

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.4
(1489 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
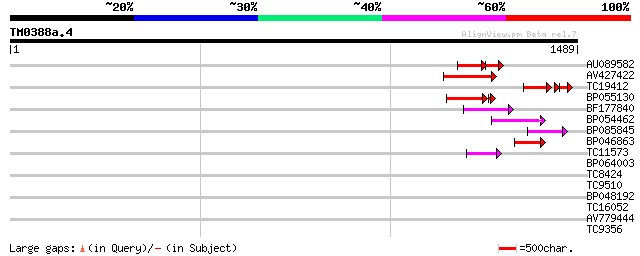
Score E
Sequences producing significant alignments: (bits) Value
AU089582 104 5e-37
AV427422 133 2e-31
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 94 9e-26
BP055130 100 6e-23
BF177840 91 1e-18
BP054462 81 1e-15
BP085845 56 4e-08
BP046863 53 3e-07
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 50 3e-06
BP064003 31 1.3
TC8424 weakly similar to GB|BAC22265.1|24414015|AP003803 arm rep... 30 3.7
TC9510 similar to UP|O22709 (O22709) F8A5.23 protein, partial (49%) 29 4.8
BP048192 29 4.8
TC16052 homologue to UP|Q9LIN9 (Q9LIN9) Similarity to the auxin-... 29 6.2
AV779444 28 8.2
TC9356 weakly similar to UP|AGOL_ARATH (Q9SJK3) Argonaute-like p... 28 8.2
>AU089582
Length = 383
Score = 104 bits (260), Expect(2) = 5e-37
Identities = 48/76 (63%), Positives = 65/76 (85%)
Frame = +2
Query: 1177 AEIHIAKIVRLHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTI 1236
A+I++ +IV LHGVP SI+SDR ++FTS FW + Q ALGT+L++S+A++PQTDGQ+ERTI
Sbjct: 23 AKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSERTI 202
Query: 1237 QSLEDLLRACVLDHKG 1252
Q LED+LRACV D +G
Sbjct: 203 QILEDMLRACVXDLRG 250
Score = 68.6 bits (166), Expect(2) = 5e-37
Identities = 28/46 (60%), Positives = 38/46 (81%)
Frame = +3
Query: 1251 KGSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDG 1296
+GSWD+ L L+EF YNNS+ +SI MAP+EALYGRRC++P+ W + G
Sbjct: 246 EGSWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSPIGWFEVG 383
>AV427422
Length = 417
Score = 133 bits (334), Expect = 2e-31
Identities = 66/139 (47%), Positives = 92/139 (65%)
Frame = +1
Query: 1139 TALPLTRRRFDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDR 1198
T LP ++ ++A+ VVVDRL+K +HFVP+ Y + +A+I + ++VRLHGVP SIVSDR
Sbjct: 4 TGLPKSKG-YEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDR 180
Query: 1199 DSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELL 1258
D F S FW L + GTKL++S+AY+P++DGQTE + LE LR + D SW +
Sbjct: 181 DPLFMSNFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWV 360
Query: 1259 PLIEFTYNNSFHASIGMAP 1277
P E+ YN S+H S G P
Sbjct: 361 PWAEYWYNTSYHVSTGQTP 417
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 94.4 bits (233), Expect(3) = 9e-26
Identities = 44/74 (59%), Positives = 57/74 (76%)
Frame = +3
Query: 1349 RVTPMTGVGRAIKSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVSQLRKYM 1408
RV+PM GV R K KL+P+FIGP+++ ERVG V+YR+ALPP LS +H V HVS LRKY+
Sbjct: 3 RVSPMKGVLRFGKKGKLSPRFIGPFEVLERVGSVSYRLALPPDLSAVHPVFHVSMLRKYL 182
Query: 1409 ADDSHVLEPDDIQL 1422
D SHV+ +D+QL
Sbjct: 183 YDPSHVIRHEDVQL 224
Score = 38.5 bits (88), Expect(3) = 9e-26
Identities = 19/35 (54%), Positives = 24/35 (68%), Gaps = 1/35 (2%)
Frame = +2
Query: 1445 NK*VSLVKVVWNQATGD-ATWELEDKMRESHPDLF 1478
+K V VKV+W +G+ ATWE ED MRE +P LF
Sbjct: 293 SKDVGSVKVLWRGPSGEEATWEAEDIMREKYPHLF 397
Score = 22.7 bits (47), Expect(3) = 9e-26
Identities = 9/13 (69%), Positives = 10/13 (76%)
Frame = +1
Query: 1432 PIKIVDRSTKRLR 1444
P+ IVDR KRLR
Sbjct: 253 PVAIVDRQVKRLR 291
>BP055130
Length = 567
Score = 100 bits (250), Expect(2) = 6e-23
