

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388b.1
(949 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
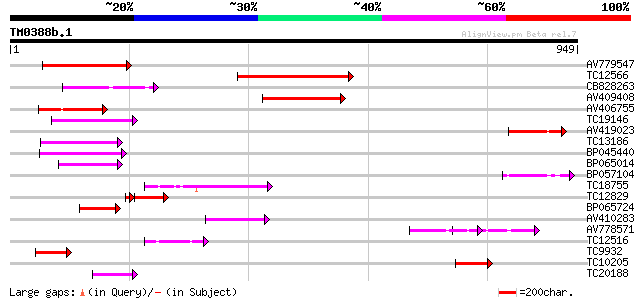
Score E
Sequences producing significant alignments: (bits) Value
AV779547 160 7e-40
TC12566 similar to UP|Q84XI1 (Q84XI1) RCa1 (Fragment), partial (... 150 1e-36
CB828263 135 2e-32
AV409408 130 1e-30
AV406755 127 1e-29
TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-contain... 119 2e-27
AV419023 117 9e-27
TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidat... 108 5e-24
BP045440 103 1e-22
BP065014 87 1e-17
BP057104 82 5e-16
TC18755 similar to UP|AAN85378 (AAN85378) Resistance protein (Fr... 69 3e-12
TC12829 67 3e-12
BP065724 69 3e-12
AV410283 67 2e-11
AV778571 64 1e-10
TC12516 similar to UP|Q9M5V7 (Q9M5V7) Disease resistance-like pr... 60 1e-09
TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like... 59 4e-09
TC10205 weakly similar to UP|Q8RL77 (Q8RL77) HpaG, partial (10%) 54 9e-08
TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 52 4e-07
>AV779547
Length = 556
Score = 160 bits (406), Expect = 7e-40
Identities = 79/149 (53%), Positives = 99/149 (66%)
Frame = +1
Query: 56 ISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSIVVFSK 115
+SFRG DTRN F DHL L KG FKDD L+KG++IS +L+QAI S++ IVVFSK
Sbjct: 109 VSFRGEDTRNNFTDHLLGALYGKGFVTFKDDTMLRKGKNISTELIQAIEGSQILIVVFSK 288
Query: 116 NYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRFKHDA 175
YA S WCL E+A IA+ +QTV PV DV PS VR Q+G Y AF+ H RFK D
Sbjct: 289 YYASSTWCLQELAKIADGIVGKRQTVLPVVCDVTPSEVRKQSGNYGEAFLKHEERFKEDL 468
Query: 176 DRVDRWKRAMRSLAGSAGWDVRNKPEFRE 204
+ V RW++A+ +A +GWDV NKPE +
Sbjct: 469 EMVQRWRKALAEVADLSGWDVTNKPEHEQ 555
>TC12566 similar to UP|Q84XI1 (Q84XI1) RCa1 (Fragment), partial (28%)
Length = 918
Score = 150 bits (378), Expect = 1e-36
Identities = 76/195 (38%), Positives = 120/195 (60%)
Frame = +1
Query: 381 VYEVPLMNNNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRNA 440
+YEV M+ D+ +LF FK ++ +L ++L YA+G+PLA++V G FL R
Sbjct: 10 IYEVKEMSFEDSLQLFSLNAFKRNDPLETYIDLSKKLLSYAKGIPLALKVLGLFLYGRER 189
Query: 441 MQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIACFFKGEKENYVKRILDACG 500
W L +L+ PD ++++VL++S++GL + K+IFL IACF+ G E V + LD+CG
Sbjct: 190 KAWESQLVKLEKLPDPEIINVLKLSYDGLDDDQKDIFLDIACFYLGHLEESVAQTLDSCG 369
Query: 501 LHPHIGIQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFPEEPGSWSRLWLYQHFHHV 560
IG++ + +R LI+I I MH++++ +GK+IVRQQ P +PG SRLW + V
Sbjct: 370 FFADIGMEVLKDRCLISILEGRIVMHDLIEVMGKEIVRQQCPNDPGKRSRLWNDDQTYDV 549
Query: 561 LMSEMGTNKVKAIVL 575
L GT+ ++ I L
Sbjct: 550 LGKNKGTDAIQCIFL 594
Score = 38.1 bits (87), Expect = 0.007
Identities = 21/49 (42%), Positives = 29/49 (58%)
Frame = +2
