

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0383.12
(339 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
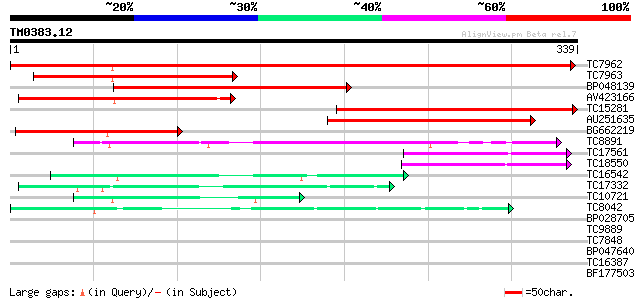
Sequences producing significant alignments: (bits) Value
TC7962 homologue to UP|Q84V97 (Q84V97) Alcohol dehydrogenase 1 ... 375 e-105
TC7963 homologue to UP|Q84V97 (Q84V97) Alcohol dehydrogenase 1 ... 145 1e-35
BP048139 142 6e-35
AV423166 140 3e-34
TC15281 similar to UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1 (F... 137 3e-33
AU251635 128 1e-30
BG662219 119 6e-28
TC8891 similar to GB|AAP68279.1|31711846|BT008840 At1g72680 {Ara... 72 2e-13
TC17561 66 1e-11
TC18550 64 5e-11
TC16542 similar to UP|Q93X81 (Q93X81) Sorbitol dehydrogenase, pa... 61 2e-10
TC17332 similar to UP|MTD_FRAAN (Q9ZRF1) Probable mannitol dehyd... 61 3e-10
TC10721 similar to UP|MTD_MEDSA (O82515) Probable mannitol dehyd... 53 6e-08
TC8042 homologue to UP|P93619 (P93619) Probable NADP-dependent o... 45 2e-05
BP028705 36 0.008
TC9889 similar to UP|Q9AYU1 (Q9AYU1) Quinone-oxidoreductase QR1 ... 31 0.34
TC7848 homologue to UP|Q9AVR8 (Q9AVR8) Sucrose synthase isoform ... 30 0.44
BP047640 29 0.99
TC16387 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, part... 28 2.9
BF177503 27 4.9
>TC7962 homologue to UP|Q84V97 (Q84V97) Alcohol dehydrogenase 1 , complete
Length = 1489
Score = 375 bits (962), Expect = e-105
Identities = 178/342 (52%), Positives = 243/342 (71%), Gaps = 4/342 (1%)
Frame = +1
Query: 1 MSSSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH 60
++ ST+ QVI C+AAV+W AG+PLVIEEV+V+PPQ E+R+K++ TSLC +D+ WE+
Sbjct: 106 IAMSTTAGQVIKCRAAVSWEAGKPLVIEEVEVAPPQAGEVRLKILYTSLCHTDVYFWEAK 285
Query: 61 ---PIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLG 117
P+FPRIFGHEA GIVESVG GVT K GDH L VF GEC EC C S +SN+C +L
Sbjct: 286 GQTPLFPRIFGHEAGGIVESVGEGVTHLKPGDHALPVFTGECGECPHCKSEESNMCDLLR 465
Query: 118 LERN-GLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSC 176
+ + G+M SD ++RFSIKGKP+YH+ G S+FSEYTV+H+GC K++P APL+K+C+LSC
Sbjct: 466 INTDRGVMISDNQSRFSIKGKPIYHFVGTSTFSEYTVLHAGCVAKINPAAPLDKVCILSC 645
Query: 177 GVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKA 236
G+ G GA NVA GS+V IFGLG VGL+ A+GA++ GASRIIGVD + E AK
Sbjct: 646 GICTGFGATVNVAKPKPGSSVAIFGLGAVGLAAAEGARVSGASRIIGVDLVSSRFEGAKK 825
Query: 237 FGITEVVDPNSYEEPIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGV 296
FG+ E V+P +++P+ +VI +T+GG D + EC G + +A + DGWG+ V +GV
Sbjct: 826 FGVNEFVNPKDHDKPVQEVIAEMTNGGVDRAVECTGSIQAMISAFECVHDGWGVAVLVGV 1005
