

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
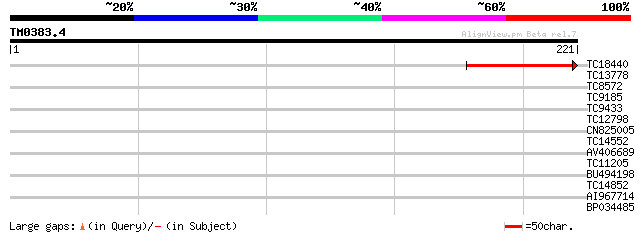
Query= TM0383.4
(221 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18440 88 1e-18
TC13778 29 0.73
TC8572 similar to UP|Q9SCM3 (Q9SCM3) 40S ribosomal protein S2 ho... 28 1.6
TC9185 28 1.6
TC9433 similar to UP|Q9M4Y9 (Q9M4Y9) AP2-related transcription f... 27 3.6
TC12798 similar to UP|O39876 (O39876) HBc antigen, partial (8%) 26 4.7
CN825005 26 4.7
TC14552 similar to UP|THM4_ARATH (Q9SEU6) Thioredoxin M-type 4, ... 26 6.2
AV406689 26 6.2
TC11205 homologue to UP|Q8LDN3 (Q8LDN3) F-box protein AtFBL5, pa... 26 6.2
BU494198 26 6.2
TC14852 similar to UP|Q9LZI2 (Q9LZI2) DTDP-glucose 4-6-dehydrata... 25 8.1
AI967714 25 8.1
BP034485 25 8.1
>TC18440
Length = 415
Score = 87.8 bits (216), Expect = 1e-18
Identities = 43/43 (100%), Positives = 43/43 (100%)
Frame = +2
Query: 179 SLSGSTKVEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDDE 221
SLSGSTKVEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDDE
Sbjct: 2 SLSGSTKVEDLSSGSHDADAKPKINNSGGGLLGLAYASSDDDE 130
>TC13778
Length = 521
Score = 28.9 bits (63), Expect = 0.73
Identities = 19/71 (26%), Positives = 32/71 (44%)
Frame = +1
Query: 93 RAASLVDSSPASVPLPLRVEPKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSV 152
+A ++V SS SV + + P P + ++ + PK+ +SES S SD
Sbjct: 10 KAVAVVKSSTKSVTVTKKSTPSPAPSLTKK-ATNSASKAPPKKRKSESISSEDDDSDYGT 186
Query: 153 TNRKPDHGRLK 163
++ P R K
Sbjct: 187 GSKAPAKKRKK 219
>TC8572 similar to UP|Q9SCM3 (Q9SCM3) 40S ribosomal protein S2 homolog
(At3g57490), partial (91%)
Length = 1076
Score = 27.7 bits (60), Expect = 1.6
Identities = 28/87 (32%), Positives = 38/87 (43%), Gaps = 4/87 (4%)
Frame = -3
Query: 86 RAFAAFKRAASLVD----SSPASVPLPLRVEPKPKSGIRQQDLLKKVVEIKPKRPRSESK 141
R F + R S V SS + P PL P+P S R + + +PK PRS
Sbjct: 255 RIFPSLTRRPSFVTGTHFSSSSRRPAPLLRPPRPLSPPRPRP-----PKPRPKPPRSPPP 91
Query: 142 QSTSASSDVSVTNRKPDHGRLKETGQS 168
+S S + NR+ R KET +S
Sbjct: 90 RSPPPRSAIDAVNRECGAER-KETLKS 13
>TC9185
Length = 555
Score = 27.7 bits (60), Expect = 1.6
Identities = 21/72 (29%), Positives = 31/72 (42%), Gaps = 6/72 (8%)
Frame = +1
Query: 108 PLRVEPKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRK------PDHGR 161
PL+ EP P S +D KP P S+ ST++SS S + + R
Sbjct: 88 PLKYEPPPVSSSGTKDK-------KPDVPSVNSQSSTASSSQSSANQTQGKLVFGSNPAR 246
Query: 162 LKETGQSLSGVK 173
K+TG++ K
Sbjct: 247 TKDTGKAKEAAK 282
>TC9433 similar to UP|Q9M4Y9 (Q9M4Y9) AP2-related transcription factor
(Ethylene responsive protein), partial (24%)
Length = 572
Score = 26.6 bits (57), Expect = 3.6
Identities = 24/68 (35%), Positives = 30/68 (43%), Gaps = 3/68 (4%)
Frame = -3
Query: 83 TEKRAFAAFKRAASLVDSSPASVPLPLRV---EPKPKSGIRQQDLLKKVVEIKPKRPRSE 139
T + A A+ RAA + S A + PLRV EP P V + KR SE
Sbjct: 552 TAEDAALAYDRAAYRMRGSRAMLNFPLRVNSGEPDP-------------VRVASKRSSSE 412
Query: 140 SKQSTSAS 147
S S+S S
Sbjct: 411 SCSSSSES 388
>TC12798 similar to UP|O39876 (O39876) HBc antigen, partial (8%)
Length = 735
Score = 26.2 bits (56), Expect = 4.7
Identities = 18/49 (36%), Positives = 27/49 (54%), Gaps = 3/49 (6%)
Frame = +2
Query: 165 TGQSLSGVKKVEEQSLSGS---TKVEDLSSGSHDADAKPKINNSGGGLL 210
+G S+S KKVE +++SGS TK G + + P +S GGL+
Sbjct: 122 SGGSVSARKKVEPRNVSGSRAGTKRSGSIDGLASSVSVPPRRSSTGGLV 268
>CN825005
Length = 665
Score = 26.2 bits (56), Expect = 4.7