Query: 611 SLHFLSNNLQYLLWHGYPFASLPSNFEPFRLVELNMPYSSIQRLWEGRK 659
SL L N+LQ+L W G+P P +F P LV+L+M S ++ GRK
Sbjct: 722 SLGSLPNHLQFLRWDGFPLRYFPLDFCPQNLVKLDMRDSHLEHF--GRK 862
>CB828263
Length = 570
Score = 135 bits (341), Expect = 2e-32
Identities = 70/162 (43%), Positives = 97/162 (59%), Gaps = 1/162 (0%)
Frame = +2
Query: 89 LQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDV 148
LQ+G+ IS+ LL+AI +++S++VFSKNY S+WCLDE+ I EC Q V PVFYD+
Sbjct: 8 LQRGDEISSSLLRAIEEAKLSVIVFSKNYGNSKWCLDELVKILECKRTRGQIVLPVFYDI 187
Query: 149 DPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDVR-NKPEFREIEN 207
DPS VRNQ G Y AFV H D+V +W+ A+R A +GWD N+ E IE
Sbjct: 188 DPSHVRNQTGTYAEAFVKH-----GQVDKVQKWREALREAANLSGWDCSVNRMESEVIEK 352
Query: 208 IVEAVIEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYD 249
I + V+E L R + G D+ ++ E L +L E+Y+
Sbjct: 353 IAKDVLEKLNRVYVGDLDE------KIAKFEQLAQLQREFYE 460
>AV409408
Length = 430
Score = 130 bits (326), Expect = 1e-30
Identities = 62/139 (44%), Positives = 93/139 (66%)
Frame = +2
Query: 423 GLPLAIRVTGSFLCTRNAMQWRDALDRLKNNPDNKVMDVLQISFEGLHSEDKEIFLHIAC 482
GLPLA+ V GSFLC R+ W DAL ++K P + +++ L+IS++ L E K +FL IAC
Sbjct: 5 GLPLALEVLGSFLCGRSLSDWEDALIKIKKVPHDDILNKLRISYDMLEDEHKTLFLDIAC 184
Query: 483 FFKGEKENYVKRILDACGLHPHIGIQNMIERSLITIRNQEIHMHEMVQDLGKKIVRQQFP 542
FFKG ++ V ++L +CGLHP +GI +IE+SL+T + I MH++++++GK V + P
Sbjct: 185 FFKGWYKHKVIQLLGSCGLHPTVGINVLIEKSLLTFDGRVIGMHDLLEEMGKTEVFLESP 364
Query: 543 EEPGSWSRLWLYQHFHHVL 561
+PG SRLW + VL
Sbjct: 365 NDPGRRSRLWSLEDIDQVL 421
>AV406755
Length = 425
Score = 127 bits (318), Expect = 1e-29
Identities = 60/116 (51%), Positives = 80/116 (68%)
Frame = +3
Query: 49 RYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRV 108
++ YDVF+SFRG ++R +F HLY L GI VF D++ LQ+GE IS+ LL+AI +SR+
Sbjct: 75 KWSYDVFLSFRGEESRRSFTSHLYTALKNAGIKVFMDNE-LQRGEDISSSLLKAIEDSRI 251
Query: 109 SIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAF 164
+I++FS NY S+WCLDE+ I EC Q V PVFY+VDPS +R Q G AF
Sbjct: 252 AIIIFSTNYTGSKWCLDELEKIIECQRTIGQEVMPVFYNVDPSDIRKQRGTVGEAF 419
>TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing protein
TSDC, partial (12%)
Length = 701
Score = 119 bits (299), Expect = 2e-27
Identities = 63/144 (43%), Positives = 87/144 (59%), Gaps = 1/144 (0%)
Frame = +1
Query: 71 LYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAI 130
LY L R+G F DD+ L G I+ LL A SR+SI+VFS+NY S WCLDE+ I
Sbjct: 1 LYHALRREGFKTFMDDEGLVGGNEIAKSLLDASEKSRLSIIVFSENYGYSSWCLDELVKI 180
Query: 131 AECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAG 190
EC E KQ V+P+FY V+ S V +Q YE+A + H + D+D V +W+ A+ ++A
Sbjct: 181 VECMETKKQLVWPIFYKVEKSDVTSQENSYEHAMIGHEKNYGKDSDEVKKWRSALSAMAK 360
Query: 191 SAGWDVR-NKPEFREIENIVEAVI 213
G +V+ NK E+ I+ IVE I
Sbjct: 361 LEGHNVKENKYEYEFIKKIVEKAI 432