Query: 297 PKVKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDKYV 338
P H L RTLKG+ +G +KP++DLP++V+ Y+
Sbjct: 1006PNQDDAFQTHPVNFLNERTLKGTFYGNYKPRTDLPNVVEMYM 1131
>TC7963 homologue to UP|Q84V97 (Q84V97) Alcohol dehydrogenase 1 , partial
(33%)
Length = 694
Score = 145 bits (365), Expect = 1e-35
Identities = 73/126 (57%), Positives = 94/126 (73%), Gaps = 4/126 (3%)
Frame = +3
Query: 15 AAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESH---PIFPRIFGHEA 71
AAV+W AG+PLVIEEV+V+PPQ E+R+K++ TSLC +D+ WE+ P+FPRIFGHEA
Sbjct: 315 AAVSWEAGKPLVIEEVEVAPPQAGEVRLKILYTSLCHTDVYFWEAKGQTPLFPRIFGHEA 494
Query: 72 SGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERN-GLMHSDQKT 130
GIVESVG GVT K DH L VF GEC E C S +SN+C +L + + G+M SD+++
Sbjct: 495 GGIVESVGEGVTHLKPVDHALPVFTGECGELPHCKSEESNMCDLLRINTDRGVMISDKQS 674
Query: 131 RFSIKG 136
RFSIKG
Sbjct: 675 RFSIKG 692
>BP048139
Length = 567
Score = 142 bits (359), Expect = 6e-35
Identities = 70/143 (48%), Positives = 93/143 (64%), Gaps = 1/143 (0%)
Frame = -2
Query: 63 FPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERNG 122
FPRI GHEA G+VESVG VTE +GD V+ +++ +C EC C S KSN C + +
Sbjct: 431 FPRILGHEAVGVVESVGKDVTEVTKGDVVIPIYLPDCGECTDCKSTKSNRCTNFPFKVSP 252
Query: 123 LMHSDQKTRFS-IKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSCGVAAG 181
M D TRF+ + G+ ++H+ +SSFSEYTVV K+ P P ++ CLLSCG++ G
Sbjct: 251 SMPRDGTTRFTDLNGEIIHHFMFISSFSEYTVVDVANVTKIDPEIPPDRACLLSCGISTG 72
Query: 182 LGAAWNVADVSKGSTVVIFGLGT 204
+GAAW A V GSTV IFGLG+
Sbjct: 71 VGAAWRTASVEPGSTVAIFGLGS 3
>AV423166
Length = 452
Score = 140 bits (353), Expect = 3e-34
Identities = 66/132 (50%), Positives = 90/132 (68%), Gaps = 2/132 (1%)
Frame = +2
Query: 6 STPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESHP--IF 63
S QVITCKAA+ WG GEP+ +EE+QV PP+ E+RVK++C S+C +DI++ P IF
Sbjct: 59 SKSQVITCKAAICWGLGEPVTVEEIQVDPPKATEVRVKMLCASICHTDITSIHGFPHGIF 238
Query: 64 PRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERNGL 123
P GHE GIVESVG VT KEGD ++ +IGEC EC+ C SG++N+C + +G+
Sbjct: 239 PLALGHEGVGIVESVGDQVTNLKEGDMIIPTYIGECQECENCVSGETNLCMKYPIALSGM 418
Query: 124 MHSDQKTRFSIK 135
M D +R SI+
Sbjct: 419 M-PDNTSRMSIR 451
>TC15281 similar to UP|Q8LJR2 (Q8LJR2) Alcohol dehydrogenase 1 (Fragment),
partial (50%)
Length = 785
Score = 137 bits (345), Expect = 3e-33
Identities = 66/144 (45%), Positives = 95/144 (65%)
Frame = +1
Query: 196 TVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYEEPIAQV 255
