

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.2
(422 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
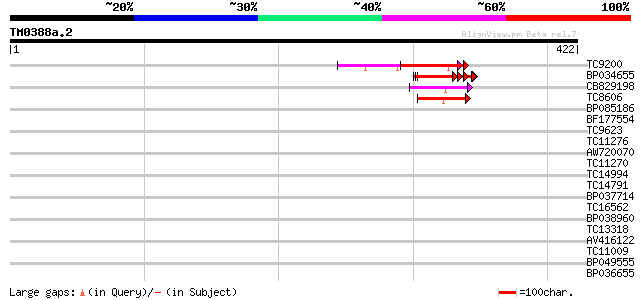
Score E
Sequences producing significant alignments: (bits) Value
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 45 2e-05
BP034655 44 4e-05
CB829198 42 1e-04
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 42 2e-04
BP085186 39 0.001
BF177554 39 0.002
TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxid... 38 0.004
TC11276 weakly similar to UP|HDA1_CHICK (P56517) Histone deacety... 37 0.005
AW720070 37 0.006
TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%) 37 0.006
TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , p... 37 0.006
TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), par... 37 0.006
BP037714 36 0.010
TC16562 similar to GB|AAM78073.1|21928089|AY125563 AT4g24660/F22... 36 0.014
BP038960 36 0.014
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 36 0.014
AV416122 35 0.023
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 35 0.023
BP049555 35 0.030
BP036655 34 0.040
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 45.1 bits (105), Expect = 2e-05
Identities = 37/103 (35%), Positives = 50/103 (47%), Gaps = 10/103 (9%)
Frame = +1
Query: 245 SKDKHRPIFPHLICSLIHA-----VRDRARRPVHVIEKRLPVAPLLSK-----ERIAQLY 294
SK ++ PH + SL + V + + ++ K A L K E Q Y
Sbjct: 19 SKSENPSFNPHSLASLQESPNEAMVNESITKKRKLVPKSSQSAELPFKLQNKLEAEVQEY 198
Query: 295 KEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E + +E EEEE+ EE E+ EEEEEEEEEEE VE E
Sbjct: 199 EEVEEEVEEEVEEEEEEEEEE-EEGEEEEEEEEEEEEVEAEAE 324
Score = 40.0 bits (92), Expect = 7e-04
Identities = 21/54 (38%), Positives = 35/54 (63%), Gaps = 4/54 (7%)
Frame = +1
Query: 292 QLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEE----EEEEENVEVVNEVVTP 341
++ +E + +E+EEEE+ EE E++EEEEEE E EEE+ E + +++ P
Sbjct: 202 EVEEEVEEEVEEEEEEEEEEEEGEEEEEEEEEEEEVEAEAEEEDDEPIQKLLEP 363
Score = 32.0 bits (71), Expect = 0.20
Identities = 26/95 (27%), Positives = 46/95 (48%), Gaps = 2/95 (2%)
Frame = +1
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV-NEVVTPPRAD 345
+E + + +E E+EEEE+ EE E + EEE++E ++ +E + E + AD
Sbjct: 220 EEEVEEEEEEEEEEEEGEEEEEEEEEEEEVEAEAEEEDDEPIQKLLEPMGKEQIISLLAD 399
Query: 346 PSPAWMEAAFG-RMFLRQDRLHRDLDLHWRGGSTS 379
+ + A R +D HR + +H G T+
Sbjct: 400 AASNHRDVADRIRKVADEDVSHRKIFVHGLGWDTT 504
>BP034655
Length = 517
Score = 44.3 bits (103), Expect = 4e-05
Identities = 20/33 (60%), Positives = 26/33 (78%)
Frame = +1
Query: 301 MPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ +DEEEE+ EE E++EEEEEEEEEEE+ E
Sbjct: 130 LSDDDEEEEEEEEEEEEEEEEEEEEEEEEEDEE 228
Score = 43.1 bits (100), Expect = 9e-05
Identities = 20/39 (51%), Positives = 29/39 (74%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTP 341