>AV419023
Length = 410
Score = 117 bits (293), Expect = 9e-27
Identities = 59/96 (61%), Positives = 71/96 (73%)
Frame = +3
Query: 836 LIFLDLGFCSLSEVPHALGEIECLERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCSKLEF 895
L+ LDL FCS+ EVP+A+G + CLERLNL+GNNFV LPS++ L LAYLNL+HC KL
Sbjct: 3 LVSLDLSFCSILEVPNAIGRLWCLERLNLQGNNFVELPSTISRLRRLAYLNLSHCIKLGD 182
Query: 896 LSELQLCDIASEGGRYFRTLSGSHNHRSGLYIFNCP 931
L L + A GGRY+ T S S +HRSGLYIFNCP
Sbjct: 183 LPLLP-SECAPLGGRYYETTSESRDHRSGLYIFNCP 287
Score = 33.9 bits (76), Expect = 0.12
Identities = 25/66 (37%), Positives = 37/66 (55%)
Frame = +3
Query: 762 LDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHL 821
LD+ C S+ V +IG L LE L+L+ N +P +++ + L L+ C+KL L
Sbjct: 12 LDLSFC-SILEVPNAIGRLWCLERLNLQGN-NFVELPSTISRLRRLAYLNLSHCIKLGDL 185
Query: 822 PLGLPS 827
PL LPS
Sbjct: 186 PL-LPS 200
>TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate
resistance protein KR1, partial (6%)
Length = 586
Score = 108 bits (269), Expect = 5e-24
Identities = 60/138 (43%), Positives = 83/138 (59%)
Frame = +1
Query: 52 YDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSIV 111
YDVFIS SDTR F LY L KGI F DD + + +S +AI +SR++IV
Sbjct: 103 YDVFISLITSDTRFGFSGDLYTALSDKGILTFIDDGLPRPDDIMSTVNSKAIESSRIAIV 282
Query: 112 VFSKNYAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRF 171
V S+NYA S +CLDE+ I + E+ ++PVFY V+PS VR Q G + AF H RF
Sbjct: 283 VISENYASSSFCLDELMYILKRREEKGLLIWPVFYQVEPSDVRYQKGGFGKAFASHEERF 462
Query: 172 KHDADRVDRWKRAMRSLA 189
D+ +V +W+ A++ +A
Sbjct: 463 GSDSAKVRKWRNALKQVA 516
>BP045440
Length = 535
Score = 103 bits (258), Expect = 1e-22
Identities = 59/147 (40%), Positives = 83/147 (56%), Gaps = 2/147 (1%)
Frame = -3
Query: 51 KYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAIRNSRVSI 110
+Y +FISFRG DTR+ F LY L R+G FKDD+ L+ G I +LL I SR +I
Sbjct: 530 RYQIFISFRGMDTRDDFTKPLYQGLCREGFVTFKDDESLEGGSPIE-KLLDDIEESRFAI 354
Query: 111 VVFSKNYAESRWCLDEMAAIAEC--CEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHM 168
++ SKNYAES WCL E++ I EC E Q V P +V P+ VR+ Y +A H
Sbjct: 353 IILSKNYAESEWCLKELSKILECKDQEGKNQLVLPSSTEVTPTEVRHVINRYADAMAKHE 174
Query: 169 LRFKHDADRVDRWKRAMRSLAGSAGWD 195
+ D+ + WK+A+ + G++
Sbjct: 173 -KNGIDSKTIQAWKKALFDVCTLTGFE 96
>BP065014
Length = 498
Score = 87.0 bits (214), Expect = 1e-17
Identities = 43/107 (40%), Positives = 64/107 (59%)
Frame = -2
Query: 83 FKDDKKLQKGESISAQLLQAIRNSRVSIVVFSKNYAESRWCLDEMAAIAECCEDFKQTVF 142
FKDD L+ G I +LL AI SRV+IVV S+N+A+S+WCL E+ I +C + K V
Sbjct: 497 FKDDLSLEDGAPIE-ELLGAIEESRVAIVVLSENFADSKWCLKELVKILDCRKKKKPLVL 321
Query: 143 PVFYDVDPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLA 189
P+FY V+P+ +R+ G Y A H + D+ V W +A+ ++
Sbjct: 320 PIFYKVEPTNIRHLKGRYGEAMAEHEKKLGKDSQTVREWTQALSDIS 180
