps = 3/144 (2%)
Frame = +1
Query: 222 FMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILL 278
FMP+GS+ENHLFRR P WS+R+KIALGAAKGLAFLH + +IYRDFKTSNILL
Sbjct: 1 FMPKGSMENHLFRRGSYFQPFSWSLRLKIALGAAKGLAFLHSTEPK-VIYRDFKTSNILL 177
Query: 279 DAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLL 338
D +Y+AKLSDFGLA+DGP G+K+HVSTRVMGT GYAAPEY+ TGHL++ SDVYSFGVVLL
Sbjct: 178 DTKYSAKLSDFGLARDGPVGDKSHVSTRVMGTRGYAAPEYLATGHLTANSDVYSFGVVLL 357
Query: 339 EMLTGRRSIDKKRPNGEHNLVEWA 362
E+++GRR+IDK + GEHNLVEWA
Sbjct: 358 EIISGRRAIDKNQLAGEHNLVEWA 429
>AV426813
Length = 431
Score = 211 bits (538), Expect = 2e-55
Identities = 103/144 (71%), Positives = 118/144 (81%), Gaps = 3/144 (2%)
Frame = +3
Query: 162 APVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYE 221
AP KPGTG +AVK LN +G QGH EWL E+NYLG L HPNLVKLIG+ IEDD R+LVYE
Sbjct: 3 APTKPGTGFVIAVKRLNQDGSQGHSEWLTEINYLGQLRHPNLVKLIGYSIEDDHRILVYE 182
Query: 222 FMPRGSLENHLFRRP---LPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILL 278
F+ +GSL+NHLFRR PL W+IRMKIAL AAKGLAFLH D + +IYRDFKTSNIL+
Sbjct: 183 FLAKGSLDNHLFRRASYFQPLSWNIRMKIALDAAKGLAFLHSD-EVEVIYRDFKTSNILI 359
Query: 279 DAEYNAKLSDFGLAKDGPEGEKTH 302
D+ YNAKLSDFGLAKDGP G+K+H
Sbjct: 360 DSNYNAKLSDFGLAKDGPAGDKSH 431
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 207 bits (526), Expect = 4e-54
Identities = 120/290 (41%), Positives = 175/290 (59%), Gaps = 1/290 (0%)
Frame = +3
Query: 142 LLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHP 201
LLG GGFG V+ G ++ G VA+K N QG E+ E+ L L H
Sbjct: 12 LLGVGGFGKVYYGEVDG----------GTKVAIKRGNPLSEQGVHEFQTEIEMLSKLRHR 161
Query: 202 NLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFR-RPLPLPWSIRMKIALGAAKGLAFLHE 260
+LV LIG+C E+ + +LVY+ M G+L HL++ + PLPW R++I +GAA+GL +LH
Sbjct: 162 HLVSLIGYCEENTEMILVYDHMAYGTLREHLYKTQKPPLPWKQRLEICIGAARGLHYLHT 341
Query: 261 DSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVM 320
++ II+RD KT+NILLD ++ AK+SDFGL+K GP + THVST V G++GY PEY
Sbjct: 342 GAKYTIIHRDVKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFR 521
Query: 321 TGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRL 380
L+ KSDVYSFGVVL E+L R +++ + +L EWA ++ + QI+DP L
Sbjct: 522 RQQLTDKSDVYSFGVVLFEILCARPALNPSLAKEQVSLAEWASHCY-NKGILDQILDPYL 698
Query: 381 EGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISS 430
+G + + +K A+ A +C+S RP M +V+ L+ L++ A S
Sbjct: 699 KGKIAPECFKKFAETAMKCVSDQGIERPSMGDVLWNLEFALQLQESAEES 848
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 206 bits (524), Expect = 7e-54
Identities = 115/275 (41%), Positives = 164/275 (58%), Gaps = 3/275 (1%)
Frame = +2
Query: 95 SSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKG 154
SS + E + P F V +S R FT+ L AT F ++ LGEGGFG V+ G
Sbjct: 185 SSFSCCGSEKVEEGPTSFGS---VNNSWRIFTYKELHAATGGFSDDNKLGEGGFGSVYWG 355
Query: 155 WIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDD 214
+ GL +AVK L + E+ E+ LG + H NL+ L G+C+ DD
Sbjct: 356 ----------RTSDGLQIAVKKLKAMNSKAEMEFAVEVEVLGRVRHKNLLGLRGYCVGDD 505
Query: 215 QRLLVYEFMPRGSLENHL---FRRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDF 271
QRL+VY++MP SL +HL F + L W RMKIA+G+A+G+ +LH + II+RD
Sbjct: 506 QRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIAIGSAEGILYLHHEVTPHIIHRDI 685
Query: 272 KTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVY 331
K SN+LL++++ ++DFG AK PEG +H++TRV GT GY APEY M G +S DVY