TV +FGLG VGL+ A+GA+L GASRIIGVD + E AK FG+T+ V+P +++P+ QV
Sbjct: 1 TVAVFGLGAVGLAAAEGARLSGASRIIGVDLVAGRFEQAKKFGVTDFVNPKDHDKPVQQV 180
Query: 256 INRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMGRT 315
I +T+G D + EC G+ +A + DGWG+ V +GVP H + GRT
Sbjct: 181 IVEMTNG*VDRAIECTGNIQATISAFECLHDGWGVAVIVGVPTKDAEFKTHPIHFMEGRT 360
Query: 316 LKGSMFGGWKPKSDLPSLVDKYVN 339
LKG+ +G ++ +SD+PSLV+KY+N
Sbjct: 361 LKGTFYGNFQTRSDIPSLVEKYMN 432
>AU251635
Length = 383
Score = 128 bits (322), Expect = 1e-30
Identities = 63/124 (50%), Positives = 84/124 (66%)
Frame = +3
Query: 191 VSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYEE 250
V GS V +FGLGTVGL+VA+GAK GASRIIG+D + K E AK FG+TE ++P +E+
Sbjct: 6 VEPGSIVAVFGLGTVGLAVAEGAKAAGASRIIGIDIDSNKFERAKNFGVTEFINPKEHEK 185
Query: 251 PIAQVINRITDGGADFSFECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLL 310
PI QVI +TDGG D+SFEC+G+ ++ AL+ C GWG +V +GV +S L
Sbjct: 186 PIQQVIVDLTDGGVDYSFECIGNVSVMRAALECCHKGWGTSVIVGVAASGQEISTRPFQL 365
Query: 311 LMGR 314
+ GR
Sbjct: 366 VTGR 377
>BG662219
Length = 366
Score = 119 bits (299), Expect = 6e-28
Identities = 57/103 (55%), Positives = 74/103 (71%), Gaps = 3/103 (2%)
Frame = +2
Query: 4 STSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAW---ESH 60
+T+ QVITCKAAVAW +PL IE+VQV+PPQ E+RVK++ T+LC +D W +
Sbjct: 56 ATTQGQVITCKAAVAWEPNKPLTIEDVQVAPPQAGEVRVKILYTALCHTDAYTWSGKDPE 235
Query: 61 PIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECK 103
+FP I GHEA+G+VESVG GVTE + GDHV+ + EC ECK
Sbjct: 236 GLFPCILGHEAAGVVESVGEGVTEVQPGDHVIPCYQAECRECK 364
>TC8891 similar to GB|AAP68279.1|31711846|BT008840 At1g72680 {Arabidopsis
thaliana;} , complete
Length = 1636
Score = 71.6 bits (174), Expect = 2e-13
Identities = 78/303 (25%), Positives = 124/303 (40%), Gaps = 11/303 (3%)
Frame = +1
Query: 39 EIRVKVVCTSLCRSDISAWE----SHPIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LT 93
++ VK+ +C +D+ AW + ++P + GHE +GIV VG V F GDHV +
Sbjct: 199 DVYVKISHCGVCYADV-AWTRNTLGNSVYPCVPGHEIAGIVTKVGSNVHGFSVGDHVGVG 375
Query: 94 VFIGECMECKPCTSGKSNVCQVLG--LERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEY 151
++ C +C+ C G + C V G NG+ H T+ +S++
Sbjct: 376 TYVNSCRDCEFCDDGLEHHC-VKGSVFTFNGVDHDGTITK--------------GGYSKF 510
Query: 152 TVVHSGCAVKVSPNAPLEKICLLSCGVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQ 211
VVH + + PL L C G ++ + GLG +G +
Sbjct: 511 IVVHERYCFLIPKSYPLASAAPLLCAGITVYSPMMRHKMNQPGKSLGVIGLGGLGHMAVK 690
Query: 212 GAKLKGASRIIGVDNNPQKCENAKAFGITEVVDPNSYEE--PIAQVINRITD-GGADFSF 268