+E+EEEE+ EE E++EEEEEEE+EE++ E +E P
Sbjct: 148 EEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYIP 264
Score = 42.7 bits (99), Expect = 1e-04
Identities = 19/31 (61%), Positives = 26/31 (83%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E+EEEE+ EE E++EEEEEEEE+EE+ E
Sbjct: 145 EEEEEEEEEEEEEEEEEEEEEEEEEDEEDDE 237
Score = 42.4 bits (98), Expect = 1e-04
Identities = 20/34 (58%), Positives = 26/34 (75%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+D+EEE+ EE E++EEEEEEEEEEE E +E
Sbjct: 136 DDDEEEEEEEEEEEEEEEEEEEEEEEEEDEEDDE 237
Score = 42.0 bits (97), Expect = 2e-04
Identities = 20/46 (43%), Positives = 30/46 (64%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSP 348
+E+EEEE+ EE E++EEEEE+EE++E E + + P SP
Sbjct: 154 EEEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYIPVAPSHRVSP 291
Score = 41.2 bits (95), Expect = 3e-04
Identities = 19/34 (55%), Positives = 26/34 (75%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
E+EEEE+ EE E++EEEEEEEEE+E + +E
Sbjct: 145 EEEEEEEEEEEEEEEEEEEEEEEEEDEEDDEEDE 246
Score = 40.8 bits (94), Expect = 4e-04
Identities = 19/45 (42%), Positives = 29/45 (64%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPS 347
+E+EEEE+ EE E++EEE+EE++EE+ E V R P+
Sbjct: 160 EEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYIPVAPSHRVSPN 294
>CB829198
Length = 562
Score = 42.4 bits (98), Expect = 1e-04
Identities = 23/52 (44%), Positives = 30/52 (57%), Gaps = 5/52 (9%)
Frame = +3
Query: 298 TARMPQEDEEEEDPEEPAGEDKEEEE-----EEEEEEENVEVVNEVVTPPRA 344
T PQE+E+ +D EEP +D EE + EE EEEE+ + NE P RA
Sbjct: 213 TEHQPQEEEQHDDQEEPQNDDVEESQHSDSKEETEEEEDEDGDNENNKPKRA 368
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 41.6 bits (96), Expect = 2e-04
Identities = 19/43 (44%), Positives = 29/43 (67%), Gaps = 3/43 (6%)
Frame = -2
Query: 304 EDEEEEDPEEPAGEDKEE---EEEEEEEEENVEVVNEVVTPPR 343
+D+E++D E+ GED+EE EEE+ E+EE E E + PP+
Sbjct: 386 DDDEDDDEEDDGGEDEEEEGVEEEDNEDEEEDEEDEEALQPPK 258
Score = 29.3 bits (64), Expect = 1.3
Identities = 13/32 (40%), Positives = 23/32 (71%), Gaps = 1/32 (3%)
Frame = -2
Query: 303 QEDEEEEDPEEPAGEDKEEEE-EEEEEEENVE 333
++ E++ED ++ +D EE++ E+EEEE VE
Sbjct: 416 EDGEDQEDDDDDDEDDDEEDDGGEDEEEEGVE 321
>BP085186
Length = 388
Score = 39.3 bits (90), Expect = 0.001
Identities = 18/30 (60%), Positives = 23/30 (76%)
Frame = -1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
E+EEEE+ EE E++EEEE E EEE+N E
Sbjct: 259 EEEEEEEEEEEEEEEEEEEEAEPEEEKNEE 170
Score = 38.5 bits (88), Expect = 0.002
Identities = 17/30 (56%), Positives = 23/30 (76%)
Frame = -1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENV 332
+E+EEEE+ EE E++ E EEE+ EEENV
Sbjct: 250 EEEEEEEEEEEEEEEEEAEPEEEKNEEENV 161
Score = 38.5 bits (88), Expect = 0.002
Identities = 18/34 (52%), Positives = 24/34 (69%)