>BP057104
Length = 496
Score = 81.6 bits (200), Expect = 5e-16
Identities = 51/123 (41%), Positives = 67/123 (54%), Gaps = 2/123 (1%)
Frame = -3
Query: 825 LPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLERLNLEGNNFVSLPSSLRWLSSLAY 884
LP L F+ L LDL FC+L +VP A+G + LE L+L GNNF+SLP S++ LS L
Sbjct: 338 LPDLPHFSC--LQHLDLSFCNLRQVPDAIGWLHSLEDLDLGGNNFISLPPSIKELSKLRV 165
Query: 885 LNLAHCSKLEFLSELQLCDIASEGGRYFRTLSGSHNHRSGLYIFNCPTLAITG--LNLAL 942
LNL HC +L + EL + G R +GLYIFNCP L + N+
Sbjct: 164 LNLEHCKQLRYFPELP----SRTQGWVVR------RPHAGLYIFNCPKLRVIEHCYNVVF 15
Query: 943 LWL 945
W+
Sbjct: 14 SWM 6
Score = 31.6 bits (70), Expect = 0.62
Identities = 27/67 (40%), Positives = 35/67 (51%), Gaps = 4/67 (5%)
Frame = -3
Query: 660 DLPF---LKRMDLSNSKYLTETPNFEG-SRRLERLDLTGCTNLLQVHPSIGLLTKLAFLS 715
DLP L+ +DLS L + P+ G LE LDL G N + + PSI L+KL L+
Sbjct: 332 DLPHFSCLQHLDLSFCN-LRQVPDAIGWLHSLEDLDLGG-NNFISLPPSIKELSKLRVLN 159
Query: 716 FESCSSL 722
E C L
Sbjct: 158 LEHCKQL 138
>TC18755 similar to UP|AAN85378 (AAN85378) Resistance protein (Fragment),
complete
Length = 880
Score = 69.3 bits (168), Expect = 3e-12
Identities = 65/226 (28%), Positives = 102/226 (44%), Gaps = 12/226 (5%)
Frame = +3
Query: 226 DLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDR---ISHLFEARC 282
DL+GI R + L L +VI + GMGG+GKTTL +YD I H F A
Sbjct: 192 DLVGIDRRKKKLMGCLIKPCPVR--KVISVTGMGGMGKTTLVKQVYDDPVVIKH-FRACA 362
Query: 283 FVENVSKVYRDGGVTAVQKQVLRQTVDE------MNLETYSPSEISGIVRDRLRSLKVLV 336
++ V + + + + + RQ E + LE + I++D L+ + LV
Sbjct: 363 WIT----VSQSCEIGELLRDLARQLFSEIRRPVPLGLENMRCDRLKMIIKDLLQRRRYLV 530
Query: 337 VLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHI---VYEVPLMNNNDAR 393
V D+V + + + GSR++ITTR + VY + + ++A
Sbjct: 531 VFDDVWHVREWEAVKYALPDNNCGSRIMITTRRSDLAFTSSTESKGKVYNLQPLKEDEAW 710
Query: 394 ELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFLCTRN 439
ELF RK F D+ S + +L+ +GLPLAI L T++
Sbjct: 711 ELFCRKTFHGDSCPSHLIGICTYILRKCEGLPLAIVAISGVLATKD 848
>TC12829
Length = 448
Score = 67.0 bits (162), Expect(2) = 3e-12
Identities = 33/57 (57%), Positives = 43/57 (74%)
Frame = +3
Query: 209 VEAVIEALGRKFSGFADDLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTT 265
V+A I+ LG KFS F++ L+GIQP +E + NLLKL+ E D +V+GIW GGIGKTT
Sbjct: 54 VQARIKTLGHKFSRFSNGLVGIQPHIEEI*NLLKLS*EDEDFRVLGIWRTGGIGKTT 224
Score = 21.9 bits (45), Expect(2) = 3e-12
Identities = 9/14 (64%), Positives = 10/14 (71%)
Frame = +1
Query: 195 DVRNKPEFREIENI 208
D NKPEF EIE +
Sbjct: 13 DQLNKPEFXEIEKM 54
>BP065724
Length = 495
Score = 68.9 bits (167), Expect = 3e-12
Identities = 28/69 (40%), Positives = 45/69 (64%)
Frame = -1
Query: 117 YAESRWCLDEMAAIAECCEDFKQTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRFKHDAD 176