K G + + + +K E G V + EE +A+ ++ I D SF
Sbjct: 691 FGKAFGLNVTVFSTSISKKEEALSLLGADRFVVSSDQEEMRALAKSLDFIVDTASGPHSF 870
Query: 269 ECVGDTDMITTALQSCCDGWGLTVTLGVPKVKPVMSAHYGLLLMG-RTLKGSMFGGWKPK 327
D L+S +G+ +G P + H GLL+MG RT+ GS GG K
Sbjct: 871 ------DPYMALLKS----FGVLALVGFP---GEIKIHPGLLIMGSRTVSGSGVGGTKEI 1011
Query: 328 SDL 330
D+
Sbjct: 1012RDM 1020
>TC17561
Length = 581
Score = 65.9 bits (159), Expect = 1e-11
Identities = 35/102 (34%), Positives = 60/102 (58%), Gaps = 1/102 (0%)
Frame = +3
Query: 236 AFGITEVVDPNSYEEPIAQVINRITDG-GADFSFECVGDTDMITTALQSCCDGWGLTVTL 294
AFG+T ++P + +++++ ++ G G D+S EC G ++T ++ + G G T+ +
Sbjct: 3 AFGMTHFINPGDSGKSLSELVKELSGGVGVDYSIECTGAPPLLTESVAATKVGTGKTIAV 182
Query: 295 GVPKVKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDK 336
G+ +PV+ LL GRTLKGS FGG K +DL + +K
Sbjct: 183 GMAP-EPVVPFGLLALLFGRTLKGSAFGGIKTTTDLSIIANK 305
>TC18550
Length = 688
Score = 63.5 bits (153), Expect = 5e-11
Identities = 35/103 (33%), Positives = 60/103 (57%), Gaps = 1/103 (0%)
Frame = +2
Query: 235 KAFGITEVVDPNSYEEPIAQVINRITDG-GADFSFECVGDTDMITTALQSCCDGWGLTVT 293
+AFG+T ++P + ++++ ++ G G D+SFEC G + T +L++ G G T+
Sbjct: 8 EAFGMTHSINPGDSTKSASELVKELSGGIGMDYSFECSGFAPLPTESLEATKVGTGETIA 187
Query: 294 LGVPKVKPVMSAHYGLLLMGRTLKGSMFGGWKPKSDLPSLVDK 336
+GV +P++ +L GRTLKGS+ GG K SD + +K
Sbjct: 188 IGV-GTEPIVPFAILSILYGRTLKGSILGGLKAISDFSIIANK 313
>TC16542 similar to UP|Q93X81 (Q93X81) Sorbitol dehydrogenase, partial (60%)
Length = 724
Score = 61.2 bits (147), Expect = 2e-10
Identities = 56/223 (25%), Positives = 87/223 (38%), Gaps = 9/223 (4%)
Frame = +1
Query: 25 LVIEEVQVSPPQPMEIRVKVVCTSLCRSDISAWESHPIF------PRIFGHEASGIVESV 78
L I+ ++ P ++RV++ +C SD+ + P + GHE +GI+E+V
Sbjct: 124 LKIQPFKLPTLGPHDVRVRMKAVGICGSDVHYLKKLRCADFIVKEPMVIGHECAGIIEAV 303
Query: 79 GLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIKGKP 138
G V GD V C C C G N+C + +H
Sbjct: 304 GAEVKSLVPGDRVAIEPGISCWRCDHCKLGSYNLCPDMKFFATPPVH------------- 444
Query: 139 VYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICL---LSCGVAAGLGAAWNVADVSKGS 195
S + V + K+ N LE+ + LS GV A A++ +
Sbjct: 445 -------GSLANQIVHPADLCFKLPDNVSLEEGAMCEPLSVGV-----HACRRANIGPET 588
Query: 196 TVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQKCENAKAFG 238
V+I G G +GL A+ GA RI+ VD + + AK G
Sbjct: 589 NVLIMGAGPIGLVTMLAARAFGAPRIVIVDVDDHRLSVAKTLG 717
>TC17332 similar to UP|MTD_FRAAN (Q9ZRF1) Probable mannitol dehydrogenase
(NAD-dependent mannitol dehydrogenase) , partial (63%)