Frame = -1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVN 336
+E+EEEE+ EE E++EEE E EEE+ E VN
Sbjct: 259 EEEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVN 158
Score = 36.6 bits (83), Expect = 0.008
Identities = 20/33 (60%), Positives = 24/33 (72%)
Frame = -1
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+EEEE+ EE E++EEEEEEEEE E E NE
Sbjct: 262 NEEEEEEEE---EEEEEEEEEEEEAEPEEEKNE 173
Score = 35.0 bits (79), Expect = 0.023
Identities = 17/34 (50%), Positives = 22/34 (64%)
Frame = -1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
++ + E EE E++EEEEEEEEEEE E E
Sbjct: 283 DNADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEE 182
Score = 34.7 bits (78), Expect = 0.030
Identities = 17/35 (48%), Positives = 19/35 (53%)
Frame = -1
Query: 311 PEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRAD 345
P+ GE EEEEEEEEEEE E E P +
Sbjct: 286 PDNADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEE 182
Score = 29.3 bits (64), Expect = 1.3
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = -1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+E+EEEE+ EP E EEE +E E
Sbjct: 226 EEEEEEEEEAEPEEEKNEEENVNTQEGE 143
Score = 26.9 bits (58), Expect = 6.3
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = -1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEE 329
+E+EEE +PEE E++ +E E E
Sbjct: 217 EEEEEEAEPEEEKNEEENVNTQEGEGE 137
>BF177554
Length = 246
Score = 38.5 bits (88), Expect = 0.002
Identities = 18/30 (60%), Positives = 21/30 (70%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
E ++E+ EE AGE EEEEEEEEE E E
Sbjct: 16 EQQDEDSCEEEAGESSEEEEEEEEEGEEEE 105
Score = 35.4 bits (80), Expect = 0.018
Identities = 17/33 (51%), Positives = 22/33 (66%)
Frame = +1
Query: 298 TARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
T+ ED EE+ E + E++EEEEE EEEEE
Sbjct: 10 TSEQQDEDSCEEEAGESSEEEEEEEEEGEEEEE 108
Score = 30.8 bits (68), Expect = 0.44
Identities = 14/42 (33%), Positives = 23/42 (54%)
Frame = +1
Query: 287 KERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEE 328
+E + +E + +EEEE+ + P D + EEEEEE+
Sbjct: 40 EEEAGESSEEEEEEEEEGEEEEEEDDGPGDRDAQPEEEEEED 165
Score = 30.4 bits (67), Expect = 0.57
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +1
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
+E+E EE+ EE G + + EEEEEE+ ++
Sbjct: 79 EEEEGEEEEEEDDGPGDRDAQPEEEEEEDFSLL 177
Score = 29.3 bits (64), Expect = 1.3
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEE 329
E+E E EE E++E EEEEEE++
Sbjct: 40 EEEAGESSEEEEEEEEEGEEEEEEDD 117
>TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxidase,
partial (42%)
Length = 640
Score = 37.7 bits (86), Expect = 0.004
Identities = 26/72 (36%), Positives = 34/72 (47%), Gaps = 2/72 (2%)
Frame = -1