Identities = 11/29 (37%), Positives = 17/29 (57%)
Frame = -3
Query: 135 RPRSESKQSTSASSDVSVTNRKPDHGRLK 163
R + S S+ + TNRKP++G+LK
Sbjct: 528 RLKPSSTHSSPTPHSIKETNRKPNNGKLK 442
>TC14552 similar to UP|THM4_ARATH (Q9SEU6) Thioredoxin M-type 4, chloroplast
precursor (TRX-M4), partial (55%)
Length = 1016
Score = 25.8 bits (55), Expect = 6.2
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = -3
Query: 193 SHDADAKPKINNSGGG 208
+HDADA INNSG G
Sbjct: 276 THDADACGAINNSGSG 229
>AV406689
Length = 418
Score = 25.8 bits (55), Expect = 6.2
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = -1
Query: 44 VAGERGGYLHGRGALDSDD 62
V GERGG + G GA D+D+
Sbjct: 148 VGGERGGCVAGGGAPDADE 92
>TC11205 homologue to UP|Q8LDN3 (Q8LDN3) F-box protein AtFBL5, partial (6%)
Length = 329
Score = 25.8 bits (55), Expect = 6.2
Identities = 14/37 (37%), Positives = 18/37 (47%)
Frame = +2
Query: 23 GKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALD 59
GKD + ED+ K+ VAG R G G+G D
Sbjct: 125 GKDNLRTEDLNLNLSFEKLMMVAGNRNG---GKGGGD 226
>BU494198
Length = 510
Score = 25.8 bits (55), Expect = 6.2
Identities = 23/101 (22%), Positives = 42/101 (40%), Gaps = 3/101 (2%)
Frame = +3
Query: 95 ASLVDSSPASVPLPLRVEPK---PKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVS 151
+S + S V P + K PK+G +Q+ K K K P+S K +T+ V
Sbjct: 42 SSKISRSKDDVSTPKSAKSKHETPKTGKSKQETSKIAAASKAKSPKS-GKSNTNGPGKVK 218
Query: 152 VTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTKVEDLSSG 192
+ K + + S V+ + ++ + S ++ SG
Sbjct: 219 SVSLKSTDSEDETSDDSTREVEDTKGKTSTSSKVGSEVKSG 341
>TC14852 similar to UP|Q9LZI2 (Q9LZI2) DTDP-glucose 4-6-dehydratase homolog
D18 (At3g62830/F26K9_260), partial (70%)
Length = 1470
Score = 25.4 bits (54), Expect = 8.1
Identities = 19/62 (30%), Positives = 24/62 (38%)
Frame = +1
Query: 100 SSPASVPLPLRVEPKPKSGIRQQDLLKKVVEIKPKRPRSESKQSTSASSDVSVTNRKPDH 159
SSP+S+PLP P P P RS S ST+++S T
Sbjct: 592 SSPSSLPLPPHPPPSPP---------------PPLMSRSRSLTSTTSTSSRRSTTASTPS 726
Query: 160 GR 161
GR
Sbjct: 727 GR 732
>AI967714
Length = 452
Score = 25.4 bits (54), Expect = 8.1
Identities = 22/62 (35%), Positives = 28/62 (44%)
Frame = +3
Query: 133 PKRPRSESKQSTSASSDVSVTNRKPDHGRLKETGQSLSGVKKVEEQSLSGSTKVEDLSSG 192
P R S+ +ASS S R K T +S V + +S ST ED+SSG
Sbjct: 282 PMRSGRSSQSGRAASSKGS---------RQKST---VSEVTDPSSKRVSKSTASEDISSG 425
Query: 193 SH 194
SH
Sbjct: 426 SH 431
>BP034485
Length = 485
Score = 25.4 bits (54), Expect = 8.1
Identities = 16/52 (30%), Positives = 22/52 (41%)
Frame = +3
Query: 11 NLSDPKDKKWGKGKDKIDDEDVTFQRMVAKMQEVAGERGGYLHGRGALDSDD 62
N + KK G G K+D D FQ +G +GG G+G + D
Sbjct: 216 NNKGKEGKKGGGGGGKLDKIDFDFQDFDIPSHGKSGGKGGGGGGKGNKNGHD 371
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.307 0.127 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,695,073
Number of Sequences: 28460
Number of extensions: 28860
Number of successful extensions: 127
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 127
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 127
length of query: 221
length of database: 4,897,600
effective HSP length: 87
effective length of query: 134
effective length of database: 2,421,580
effective search space: 324491720
effective search space used: 324491720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (22.0 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0383.4


