

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388b.3
(149 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
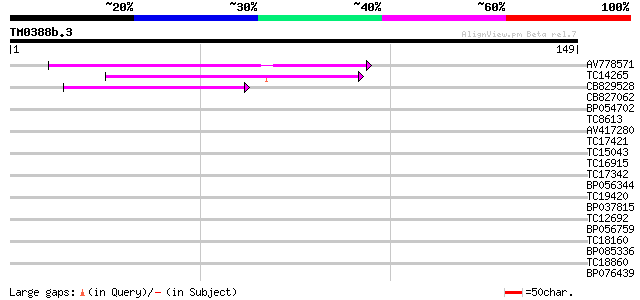
Score E
Sequences producing significant alignments: (bits) Value
AV778571 55 5e-09
TC14265 weakly similar to UP|AAN62760 (AAN62760) Disease resista... 44 2e-05
CB829528 41 8e-05
CB827062 31 0.10
BP054702 29 0.39
TC8613 similar to GB|AAC64889.1|3776572|T22H22 ESTs gb|R65052, g... 28 0.86
AV417280 27 1.5
TC17421 similar to UP|Q9M205 (Q9M205) Regulatory protein-like, p... 27 1.5
TC15043 weakly similar to UP|AAR87280 (AAR87280) Expressed prote... 27 1.5
TC16915 weakly similar to UP|Q9SDA8 (Q9SDA8) AT2G17020 protein (... 27 1.9
TC17342 similar to PIR|JC7226|JC7226 endo-1,3(4)-beta-glucanase ... 27 1.9
BP056344 26 2.5
TC19420 26 2.5
BP037815 26 3.3
TC12692 similar to UP|Q8H2C6 (Q8H2C6) Cellulose synthase, partia... 26 3.3
BP056759 25 4.3
TC18160 homologue to UP|Q9ZP40 (Q9ZP40) Plastoglobule associated... 25 4.3
BP085336 25 5.6
TC18860 similar to UP|Q940L4 (Q940L4) AT3g62590/F26K9_20, partia... 25 5.6
BP076439 25 5.6
>AV778571
Length = 452
Score = 55.1 bits (131), Expect = 5e-09
Identities = 29/85 (34%), Positives = 46/85 (54%)
Frame = +3
Query: 11 SNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGT 70
S + + + P+ G LE+L F NL+++H S+G L +L L +EGC + +
Sbjct: 39 SECESITQIPDLSGLPNLEKLSFVYSENLIKIHESVGFLNKLRILDIEGCCKIRTF---P 209
Query: 71 VCELNSLRILHLSGCTRLESTPNFI 95
+L SL L+LS C+ LES P +
Sbjct: 210 PMKLASLEELYLSHCSSLESFPEIL 284
Score = 25.0 bits (53), Expect = 5.6
Identities = 20/61 (32%), Positives = 28/61 (45%), Gaps = 2/61 (3%)
Frame = +3
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGC--TNLLQVHPSIGLLTELAFLSLEGCSS 62
L+ L LS+ L P ++E + G T + ++ SIG LT L L L GC
Sbjct: 228 LEELYLSHCSSLESFPEI--LAKMENITLIGVYHTPIKELPSSIGNLTRLRRLELHGCGM 401
Query: 63 L 63
L
Sbjct: 402 L 404
>TC14265 weakly similar to UP|AAN62760 (AAN62760) Disease resistance
protein-like protein MsR1, partial (5%)
Length = 766
Score = 43.5 bits (101), Expect = 2e-05
Identities = 26/70 (37%), Positives = 37/70 (52%), Gaps = 2/70 (2%)
Frame = +1
Query: 26 QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSL--YLGTVCELNSLRILHLS 83
+ LE L + CT+L + SIG L++L L + C SL SL +G +C L SL + +
Sbjct: 19 ENLELLRLSSCTDLKGLPDSIGRLSKLRLLDISNCISLPSLPEEIGNLCNLKSLYMTSCA 198
Query: 84 GCTRLESTPN 93
GC S N
Sbjct: 199GCELPSSIVN 228
Score = 36.2 bits (82), Expect = 0.002
Identities = 23/70 (32%), Positives = 34/70 (47%)
Frame = +1
Query: 46 IGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGF 105
IG L L L L C+ L L ++ L+ LR+L +S C L S P IGN + + +
Sbjct: 7 IGKLENLELLRLSSCTDLKGLP-DSIGRLSKLRLLDISNCISLPSLPEEIGNLCNLKSLY 183
Query: 106 DIVVPGNKIP 115
G ++P
Sbjct: 184 MTSCAGCELP 213
>CB829528
Length = 188
Score = 41.2 bits (95), Expect = 8e-05
Identities = 18/49 (36%), Positives = 26/49 (52%)
Frame = +2
Query: 15 YLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
+L E PN + L L C +L+ +H S+G L L LS +GC+ L
Sbjct: 2 FLTELPNLSAAPFLRNLSLDNCPSLVTIHESVGFLENLRSLSAKGCTQL 148
Score = 30.8 bits (68), Expect = 0.10
Identities = 18/46 (39%), Positives = 28/46 (60%)
Frame = +2
Query: 44 PSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLE 89