Query: 275 IEKRLPVAPLLSK--ERIAQLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENV 332
+E R P L K E + K W +E+EEEE+ E+ E+ EEEE E V
Sbjct: 271 LETRTSALP*LRKR*EMRGEERKHWKEER-EEEEEEEENEKKEREEYEEEESERVALVAV 95
Query: 333 EVVNEVVTPPRA 344
E + PPRA
Sbjct: 94 EGIMRAAAPPRA 59
>TC11276 weakly similar to UP|HDA1_CHICK (P56517) Histone deacetylase 1
(HD1), partial (5%)
Length = 636
Score = 37.4 bits (85), Expect = 0.005
Identities = 21/58 (36%), Positives = 35/58 (60%), Gaps = 6/58 (10%)
Frame = +3
Query: 302 PQEDEEEEDP---EEPAGEDKEEEE---EEEEEEENVEVVNEVVTPPRADPSPAWMEA 353
PQ++EE++D EEP E+ ++EE EEE+++E E + P + DP W++A
Sbjct: 60 PQKEEEKKDEPKSEEPQKEEAKKEEPKKEEEKKDEKKEEEKKKEQPAQIDPVLEWVKA 233
Score = 27.7 bits (60), Expect = 3.7
Identities = 12/32 (37%), Positives = 21/32 (65%)
Frame = +3
Query: 306 EEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
+E E EEP E+ ++EEE+++E ++ E E
Sbjct: 21 KEPEKKEEPKKEEPQKEEEKKDEPKSEEPQKE 116
>AW720070
Length = 484
Score = 37.0 bits (84), Expect = 0.006
Identities = 19/35 (54%), Positives = 25/35 (71%), Gaps = 6/35 (17%)
Frame = +1
Query: 299 ARMPQEDEEEEDPEEPAGED------KEEEEEEEE 327
+R +E+EEEE+ EE AGE+ +EEEEEEEE
Sbjct: 103 SREEEEEEEEEEEEEEAGEEGIITTTREEEEEEEE 207
Score = 31.6 bits (70), Expect = 0.26
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 1/39 (2%)
Frame = +1
Query: 308 EEDPEEPAGEDKEEEEEEEEEEENVEVVNE-VVTPPRAD 345
+E+ + + +EEEEEEEEEEE E E ++T R +
Sbjct: 73 QEEAVKGYSDSREEEEEEEEEEEEEEAGEEGIITTTREE 189
Score = 28.1 bits (61), Expect = 2.8
Identities = 13/30 (43%), Positives = 19/30 (63%)
Frame = +1
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
++E + + E++EEEEEEEEEE E
Sbjct: 73 QEEAVKGYSDSREEEEEEEEEEEEEEAGEE 162
>TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%)
Length = 735
Score = 37.0 bits (84), Expect = 0.006
Identities = 22/43 (51%), Positives = 25/43 (57%), Gaps = 8/43 (18%)
Frame = +3
Query: 303 QEDEEEEDPE--------EPAGEDKEEEEEEEEEEENVEVVNE 337
+EDEEE+ P E A E+ EEEE EEEEE EVV E
Sbjct: 357 EEDEEEQTPHPTRGRSRVEHAQEEDEEEENGEEEEEEEEVVVE 485
Score = 35.0 bits (79), Expect = 0.023
Identities = 21/45 (46%), Positives = 25/45 (54%), Gaps = 13/45 (28%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEE-------------EEEEEENVEVV 335
E +EED EE GE++EEEEE E+EEEE V VV
Sbjct: 411 EHAQEEDEEEENGEEEEEEEEVVVEDEQGNEDEDEDEEEEGVVVV 545
Score = 31.2 bits (69), Expect = 0.34
Identities = 19/55 (34%), Positives = 28/55 (50%), Gaps = 3/55 (5%)
Frame = +3
Query: 283 PLLSKERIAQLYKEWTARMPQEDEEEEDP---EEPAGEDKEEEEEEEEEEENVEV 334
P + R+ +E E+EEEE+ E+ G + E+E+EEEE VEV
Sbjct: 387 PTRGRSRVEHAQEEDEEEENGEEEEEEEEVVVEDEQGNEDEDEDEEEEGVVVVEV 551
>TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial
(89%)
Length = 1672
Score = 37.0 bits (84), Expect = 0.006
Identities = 19/28 (67%), Positives = 22/28 (77%)
Frame = +2