Identities = 53/107 (49%), Positives = 67/107 (62%)
Frame = +2
Query: 1148 FDAIWVVVDRLTKTAHFVPISLNYKVEKLAEIHIAKIVRLHGVPSSIVSDRDSRFTSRFW 1207
F I VVVDRL+K HF P NY ++AE ++ IV+LHG+P +IVSDRD FTS FW
Sbjct: 182 FTVIIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAFW 361
Query: 1208 GALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSW 1254
+ GT L +SS+Y+PQTDGQTE + LE LR V + W
Sbjct: 362 KHFFKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLW 502
Score = 25.0 bits (53), Expect(2) = 6e-23
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = +3
Query: 1258 LPLIEFTYNNSFHASIG 1274
LP E+ YN+SF SIG
Sbjct: 516 LPWAEYCYNSSFQTSIG 566
>BF177840
Length = 410
Score = 90.9 bits (224), Expect = 1e-18
Identities = 51/131 (38%), Positives = 76/131 (57%)
Frame = +2
Query: 1192 SSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHK 1251
+SIVSDRD++F S FW L +GTKL S+ +PQTDGQTE ++L LLR+ + +
Sbjct: 11 TSIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNL 190
Query: 1252 GSWDELLPLIEFTYNNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTE 1311
W+ LP IEF YN H++ +P+E +YG TPL ++ + + E
Sbjct: 191 KMWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDLLPMPNTYLLKHKDGKAKAE 370
Query: 1312 EVKRIQEKMRI 1322
VKR+ EK+++
Sbjct: 371 FVKRLHEKIKL 403
>BP054462
Length = 422
Score = 81.3 bits (199), Expect = 1e-15
Identities = 46/141 (32%), Positives = 80/141 (56%), Gaps = 1/141 (0%)
Frame = +1
Query: 1266 NNSFHASIGMAPYEALYGRRCQTPLCWHQDGEHLVIGPELVQQTTEEVKRIQEKMRIS*S 1325
N++++ S M+P++ALYGR L + +L E + ++ + S
Sbjct: 1 NSNYNRSAKMSPFQALYGREPPVLLQGTTIPSKIAAVNDLQVGRDELLSDLRANLLKSQD 180
Query: 1326 RQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAIK-SKKLTPKFIGPYQITERVGPVAY 1384
++YA+ +R+++++Q GD VFL++ P A K ++KL+P++ GPY I ++G VAY
Sbjct: 181 MMRTYANKKRRDVDYQIGDEVFLKLQPYRRRSLAKKMNEKLSPRYYGPYPIVAKIGAVAY 360
Query: 1385 RIALPPFLSHIHDVLHVSQLR 1405
R+ LP S +H V HVS L+
Sbjct: 361 RLELPAH-SRVHPVFHVSLLK 420
>BP085845
Length = 464
Score = 56.2 bits (134), Expect = 4e-08
Identities = 34/106 (32%), Positives = 57/106 (53%), Gaps = 1/106 (0%)
Frame = -2
Query: 1361 KSKKLTPKFIGPYQITERVGPVAYRIALPPFLSHIHDVLHVSQLRKYMADDSHVLEPDDI 1420
K +K K IGP++I E V +AY + +L H VLHV+ L + + D S + + +
Sbjct: 421 KKRKTESKIIGPFKILEMVCLIAY*LTHSLYLLAAHIVLHVTLLWRNLYDQSQNICHEGV 242
Query: 1421 QLKDDLTVVMPPIKIVDRSTKRLRNK*VSLVKVVWNQATG-DATWE 1465
L + + + PI +VD + +R K + VKV+W +G + +WE
Sbjct: 241 XLGEHWSHMEHPIVMVDMKVRCMRPKNIDDVKVIWRGLSG*EKSWE 104
>BP046863
Length = 580
Score = 53.1 bits (126), Expect = 3e-07
Identities = 30/82 (36%), Positives = 51/82 (61%), Gaps = 1/82 (1%)
Frame = +1
Query: 1326 RQKSYADNRRKELEFQAGDHVFLRVTPMTGVGRAI-KSKKLTPKFIGPYQITERVGPVAY 1384
+ ++ A+ R+ +++Q G+ VFL++ P A K++KL+P+F GP+++ ERV VAY
Sbjct: 337 QMRAQANKHRRYVDYQVGNWVFLKLQPYKLQNLAQRKNQKLSPRFYGPFKVLERVVQVAY 516
Query: 1385 RIALPPFLSHIHDVLHVSQLRK 1406
+ L S +H V H+S L K
Sbjct: 517 *LDLXS-ESRVHPVFHLSLLEK 579
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 49.7 bits (117), Expect = 3e-06
Identities = 26/90 (28%), Positives = 48/90 (52%)
Frame = +2
Query: 1201 RFTSRFWGALQQALGTKLRLSSAYNPQTDGQTERTIQSLEDLLRACVLDHKGSWDELLPL 1260
+FTS+ +G ++R SS +PQT+GQTE + + ++ + + +G W + LP+