P++ L LSL+ C SLV+++ +V L +LR L GCT+L+
Sbjct: 17 PNLSAAPFLRNLSLDNCPSLVTIH-ESVGFLENLRSLSAKGCTQLK 151
>CB827062
Length = 245
Score = 30.8 bits (68), Expect = 0.10
Identities = 14/43 (32%), Positives = 20/43 (45%)
Frame = +2
Query: 107 IVVPGNKIPYWFDSQFKGGSRIREVDHVYEQDHWLGFAFCAVF 149
IV+PG + P WF S ++ V +W+G A C F
Sbjct: 14 IVIPGGEFPRWFSELNVVYSIRGDLSPVMPNQNWIGLACCFTF 142
>BP054702
Length = 528
Score = 28.9 bits (63), Expect = 0.39
Identities = 19/45 (42%), Positives = 22/45 (48%)
Frame = -3
Query: 48 LLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLESTP 92
L T L FL L SSL L + L+SL LH+ C L S P
Sbjct: 400 LPTSLVFLGLYNFSSLKFLEGNGLQHLSSLHQLHIDKCPSLVSLP 266
>TC8613 similar to GB|AAC64889.1|3776572|T22H22 ESTs gb|R65052,
gb|AA712146, gb|H76533, gb|H76282, gb|AA650771,
gb|H76287, gb|AA650887, gb|N37383, gb|Z29721 and
gb|Z29722 come from this gene. {Arabidopsis thaliana;},
partial (68%)
Length = 939
Score = 27.7 bits (60), Expect = 0.86
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = -3
Query: 118 FDSQFKGGSRIREVDHVYEQDHWLGFAFCAVF 149
FD+ S +R + +E++ W GF FC VF
Sbjct: 112 FDAGEWRESHVRRKNGSHEEEDWEGFNFCRVF 17
>AV417280
Length = 427
Score = 26.9 bits (58), Expect = 1.5
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 2/59 (3%)
Frame = +3
Query: 27 RLERLDFTGCT--NLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLS 83
R+ LD +G + + + PS+G L L LSL +L+ + +L SL +++S
Sbjct: 219 RVNDLDLSGFSLPSPHPIPPSVGDLPHLNILSLRNIPNLIGPIPSAITKLTSLHYIYIS 395
>TC17421 similar to UP|Q9M205 (Q9M205) Regulatory protein-like, partial
(52%)
Length = 548
Score = 26.9 bits (58), Expect = 1.5
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = -3
Query: 11 SNSKYLMETPNFDGSQRLERLDFTGCTNLL 40
S+S++ + P F S + ERL GC NLL
Sbjct: 294 SSSQFSLFWPCFSQSVKRERLSLMGCWNLL 205
>TC15043 weakly similar to UP|AAR87280 (AAR87280) Expressed protein (With
alternative splicing), partial (10%)
Length = 1269
Score = 26.9 bits (58), Expect = 1.5
Identities = 17/48 (35%), Positives = 22/48 (45%)
Frame = +2
Query: 54 FLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHF 101
F+SL S L +YL C + LHL C + E+T G P F
Sbjct: 815 FVSLLSSSILCGVYLFP*CTIFISSQLHLPFCEKTETTTLVRGGPHIF 958
>TC16915 weakly similar to UP|Q9SDA8 (Q9SDA8) AT2G17020 protein
(At2g17020/At2g17020), partial (29%)
Length = 1011
Score = 26.6 bits (57), Expect = 1.9
Identities = 24/64 (37%), Positives = 32/64 (49%), Gaps = 3/64 (4%)
Frame = +3
Query: 28 LERLDFTGCTNLLQ-VHPSIGLLTELAFLSLEGCSSLVSLYLGTV--CELNSLRILHLSG 84
L+ LD C +L +IG LT+L L L+G S + L + +NSL L L G
Sbjct: 54 LKILDLRYCRSLGDDALQAIGTLTKLKVLLLDG-SDITDAGLSCLRPSVINSLYALSLRG 230
Query: 85 CTRL 88
C RL
Sbjct: 231 CKRL 242
>TC17342 similar to PIR|JC7226|JC7226 endo-1,3(4)-beta-glucanase - garden
pea {Pisum sativum;} , partial (24%)
Length = 499
Score = 26.6 bits (57), Expect = 1.9
Identities = 21/71 (29%), Positives = 32/71 (44%), Gaps = 8/71 (11%)
Frame = -1
Query: 52 LAFLSLEG---CSSLVSLYLGTVCELNSLRILHLSGCTRLESTP-NFIGNPSHFRCGFD- 106
+ +L++ G CSSL + G + +R T L++TP G H + D
Sbjct: 496 IPYLNINGVWICSSLEKIISGPANRFSGIRQFTFHHPTELKNTPRKHGGGEGHGKAKLDV 317
Query: 107 ---IVVPGNKI 114
IVVP N+I
Sbjct: 316 VPCIVVPSNQI 284
>BP056344
Length = 397