Query: 306 EEEEDPEEPAGEDKEEEEEEEEEEENVE 333
EEEE+ EE E++EEEEEEEEEEE E
Sbjct: 161 EEEEEVEE---EEEEEEEEEEEEEEEAE 235
Score = 36.6 bits (83), Expect = 0.008
Identities = 17/27 (62%), Positives = 21/27 (76%)
Frame = +2
Query: 305 DEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+EEEE EE E++EEEEEEEE E+N
Sbjct: 161 EEEEEVEEEEEEEEEEEEEEEEEAEKN 241
Score = 34.3 bits (77), Expect = 0.040
Identities = 18/38 (47%), Positives = 24/38 (62%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVT 340
+E+EEE + EE E++EEEEEEE E+ N V T
Sbjct: 158 REEEEEVEEEEEEEEEEEEEEEEEAEKNGG*SRNRVST 271
Score = 28.5 bits (62), Expect = 2.2
Identities = 14/23 (60%), Positives = 16/23 (68%)
Frame = +2
Query: 315 AGEDKEEEEEEEEEEENVEVVNE 337
A E++EE EEEEEEEE E E
Sbjct: 155 AREEEEEVEEEEEEEEEEEEEEE 223
>TC14791 similar to UP|Q9LJ04 (Q9LJ04) ESTs AU082210(C53655), partial (37%)
Length = 1779
Score = 37.0 bits (84), Expect = 0.006
Identities = 18/43 (41%), Positives = 26/43 (59%), Gaps = 3/43 (6%)
Frame = +2
Query: 298 TARMPQED---EEEEDPEEPAGEDKEEEEEEEEEEENVEVVNE 337
T PQ D EEE+DPE PA + E ++++E E+N + NE
Sbjct: 251 TEDQPQIDDAVEEEQDPEAPATTESEPQQQQETLEDNADAPNE 379
Score = 28.1 bits (61), Expect = 2.8
Identities = 23/71 (32%), Positives = 34/71 (47%), Gaps = 8/71 (11%)
Frame = +2
Query: 317 EDKEEEEEEEEEEENVEVVNEVVTPPR--------ADPSPAWMEAAFGRMFLRQDRLHRD 368
E++EEEE+ E EEE VE + E T + A+ P ++E R D HR
Sbjct: 440 EEEEEEEDLELEEEPVERLLEPFTREQLHSLVKQAAEKYPDFVENV--RQLADVDPAHRK 613
Query: 369 LDLHWRGGSTS 379
+ +H G T+
Sbjct: 614 IFVHGLGWDTT 646
>BP037714
Length = 463
Score = 36.2 bits (82), Expect = 0.010
Identities = 19/50 (38%), Positives = 29/50 (58%), Gaps = 6/50 (12%)
Frame = +1
Query: 300 RMPQEDEEEEDPEEPAGEDKEEEEEEEEEEEN------VEVVNEVVTPPR 343
R + + EEEDPE+ ED+ EEE EEE ++N +E+ N + P+
Sbjct: 202 RQKEAEHEEEDPEQ-VSEDESEEESEEETKKNKGTQGIIEIENPNLVKPK 348
>TC16562 similar to GB|AAM78073.1|21928089|AY125563 AT4g24660/F22K18_140
{Arabidopsis thaliana;}, partial (29%)
Length = 621
Score = 35.8 bits (81), Expect = 0.014
Identities = 20/42 (47%), Positives = 25/42 (58%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRAD 345
+ + E D E E++EEEEEEEEEE VE + V PP D
Sbjct: 114 DSDMEFDDHEDLEEEEEEEEEEEEEEGEVE-MGFTVAPPGFD 236
>BP038960
Length = 524
Score = 35.8 bits (81), Expect = 0.014
Identities = 18/44 (40%), Positives = 24/44 (53%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADP 346
++ E+ EE A D +EEEEEEEEEE+ V+ P P
Sbjct: 288 EQPRVEDSEEEAADYDDDEEEEEEEEEEDEHADRVAVSGPGKGP 419
Score = 32.3 bits (72), Expect = 0.15
Identities = 13/31 (41%), Positives = 22/31 (70%)
Frame = +3
Query: 299 ARMPQEDEEEEDPEEPAGEDKEEEEEEEEEE 329
A P+ ++ EE+ + +++EEEEEEEE+E
Sbjct: 285 AEQPRVEDSEEEAADYDDDEEEEEEEEEEDE 377
Score = 32.0 bits (71), Expect = 0.20
Identities = 17/37 (45%), Positives = 22/37 (58%)