YA+S WCL+E+ I EC + Q V+P+FY V+PS VR+Q YE A + R+ D++
Sbjct: 495 YADSSWCLEEVVKILECKQKNDQLVWPIFYKVEPSDVRHQRNSYEKAMIKQETRYGKDSE 316
Query: 177 RVDRWKRAM 185
+V +W+ ++
Sbjct: 315 KVKKWRSSL 289
>AV410283
Length = 426
Score = 66.6 bits (161), Expect = 2e-11
Identities = 39/107 (36%), Positives = 59/107 (54%)
Frame = +1
Query: 329 LRSLKVLVVLDNVDQLVQLQEFAVNPGLFQKGSRMIITTRDEHILKVYGAHIVYEVPLMN 388
L+ KVL++LD+VD++ QL N F KGSR++ITTR+ +L + YEV +
Sbjct: 82 LQGNKVLLILDDVDEIQQLDFLMGNREWFHKGSRVVITTRNTQVLPESYVDMFYEVRELE 261
Query: 389 NNDARELFYRKGFKSDNLSSRCAELVPEVLKYAQGLPLAIRVTGSFL 435
+ A LF + + + L +++K GLPLA+ V GSFL
Sbjct: 262 LSAALALFCHHAMRRKKPAEGFSNLSKQIVKKTGGLPLALEVIGSFL 402
>AV778571
Length = 452
Score = 63.5 bits (153), Expect = 1e-10
Identities = 41/123 (33%), Positives = 60/123 (48%), Gaps = 1/123 (0%)
Frame = +3
Query: 670 SNSKYLTETPNFEGSRRLERLDLTGCTNLLQVHPSIGLLTKLAFLSFESCSSLVSLDLGS 729
S + +T+ P+ G LE+L NL+++H S+G L KL L E C + +
Sbjct: 39 SECESITQIPDLSGLPNLEKLSFVYSENLIKIHESVGFLNKLRILDIEGCCKIRTFPPMK 218
Query: 730 LCVLYSLAVLHLSGCTKLESTPNFTG-VENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSL 788
L SL L+LS C+ LES P +EN+ + + + + SIG LTRL L L
Sbjct: 219 LA---SLEELYLSHCSSLESFPEILAKMENITLIGVYH-TPIKELPSSIGNLTRLRRLEL 386
Query: 789 RDC 791
C
Sbjct: 387 HGC 395
Score = 55.8 bits (133), Expect = 3e-08
Identities = 38/145 (26%), Positives = 63/145 (43%)
Frame = +3
Query: 742 SGCTKLESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSV 801
S C + P+ +G+ NLE L +L + +S+G L +L L + C + P
Sbjct: 39 SECESITQIPDLSGLPNLEKLSFVYSENLIKIHESVGFLNKLRILDIEGCCKIRTFP--P 212
Query: 802 NNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLER 861
+ SL L C L+ P L + TL + + + E+P ++G + L R
Sbjct: 213 MKLASLEELYLSHCSSLESFPEILAKMENITL-----IGVYHTPIKELPSSIGNLTRLRR 377
Query: 862 LNLEGNNFVSLPSSLRWLSSLAYLN 886
L L G + LPS + + L L+
Sbjct: 378 LELHGCGMLQLPSGMAM*TELEELS 452
Score = 27.7 bits (60), Expect = 8.9
Identities = 24/87 (27%), Positives = 44/87 (49%), Gaps = 2/87 (2%)
Frame = +3
Query: 815 CLKLKHLP--LGLPSLSPFTLQSLIFLDLGFCSLSEVPHALGEIECLERLNLEGNNFVSL 872
C + +P GLP+L S ++ + +L ++ ++G + L L++EG +
Sbjct: 45 CESITQIPDLSGLPNLEKL---SFVYSE----NLIKIHESVGFLNKLRILDIEGCCKIRT 203
Query: 873 PSSLRWLSSLAYLNLAHCSKLEFLSEL 899
++ L+SL L L+HCS LE E+
Sbjct: 204 FPPMK-LASLEELYLSHCSSLESFPEI 281
>TC12516 similar to UP|Q9M5V7 (Q9M5V7) Disease resistance-like protein
(Fragment), partial (36%)
Length = 859
Score = 60.5 bits (145), Expect = 1e-09
Identities = 35/108 (32%), Positives = 65/108 (59%)
Frame = +3
Query: 226 DLIGIQPRVETLENLLKLNSEYYDCQVIGIWGMGGIGKTTLATVLYDRISHLFEARCFVE 285
+LIG+ + LE+LL S+ D +VIGIWGMGGIGKTT+A +++R +E CF+
Sbjct: 543 ELIGVDKSIAHLESLLCQESK--DVRVIGIWGMGGIGKTTIAEEIFNRRHSQYEGICFLS 716
Query: 286 NVSKVYRDGGVTAVQKQVLRQTVDEMNLETYSPSEISGIVRDRLRSLK 333