Sbjct: 686 KASNVLLNSDFEPLVADFGFAKLIPEGV-SHMTTRVKGTLGYLAPEYAMWGKVSESCDVY 862
Query: 332 SFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVL 366
SFG++LLE++TGR+ I+K + + EWA P++
Sbjct: 863 SFGILLLELVTGRKPIEKLPGGVKRTITEWAEPLI 967
>TC9462 similar to UP|O49840 (O49840) Protein kinase (AT2G02800/T20F6.6) ,
partial (37%)
Length = 576
Score = 202 bits (513), Expect = 1e-52
Identities = 110/211 (52%), Positives = 138/211 (65%), Gaps = 3/211 (1%)
Frame = +1
Query: 42 FFGSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSN 101
F G+C+ S KVD++ S ST + SG K T P S ++ +
Sbjct: 1 FMGNCLDSSAKVDAAQSSRST---------------SASGISKTT-----PSSISISSYS 120
Query: 102 EESIASTPKLFSEELKVASS--LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEEN 159
E+S AS+ E ++ SS L+ FTFN LK ATRNFRP+SLLGEGGFG V+KGWI+E+
Sbjct: 121 EKSNASSLPTPRSEGEILSSPNLKAFTFNELKNATRNFRPDSLLGEGGFGYVYKGWIDEH 300
Query: 160 GTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLV 219
KPG+G+ VAVK L G QGHKEWL E+NYLG L HPNLVKLIG+C++ + RLLV
Sbjct: 301 SFTAAKPGSGMVVAVKRLKPEGFQGHKEWLTEVNYLGQLYHPNLVKLIGYCLDGENRLLV 480
Query: 220 YEFMPRGSLENHLFRR-PLPLPWSIRMKIAL 249
YEFMP+GSL+NHLF R P PL WS R K+A+
Sbjct: 481 YEFMPKGSLDNHLFTRGPQPLSWSXRXKLAI 573
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 193 bits (491), Expect = 5e-50
Identities = 106/265 (40%), Positives = 160/265 (60%), Gaps = 3/265 (1%)
Frame = +1
Query: 123 RKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGH 182
R F+ L AT NF ++ LGEGGFG V+ G + + G +AVK L +
Sbjct: 22 RVFSLKELHSATNNFNYDNKLGEGGFGSVYWGQLWD----------GSQIAVKRLKVWSN 171
Query: 183 QGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLP---L 239
+ E+ E+ L + H NL+ L G+C E +RL+VY++MP SL +HL + L
Sbjct: 172 KADMEFAVEVEILARVRHKNLLSLRGYCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLL 351
Query: 240 PWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGE 299
W+ RM IA+G+A+G+ +LH + II+RD K SN+LLD+++ A+++DFG AK P+G
Sbjct: 352 DWNRRMNIAIGSAEGIVYLHHQATPHIIHRDIKASNVLLDSDFQARVADFGFAKLIPDGA 531
Query: 300 KTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLV 359
THV+TRV GT GY APEY M G + DV+SFG++LLE+ +G++ ++K + ++
Sbjct: 532 -THVTTRVKGTLGYLAPEYAMLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSIN 708
Query: 360 EWARPVLGHRRMFFQIIDPRLEGHF 384
+WA P L + F + DPRL G +
Sbjct: 709 DWALP-LACAKKFTEFADPRLNGEY 780
>AI967315
Length = 1308
Score = 189 bits (480), Expect = 9e-49
Identities = 119/322 (36%), Positives = 181/322 (55%), Gaps = 4/322 (1%)
Frame = +1
Query: 101 NEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENG 160
N++S P EE S + F++ L AT F E+++G+GG+ V+KG +E
Sbjct: 46 NDDSSLVVPPY--EEHPPRPSWKCFSYEELFHATNGFSSENMVGKGGYAEVYKGRLE--- 210
Query: 161 TAPVKPGTGLTVAVKILNHN--GHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLL 218
+G +AVK L + KE+L E+ +G + H N++ L+G CI D+ L
Sbjct: 211 -------SGDEIAVKRLTRTCRDERKEKEFLTEIGTIGHVCHSNVMPLLGCCI-DNGLYL 366
Query: 219 VYEFMPRGSLENHLFRRPL-PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNIL 277
V+E GS+ + + + PL W R KI LG A+GL +LH+ QR II+RD K SNIL
Sbjct: 367 VFELSTVGSVASLIHDEKMAPLDWKTRYKIVLGTARGLHYLHKGCQRRIIHRDIKASNIL 546
Query: 278 LDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVL 337
L ++ ++SDFGLAK P H + GT+G+ APEY M G + K+DV++FGV L
Sbjct: 547 LTEDFEPQISDFGLAKWLPSQWTHHSIAPIEGTFGHLAPEYYMHGVVDEKTDVFAFGVFL 726