Frame = +3
Query: 292 QLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEE 328
QL K + ED EEE + E++EEEEEEE+E
Sbjct: 267 QLQKGEAEQPRVEDSEEEAADYDDDEEEEEEEEEEDE 377
Score = 30.4 bits (67), Expect = 0.57
Identities = 20/64 (31%), Positives = 30/64 (46%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPPRADPSPAWMEAAFGRMFLRQD 363
E ED EE A D +++EEEEEEEE + + V P E + +R+
Sbjct: 288 EQPRVEDSEEEAA-DYDDDEEEEEEEEEEDEHADRVAVSGPGKGPILREGFYEVEAIRRK 464
Query: 364 RLHR 367
R+ +
Sbjct: 465 RIRK 476
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 35.8 bits (81), Expect = 0.014
Identities = 12/32 (37%), Positives = 26/32 (80%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVV 335
ED++++D ++ GE++E++++++EEEE E V
Sbjct: 294 EDDDDDDDDDDDGEEEEDDDDDDEEEEGSEEV 389
Score = 32.7 bits (73), Expect = 0.12
Identities = 11/31 (35%), Positives = 24/31 (76%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ED+E+ED ++ +D + EEEE++++++ E
Sbjct: 276 EEDDEDEDDDDDDDDDDDGEEEEDDDDDDEE 368
Score = 32.7 bits (73), Expect = 0.12
Identities = 11/29 (37%), Positives = 23/29 (78%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
++D+EEED E+ +D ++++++ EEEE+
Sbjct: 261 EDDDEEEDDEDEDDDDDDDDDDDGEEEED 347
Score = 32.3 bits (72), Expect = 0.15
Identities = 12/37 (32%), Positives = 26/37 (69%)
Frame = +3
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVV 339
++DE+E+D ++ +D EEEE++++++ E +E V
Sbjct: 279 EDDEDEDDDDDDDDDDDGEEEEDDDDDDEEEEGSEEV 389
Score = 32.0 bits (71), Expect = 0.20
Identities = 11/28 (39%), Positives = 22/28 (78%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
+DEEE+D +E +D +++++ EEEE++
Sbjct: 267 DDEEEDDEDEDDDDDDDDDDDGEEEEDD 350
Score = 29.6 bits (65), Expect = 0.98
Identities = 15/44 (34%), Positives = 26/44 (59%), Gaps = 1/44 (2%)
Frame = +3
Query: 289 RIAQLYKEWTARMPQEDEEEED-PEEPAGEDKEEEEEEEEEEEN 331
R+ +L+ A + + E D P +P D EEEE+++EEE++
Sbjct: 156 RLIELHNGLQAWLVHDPEIYPDGPPKPVQTDNEEEEDDDEEEDD 287
>AV416122
Length = 414
Score = 35.0 bits (79), Expect = 0.023
Identities = 17/28 (60%), Positives = 22/28 (77%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
E EEEED E EDKEEE+EEE++E++
Sbjct: 297 EGEEEEDDE----EDKEEEDEEEDDEDD 368
Score = 30.8 bits (68), Expect = 0.44
Identities = 13/30 (43%), Positives = 20/30 (66%)
Frame = +3
Query: 302 PQEDEEEEDPEEPAGEDKEEEEEEEEEEEN 331
P D + E EE E+ +EEE+EEE++E+
Sbjct: 276 PDNDVQFEGEEEEDDEEDKEEEDEEEDDED 365
Score = 29.6 bits (65), Expect = 0.98
Identities = 11/30 (36%), Positives = 22/30 (72%)
Frame = +3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+D E ++ + GE++E++EE++EEE+ E
Sbjct: 264 DDNEPDNDVQFEGEEEEDDEEDKEEEDEEE 353
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 35.0 bits (79), Expect = 0.023
Identities = 13/30 (43%), Positives = 22/30 (73%)
Frame = +2