Score = 26.2 bits (56), Expect = 2.5
Identities = 14/39 (35%), Positives = 18/39 (45%)
Frame = -2
Query: 20 PNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLE 58
P F + LE ++ C N L VH L FLSL+
Sbjct: 384 PRFSKASELEEIELYACRNALSVH--------LPFLSLD 292
>TC19420
Length = 532
Score = 26.2 bits (56), Expect = 2.5
Identities = 13/28 (46%), Positives = 18/28 (63%), Gaps = 1/28 (3%)
Frame = +2
Query: 72 CELNSLRILHLSGC-TRLESTPNFIGNP 98
C +L+ +L+ C TRL+ TP F GNP
Sbjct: 113 CTFETLKPPNLANCFTRLQITPIFGGNP 196
>BP037815
Length = 546
Score = 25.8 bits (55), Expect = 3.3
Identities = 13/43 (30%), Positives = 23/43 (53%)
Frame = +3
Query: 42 VHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSG 84
+ PSI LT+L+ +++ + G + L SL+ + LSG
Sbjct: 366 ISPSICNLTQLSSITISDWKGISGTIPGCITTLRSLQTIDLSG 494
>TC12692 similar to UP|Q8H2C6 (Q8H2C6) Cellulose synthase, partial (4%)
Length = 632
Score = 25.8 bits (55), Expect = 3.3
Identities = 15/44 (34%), Positives = 23/44 (52%), Gaps = 2/44 (4%)
Frame = -2
Query: 56 SLEGCSSLVSLYLGTVCELNSLRILH--LSGCTRLESTPNFIGN 97
S E C + L+LG + +L S+ LH L GC + P F+ +
Sbjct: 490 SSESCFIVSLLHLGFIAQLGSVSFLHFPLLGC--FDPAPGFMAH 365
>BP056759
Length = 551
Score = 25.4 bits (54), Expect = 4.3
Identities = 11/41 (26%), Positives = 19/41 (45%)
Frame = -1
Query: 98 PSHFRCGFDIVVPGNKIPYWFDSQFKGGSRIREVDHVYEQD 138
P+ RCG + + +WF ++ G R RE ++D
Sbjct: 242 PASTRCGSSWTRNRSPVRWWFGEHWRSGERKRERPRRNDED 120
>TC18160 homologue to UP|Q9ZP40 (Q9ZP40) Plastoglobule associated protein
PG1 precursor, partial (15%)
Length = 458
Score = 25.4 bits (54), Expect = 4.3
Identities = 14/43 (32%), Positives = 23/43 (52%)
Frame = +3
Query: 66 LYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIV 108
L++ + C L +L LSGC+ + S + +P HF G I+
Sbjct: 282 LFISSCCILIG-HLLLLSGCSSINSCWRYAQSPVHFVGGHMII 407
>BP085336
Length = 369
Score = 25.0 bits (53), Expect = 5.6
Identities = 11/15 (73%), Positives = 13/15 (86%)
Frame = -2
Query: 76 SLRILHLSGCTRLES 90
SL IL+LSGC+RL S
Sbjct: 245 SLTILYLSGCSRLTS 201
>TC18860 similar to UP|Q940L4 (Q940L4) AT3g62590/F26K9_20, partial (29%)
Length = 606
Score = 25.0 bits (53), Expect = 5.6
Identities = 15/56 (26%), Positives = 22/56 (38%), Gaps = 4/56 (7%)
Frame = -3
Query: 69 GTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIV----VPGNKIPYWFDS 120
G +C L + CTR + N G+ F C I + N +P+W S
Sbjct: 199 GNICWYIPLAAS*IPLCTRTSNPSNLTGSKRRFACQDAIDSEP*ITKNLVPFWSSS 32
>BP076439
Length = 478
Score = 25.0 bits (53), Expect = 5.6
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +1
Query: 66 LYLGTVCELNSLRILHL 82
L L T+C LNS RI+H+
Sbjct: 223 LLLNTLCPLNSQRIVHI 273
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.143 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,275,066
Number of Sequences: 28460
Number of extensions: 51299
Number of successful extensions: 296
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 289
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 294
length of query: 149
length of database: 4,897,600
effective HSP length: 82
effective length of query: 67
effective length of database: 2,563,880
effective search space: 171779960
effective search space used: 171779960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0388b.3