Query: 338 LEMLTGRRSIDKKRPNGEH-NLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLA 396
LE+++GR+ +D G H +L WA+P+L + +++DPRLEG + V + A A
Sbjct: 727 LEVISGRKPVD-----GSHQSLHTWAKPILS-KWEIEKLVDPRLEGCYDVTQFNRVAFAA 888
Query: 397 AQCLSRDPKARPLMSEVVHTLK 418
+ C+ RP MSEV+ ++
Sbjct: 889 SLCIRASSTWRPTMSEVLEVME 954
>TC20060 similar to UP|Q9LZF8 (Q9LZF8) Protein kinase-like (Protein
serine/threonine kinase), partial (35%)
Length = 482
Score = 177 bits (449), Expect = 3e-45
Identities = 91/148 (61%), Positives = 110/148 (73%), Gaps = 5/148 (3%)
Frame = +1
Query: 172 VAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDD----QRLLVYEFMPRGS 227
VAVK L G QGHKEW+ E+N LG + HPNLVKL+G+C +DD QRLL+YE+MP S
Sbjct: 37 VAVKQLGRRGIQGHKEWVTEVNVLGIVEHPNLVKLVGYCADDDERGIQRLLIYEYMPNRS 216
Query: 228 LENHLF-RRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKL 286
+E+HL R PLPWS R+KIA AA+GL +LHE+ II+RDFK+SNILLD +NAKL
Sbjct: 217 VEHHLSPRSETPLPWSRRLKIAQDAARGLTYLHEEMDFQIIFRDFKSSNILLDEHWNAKL 396
Query: 287 SDFGLAKDGPEGEKTHVSTRVMGTYGYA 314
SDFGLA+ GP THVST V+GT GYA
Sbjct: 397 SDFGLARLGPAEGLTHVSTAVVGTMGYA 480
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 177 bits (449), Expect = 3e-45
Identities = 126/350 (36%), Positives = 181/350 (51%), Gaps = 8/350 (2%)
Frame = +1
Query: 94 ESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFK 153
E T+ S+ AS L S + VA S+ +F++ L AT NF ++ +G+GGFG V+
Sbjct: 967 EYETSGSSGPGTASATGLTS--IMVAKSM-EFSYQELAKATNNFSLDNKIGQGGFGAVYY 1137
Query: 154 GWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIED 213
+ TA K Q E+L EL L + H NLV+LIG+C+E
Sbjct: 1138 AELRGKKTAIKKMDV--------------QASTEFLCELKVLTHVHHLNLVRLIGYCVEG 1275
Query: 214 DQRLLVYEFMPRGSLENHLFRRPL-PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFK 272
LVYE + G+L +L PLPWS R++IAL AA+GL ++HE + I+RD K
Sbjct: 1276 SL-FLVYEHIDNGNLGQYLHGSGKEPLPWSSRVQIALDAARGLEYIHEHTVPVYIHRDVK 1452
Query: 273 TSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYS 332
++NIL+D K++DFGL K G T + TR++GT+GY PEY G +S K DVY+
Sbjct: 1453 SANILIDKNLRGKVADFGLTKLIEVGNST-LQTRLVGTFGYMPPEYAQYGDISPKIDVYA 1629
Query: 333 FGVVLLEMLTGRRSIDK-----KRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVK 387
FGVVL E+++ + ++ K G L E A +++DPRL ++ +
Sbjct: 1630 FGVVLFELISAKNAVLKTGELVAESKGLVALFEEALNKSDPCDALRKLVDPRLGENYPID 1809
Query: 388 GAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMA--ISSYHFQT 435
K AQL C +P RP M +V L L +L + SSY QT
Sbjct: 1810 SVLKIAQLGRACTRDNPLLRPSMRSLVVALMTLSSLTEDCDDESSYESQT 1959
>TC11816 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B , partial
(32%)
Length = 513
Score = 175 bits (444), Expect = 1e-44
Identities = 89/167 (53%), Positives = 113/167 (67%), Gaps = 2/167 (1%)
Frame = +3
Query: 63 HN-GNSIKSVITQSEENKSGFEKNTEETEAPPES-STTTSNEESIASTPKLFSEELKVAS 120
HN G + + I N +G T S + + + S+A TP+ E L+ +S
Sbjct: 15 HNMGACLSAQIKAESPNNTGLSSKNVSTNGNDLSCANSNGSAASVAQTPRSEGEILQ-SS 191
Query: 121 SLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHN 180
+++ FTF LK ATRNFRP+S+LGEGGFG VFKGWI+EN A KPG G+ +AVK LNH
Sbjct: 192 NVKSFTFIELKTATRNFRPDSVLGEGGFGSVFKGWIDENTLAAAKPGIGIGIAVKRLNHE 371
Query: 181 GHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGS 227
QGH+EWLAE+NYLG HP+LVKLIG+C+ED+ RLLVYEFMPRGS
Sbjct: 372 SFQGHREWLAEVNYLGQFSHPHLVKLIGYCLEDEHRLLVYEFMPRGS 512
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 172 bits (435), Expect = 1e-43