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+ +E++D + P G D +++EE+EEEE VE
Sbjct: 59 DGDEDDDDDAPGGGDDDDDEEDEEEEGGVE 148
Score = 33.1 bits (74), Expect = 0.089
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 11/39 (28%)
Frame = +2
Query: 303 QEDEEEE-----------DPEEPAGEDKEEEEEEEEEEE 330
+EDEEEE DP++ +D +++EEEEEEE+
Sbjct: 116 EEDEEEEGGVEGGRGGGGDPDDDDDDDDDDDEEEEEEED 232
Score = 30.8 bits (68), Expect = 0.44
Identities = 15/44 (34%), Positives = 25/44 (56%), Gaps = 15/44 (34%)
Frame = +2
Query: 304 EDEEEEDPEEPAG---------------EDKEEEEEEEEEEENV 332
+D++EED EE G +D ++++EEEEEEE++
Sbjct: 104 DDDDEEDEEEEGGVEGGRGGGGDPDDDDDDDDDDDEEEEEEEDL 235
Score = 29.3 bits (64), Expect = 1.3
Identities = 10/27 (37%), Positives = 20/27 (74%)
Frame = +2
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEE 330
+D +E+D ++ G ++++EE+EEEE
Sbjct: 56 DDGDEDDDDDAPGGGDDDDDEEDEEEE 136
Score = 27.3 bits (59), Expect = 4.9
Identities = 9/28 (32%), Positives = 21/28 (74%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEE 330
++D+ +ED ++ A +++++EE+EEE
Sbjct: 50 EDDDGDEDDDDDAPGGGDDDDDEEDEEE 133
>BP049555
Length = 555
Score = 34.7 bits (78), Expect = 0.030
Identities = 15/31 (48%), Positives = 23/31 (73%)
Frame = +2
Query: 303 QEDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+E EE+ E+PA E+K+EEE+ EEE++ E
Sbjct: 284 EEKAEEKAEEKPAAEEKKEEEKPEEEKKEEE 376
Score = 31.2 bits (69), Expect = 0.34
Identities = 20/58 (34%), Positives = 28/58 (47%), Gaps = 5/58 (8%)
Frame = +2
Query: 290 IAQLYKEWTARMPQ-----EDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVNEVVTPP 342
+ Q+ +E PQ E++ EE EE A E EE++EEE+ E E PP
Sbjct: 215 LEQMGEEAKQEQPQAEAKPEEKTEEKAEEKAEEKPAAEEKKEEEKPEEEKKEEEPKPP 388
>BP036655
Length = 556
Score = 34.3 bits (77), Expect = 0.040
Identities = 17/45 (37%), Positives = 25/45 (54%)
Frame = -3
Query: 292 QLYKEWTARMPQEDEEEEDPEEPAGEDKEEEEEEEEEEENVEVVN 336
++ +EW+ D+E +D E E + E E+EEEEEE V N
Sbjct: 353 EVVEEWSDDDDINDDESDDEVEEESETEIETEKEEEEEEECGVEN 219
Score = 33.5 bits (75), Expect = 0.068
Identities = 17/30 (56%), Positives = 20/30 (66%)
Frame = -3
Query: 304 EDEEEEDPEEPAGEDKEEEEEEEEEEENVE 333
+DE EE+ E +KEEEEEEE ENVE
Sbjct: 302 DDEVEEESETEIETEKEEEEEEECGVENVE 213
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,422,985
Number of Sequences: 28460
Number of extensions: 110011
Number of successful extensions: 2428
Number of sequences better than 10.0: 162
Number of HSP's better than 10.0 without gapping: 989
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1643
length of query: 422
length of database: 4,897,600
effective HSP length: 93
effective length of query: 329
effective length of database: 2,250,820
effective search space: 740519780
effective search space used: 740519780
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0388a.2