Sbjct: 20 QFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWIDELPI 199
Query: 1261 IEFTYNNSFHASIGMAPYEALYGRRCQTPL 1290
+ ++YN +SI P+ YG P+
Sbjct: 200 VLWSYNTMPQSSIKETPF*LTYGADTMLPV 289
>BP064003
Length = 509
Score = 31.2 bits (69), Expect = 1.3
Identities = 23/61 (37%), Positives = 31/61 (50%), Gaps = 3/61 (4%)
Frame = +1
Query: 907 GSMFTIYSDHKSLKYLFDQKDLNMRQRRWMEFLQD---YEFDLQ*HPGKTNVVADALSRK 963
GS F I +D+ + QK R R W +FL + +Q ++N VADALSRK
Sbjct: 115 GSRFVIKTDNVATSSFLTQKRAPTRAR-WQDFLSGGVRHGPRVQAREDQSNKVADALSRK 291
Query: 964 A 964
A
Sbjct: 292 A 294
>TC8424 weakly similar to GB|BAC22265.1|24414015|AP003803 arm repeat
containing protein homolog-like {Oryza sativa (japonica
cultivar-group);} , partial (49%)
Length = 1586
Score = 29.6 bits (65), Expect = 3.7
Identities = 12/27 (44%), Positives = 14/27 (51%)
Frame = +1
Query: 503 LSWLRRRTTPTYTTFWWCKTLWMYFQM 529
L W RRR WWC T W+ F+M
Sbjct: 769 LVWRRRR--------WWCSTAWLRFRM 825
>TC9510 similar to UP|O22709 (O22709) F8A5.23 protein, partial (49%)
Length = 694
Score = 29.3 bits (64), Expect = 4.8
Identities = 15/33 (45%), Positives = 17/33 (51%), Gaps = 2/33 (6%)
Frame = -2
Query: 1373 YQITERVGPVAYRIALPPFLS--HIHDVLHVSQ 1403
YQI GP Y +AL PF H+H LH Q
Sbjct: 303 YQIWSLTGPFLYVVALKPFQPKVHLHQQLHKHQ 205
>BP048192
Length = 551
Score = 29.3 bits (64), Expect = 4.8
Identities = 10/37 (27%), Positives = 23/37 (62%), Gaps = 3/37 (8%)
Frame = +3
Query: 910 FTIYSDHKSLKYLFDQ---KDLNMRQRRWMEFLQDYE 943
F Y+D+ ++ Y Q K++++ + RW +++DY+
Sbjct: 198 FLTYADNNAVGYFIKQGFTKEIHLEKERWQGYIKDYD 308
>TC16052 homologue to UP|Q9LIN9 (Q9LIN9) Similarity to the auxin-independent
growth promoter, partial (24%)
Length = 809
Score = 28.9 bits (63), Expect = 6.2
Identities = 17/44 (38%), Positives = 21/44 (47%)
Frame = +2
Query: 1187 LHGVPSSIVSDRDSRFTSRFWGALQQALGTKLRLSSAYNPQTDG 1230
+HG PS I FWG++ LG R S Y+P TDG
Sbjct: 197 IHGSPSKIN*T*PRTNVQVFWGSIH-GLGYFCRRCSRYSPDTDG 325
>AV779444
Length = 539
Score = 28.5 bits (62), Expect = 8.2
Identities = 14/45 (31%), Positives = 19/45 (42%)
Frame = +1
Query: 1431 PPIKIVDRSTKRLRNK*VSLVKVVWNQATGDATWELEDKMRESHP 1475
P + V S +R K S++ +VW T TW L HP
Sbjct: 343 PSLCTVAPSRWAVR*KHYSILSIVWRDGTASCTWRLTSST*PPHP 477
>TC9356 weakly similar to UP|AGOL_ARATH (Q9SJK3) Argonaute-like protein
At2g27880, partial (10%)
Length = 697
Score = 28.5 bits (62), Expect = 8.2
Identities = 21/53 (39%), Positives = 26/53 (48%), Gaps = 3/53 (5%)
Frame = -3
Query: 143 WRSTSRRLPGKTWRINSSA*DRG**P*GN---MLQDWRPFPNTFAFSKCKWMK 192
WR +RR + WR N SA DR G +L+ WRP T A C WM+
Sbjct: 245 WRG-NRRGNNRRWR-NRSADDRSGRMRGR*WRLLRRWRPCTATAAACACIWMR 93
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.342 0.148 0.511
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,036,910
Number of Sequences: 28460
Number of extensions: 422407
Number of successful extensions: 2922
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2847
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2920
length of query: 1489
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1387
effective length of database: 1,994,680
effective search space: 2766621160
effective search space used: 2766621160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0388a.4