Identities = 111/297 (37%), Positives = 163/297 (54%), Gaps = 13/297 (4%)
Frame = +2
Query: 138 RPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEW--LAELNYL 195
+ E+++G+GG G V++G + NGT VA+K L G G ++ AE+ L
Sbjct: 2195 KEENIIGKGGAGIVYRGSMP-NGT---------DVAIKRLVGQG-SGRNDYGFRAEIETL 2341
Query: 196 GDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLF-RRPLPLPWSIRMKIALGAAKG 254
G + H N+++L+G+ D LL+YE+MP GSL L + L W +R KIA+ AA+G
Sbjct: 2342 GKIRHRNIMRLLGYVSNKDTNLLLYEYMPNGSLGEWLHGAKGGHLRWEMRYKIAVEAARG 2521
Query: 255 LAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYA 314
L ++H D II+RD K++NILLDA++ A ++DFGLAK + + + + G+YGY
Sbjct: 2522 LCYMHHDCSPLIIHRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAGSYGYI 2701
Query: 315 APEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEH----NLVEWARPVLGH-- 368
APEY T + KSDVYSFGVVLLE++ GR +P GE ++V W +
Sbjct: 2702 APEYAYTLKVDEKSDVYSFGVVLLELIIGR------KPVGEFGDGVDIVGWVNKTMSELS 2863
Query: 369 ----RRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLP 421
+ ++DPRL G + + +A C+ ARP M EVVH L P
Sbjct: 2864 QPSDTALVLAVVDPRLSG-YPLTSVIHMFNIAMMCVKEMGPARPTMREVVHMLTNPP 3031
>BP032158
Length = 555
Score = 170 bits (431), Expect = 4e-43
Identities = 93/191 (48%), Positives = 122/191 (63%), Gaps = 3/191 (1%)
Frame = +2
Query: 125 FTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQG 184
F+ +K AT NF P + +GEGGFG V+KG + E G +AVK L+ QG
Sbjct: 14 FSLRQIKAATNNFDPANKIGEGGFGPVYKGVLSE----------GDVIAVKQLSSKSKQG 163
Query: 185 HKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLF---RRPLPLPW 241
++E++ E+ + L HPNLVKL G CIE +Q LLVYE+M SL LF + L L W
Sbjct: 164 NREFINEIGMISALQHPNLVKLYGCCIEGNQLLLVYEYMENNSLARALFGNEEQKLNLNW 343
Query: 242 SIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKT 301
RMKI +G AKGLA+LHE+S+ I++RD K +N+LLD + AK+SDFGLAK E E T
Sbjct: 344 RTRMKICVGIAKGLAYLHEESRLKIVHRDIKATNVLLDKDLKAKISDFGLAKLDEE-ENT 520
Query: 302 HVSTRVMGTYG 312
H+STR+ GT G
Sbjct: 521 HISTRIAGTIG 553
>TC16403 homologue to UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, partial
(55%)
Length = 876
Score = 164 bits (415), Expect = 3e-41
Identities = 85/164 (51%), Positives = 114/164 (68%)
Frame = +3
Query: 237 LPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGP 296
L L W+ R+KIA+GAA+GL +LHE ++ II+R K+SNILL + AK++DF L+ P
Sbjct: 30 LVLSWTQRVKIAVGAARGLEYLHEKAETHIIHRYIKSSNILLFDDDVAKIADFDLSNQAP 209
Query: 297 EGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEH 356
+ STRV+GT+GY APEY MTG L+SKSDVYSFGVVLLE+LTGR+ +D P G+
Sbjct: 210 DAAARLHSTRVLGTFGYHAPEYAMTGQLTSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQ 389
Query: 357 NLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCL 400
+LV WA P L ++ Q +D RL+G + VK K A +AA C+
Sbjct: 390 SLVTWATPKLSEDKV-KQCVDARLKGEYPVKSVAKMAAVAALCV 518
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,512,787
Number of Sequences: 28460
Number of extensions: 122883
Number of successful extensions: 1172
Number of sequences better than 10.0: 329
Number of HSP's better than 10.0 without gapping: 996
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1004
length of query: 490
length of database: 4,897,600
effective HSP length: 94
effective length of query: 396
effective length of database: 2,222,360
effective search space: 880054560
effective search space used: 880054560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0391.11