V G+ +++++ L T+ +++ +P+ ++ ++ R+ +K
Sbjct: 717 KVKDEL*KHGMQSLREK-LFSTLLAQDVKIDTPNGMADDIKRRIGRMK 857
>TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like, partial
(5%)
Length = 689
Score = 58.9 bits (141), Expect = 4e-09
Identities = 27/60 (45%), Positives = 40/60 (66%)
Frame = -2
Query: 44 SNGARRYKYDVFISFRGSDTRNTFVDHLYAHLVRKGIFVFKDDKKLQKGESISAQLLQAI 103
S+ ++ YDVFI+FRG DTR++FV HLY L G+ VF DD+ L++G + +L + I
Sbjct: 226 SSSNHQWIYDVFINFRGKDTRHSFVSHLYTALSNAGVNVFLDDENLERGLELEPELQKLI 47
>TC10205 weakly similar to UP|Q8RL77 (Q8RL77) HpaG, partial (10%)
Length = 586
Score = 54.3 bits (129), Expect = 9e-08
Identities = 24/61 (39%), Positives = 41/61 (66%)
Frame = +3
Query: 747 LESTPNFTGVENLEYLDIDQCVSLSTVDQSIGVLTRLEFLSLRDCLNLTNIPLSVNNMES 806
LE + T +E +++LDI C+SLS + + +G L +LE L++R C LT +P S+ ++E+
Sbjct: 3 LELPKSITNLEQVQFLDISDCISLSHLPEDMGELKKLEKLNVRGCSRLTELPTSIMDLEN 182
Query: 807 L 807
L
Sbjct: 183 L 185
Score = 39.3 bits (90), Expect = 0.003
Identities = 31/106 (29%), Positives = 56/106 (52%), Gaps = 2/106 (1%)
Frame = +3
Query: 797 IPLSVNNMESLLTLDFCGCLKLKHLPLGLPSLSPFTLQSLIFLDLGFCS-LSEVPHALGE 855
+P S+ N+E + LD C+ L HLP + L+ L L++ CS L+E+P ++ +
Sbjct: 9 LPKSITNLEQVQFLDISDCISLSHLPEDMGE-----LKKLEKLNVRGCSRLTELPTSIMD 173
Query: 856 IECLERLNLEGNNFVSLPSSLRWLSSLAYLNLAHCS-KLEFLSELQ 900
+E LE + + L ++ + S L++A L+FLS+L+
Sbjct: 174 LENLEEVECD-EEIAELWEPVQQILSDLQLHVAPTDFNLDFLSKLR 308
Score = 38.9 bits (89), Expect = 0.004
Identities = 17/54 (31%), Positives = 33/54 (60%)
Frame = +3
Query: 775 QSIGVLTRLEFLSLRDCLNLTNIPLSVNNMESLLTLDFCGCLKLKHLPLGLPSL 828
+SI L +++FL + DC++L+++P + ++ L L+ GC +L LP + L
Sbjct: 15 KSITNLEQVQFLDISDCISLSHLPEDMGELKKLEKLNVRGCSRLTELPTSIMDL 176
>TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (11%)
Length = 494
Score = 52.0 bits (123), Expect = 4e-07
Identities = 26/76 (34%), Positives = 44/76 (57%), Gaps = 1/76 (1%)
Frame = +1
Query: 139 QTVFPVFYDVDPSPVRNQNGVYENAFVFHMLRFKHDADRVDRWKRAMRSLAGSAGWDV-R 197
Q V+P+FY++ S V NQ Y +A + H F D+++V +WK A+ +A G +
Sbjct: 4 QLVWPIFYEISXSDVSNQTKSYGHAMLAHENSFGKDSEKVQKWKSALSEMAFLQGEHITE 183
Query: 198 NKPEFREIENIVEAVI 213
N+ E++ I+ IVE +
Sbjct: 184 NEYEYKLIQKIVEKAL 231
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,141,877
Number of Sequences: 28460
Number of extensions: 267159
Number of successful extensions: 1566
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 1524
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1550
length of query: 949
length of database: 4,897,600
effective HSP length: 99
effective length of query: 850
effective length of database: 2,080,060
effective search space: 1768051000
effective search space used: 1768051000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0388b.1


