

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
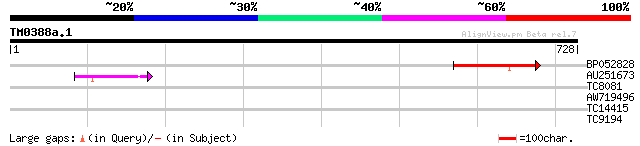
Query= TM0388a.1
(728 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP052828 98 4e-21
AU251673 42 3e-04
TC8081 similar to UP|AAQ84334 (AAQ84334) Zinc-finger protein, pa... 30 1.0
AW719496 30 1.4
TC14415 similar to UP|O04237 (O04237) Transcription factor, part... 28 4.0
TC9194 similar to UP|Q9LYP1 (Q9LYP1) Membrane protein (AT5g07250... 27 8.8
>BP052828
Length = 523
Score = 98.2 bits (243), Expect = 4e-21
Identities = 50/115 (43%), Positives = 76/115 (65%), Gaps = 3/115 (2%)
Frame = +2
Query: 570 LCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVD 629
+ DLGA IN+MP +++ L+ G + T L +Q+A+RS P G+VEDV+V V++ +FP D
Sbjct: 107 MLDLGAGINVMPTSIYNILHPGPLQHTGLIVQLANRSNARPKGVVEDVLVQVNELIFPAD 286
Query: 630 FVVLDMEEDEK---IPLILGRPFLATGRAKIDVDKGHLILPVGKEKVRFSVFNPM 681
F +LDME + K P+ILGRPF+ T + KIDVD G + + G +F++ + M
Sbjct: 287 FYILDMEGETKSSRAPIILGRPFMKTAKTKIDVDNGTMSMEFGDIIAKFNIDDAM 451
>AU251673
Length = 413
Score = 42.4 bits (98), Expect = 3e-04
Identities = 30/105 (28%), Positives = 49/105 (46%), Gaps = 5/105 (4%)
Frame = +2
Query: 84 TDTIRLSLFPFSLRDTAEEWLN----SQPQGSIT-SWEDLAEKFTTRFVPRALLRKLKND 138
+DT + L F L A +W N ++P GS +W D + +F RF+P+++ D
Sbjct: 8 SDTRAVELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVRD 187
Query: 139 IMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQ 183
Q+ + E F L R P + L + E+V +F GL+
Sbjct: 188 FERLEQAEGMTVSEYSAHFTHLSRYVP-YPLLEEERVKRFVRGLK 319
>TC8081 similar to UP|AAQ84334 (AAQ84334) Zinc-finger protein, partial
(65%)
Length = 1222
Score = 30.4 bits (67), Expect = 1.0
Identities = 14/40 (35%), Positives = 20/40 (50%)
Frame = +1
Query: 415 KQENASVVATRSGRVMSELKKKTEGEKREEIVGGEVVVPI 454
KQE A + AT G +M+ E E E +V V +P+
Sbjct: 319 KQEQAKLAATSIGNIMNGSSSSIETEPTEPVVAASVDIPV 438
>AW719496
Length = 492
Score = 30.0 bits (66), Expect = 1.4
Identities = 20/87 (22%), Positives = 37/87 (41%)
Frame = +1
Query: 423 ATRSGRVMSELKKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDK 482
A+ R+ + KKK+ EIV E V+ K + TP+ + P+ L K
Sbjct: 154 ASTKTRMKKKKKKKSMARPHHEIVAEEEVLDNKEEDITCCTPKAKRFRIPEVLTCPPAPK 333
Query: 483 KFSKFVDVFKKLHISIPFADALEQMPI 509
K +++ + +P ++ + PI
Sbjct: 334 K--------RRVTVPVPPCSSINRSPI 390
>TC14415 similar to UP|O04237 (O04237) Transcription factor, partial (83%)
Length = 1666
Score = 28.5 bits (62), Expect = 4.0
Identities = 26/116 (22%), Positives = 49/116 (41%), Gaps = 4/116 (3%)
Frame = +2
Query: 279 DSEECTETR----PEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQF 334
DSE+ +E R P + G R FSN R +G+G+GF+R+
Sbjct: 578 DSEKASERRGYGAPRGPYRGGGGGRRGGFSNGEADEEGRPRRAFERHSGTGRGSGFKREG 757
Query: 335 PSQGFQGQSLRQPQERGEGEGSGSKKSLEELVETFINRTENNYKNQEAANKNLENQ 390
+G G + + + + ++K+L + + + N + +AN+N E +
Sbjct: 758 AGRGNWGTQSDEIAQVTDEVANETEKNLSDEKPAVEDVADGN--KESSANENEEKE 919
>TC9194 similar to UP|Q9LYP1 (Q9LYP1) Membrane protein
(AT5g07250/T28J14_190), partial (78%)
Length = 1467
Score = 27.3 bits (59), Expect = 8.8
Identities = 18/44 (40%), Positives = 25/44 (55%), Gaps = 1/44 (2%)
Frame = -1
Query: 315 PNFSWRQGSSGQGNGFQRQFPSQGFQ-GQSLRQPQERGEGEGSG 357
PNFS R+G G G G R +GF + LR + +G+G+G G
Sbjct: 234 PNFSSREG--GDGFGLVR----RGFNLNEFLRN*ERKGKGKGGG 121
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,705,966
Number of Sequences: 28460
Number of extensions: 134986
Number of successful extensions: 646
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 640
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 645
length of query: 728
length of database: 4,897,600
effective HSP length: 97
effective length of query: 631
effective length of database: 2,136,980
effective search space: 1348434380
effective search space used: 1348434380
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0388a.1