Length = 736
Score = 60.8 bits (146), Expect = 3e-10
Identities = 59/234 (25%), Positives = 94/234 (39%), Gaps = 9/234 (3%)
Frame = +1
Query: 6 STPQVITCKAAVAWGAGEPL-VIEEVQVSPPQPME--IRVKVVCTSLCRSDI----SAWE 58
S P++ K A W A + V+ + S + E + KV+ +C SD+ + W
Sbjct: 46 SQPEMEHPKKAFGWAARDSSGVLSPFKFSRRETGEKDVTFKVLYCGICHSDLHMIKNEWG 225
Query: 59 SHPIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LTVFIGECMECKPCTSGKSNVCQVLG 117
S ++P + GHE +G+V VG V +FK GD V + + C C+ C N C
Sbjct: 226 SS-VYPLVPGHEIAGVVTEVGSKVQKFKVGDRVGVGCMVNSCRSCQSCADNLENYC---- 390
Query: 118 LERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKICLLSC- 176
K + K V +S+ V +++S N PLE C
Sbjct: 391 ----------PKMILTYAAKNVDGTITYGGYSDLMVSDEDFGIRISDNLPLEAAAPFLCA 540
Query: 177 GVAAGLGAAWNVADVSKGSTVVIFGLGTVGLSVAQGAKLKGASRIIGVDNNPQK 230
G+ + D G V + GLG +G + A GA ++ + +P K
Sbjct: 541 GITVYSPLKYYGLD-KPGLHVGVSGLGGLGHMAVKFAIALGA-KVTAISTSPNK 696
>TC10721 similar to UP|MTD_MEDSA (O82515) Probable mannitol dehydrogenase
(NAD-dependent mannitol dehydrogenase) , complete
Length = 1297
Score = 53.1 bits (126), Expect = 6e-08
Identities = 37/149 (24%), Positives = 60/149 (39%), Gaps = 11/149 (7%)
Frame = +3
Query: 39 EIRVKVVCTSLCRSDISAWESH---PIFPRIFGHEASGIVESVGLGVTEFKEGDHV-LTV 94
++ +K++ +C SD+ ++ +P + GHE G+V G VT+FK GD V + V
Sbjct: 189 DVAIKILFCGVCHSDLHTIKNDWGFTTYPVVPGHEIVGVVTKTGNNVTKFKVGDKVGVGV 368
Query: 95 FIGECMECKPCTSGKSNVCQVLGLERNGLMHSDQKTRFSIKGKPVYHYCGV-------SS 147
+ C C+ C N C +PVY Y
Sbjct: 369 IVDSCKSCENCDQDLENYCP----------------------QPVYTYNSPYNGTRTNGG 482
Query: 148 FSEYTVVHSGCAVKVSPNAPLEKICLLSC 176
+S++ VVH ++ N PL+ L C
Sbjct: 483 YSDFVVVHQRYVLQFPDNLPLDAGAPLLC 569
>TC8042 homologue to UP|P93619 (P93619) Probable NADP-dependent
oxidoreductase TED2 , complete
Length = 1244
Score = 45.1 bits (105), Expect = 2e-05
Identities = 73/310 (23%), Positives = 117/310 (37%), Gaps = 9/310 (2%)
Frame = +2
Query: 1 MSSSTSTPQVITCKAAVAWGAGEPLVIEEVQVSPPQPMEIRVKVVCTSL------CRSDI 54
+SSS+S +++ G + L E+V++ P+ E+RVK + R +
Sbjct: 89 LSSSSSATEMVKAIRVHELGGPQILKWEDVEIGDPKENEVRVKNKAIGVNFIDVYFRKGV 268
Query: 55 SAWESHPIFPRIFGHEASGIVESVGLGVTEFKEGDHVLTVFIGECMECKPCTSGKSNVCQ 114
+ P P G EA G+V +VG G T + GD V
Sbjct: 269 YKASTSPFTP---GMEAVGVVTAVGGGPTGVQVGDLV----------------------- 370
Query: 115 VLGLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKV-SPNAPLEKICL 173
G+P + S+ E ++H+G V + S P +
Sbjct: 371 ------------------GYAGQP------IGSYVEEQILHAGKVVPIPSSIDPSVAASV 478
Query: 174 LSCGVAAGLGAAWNVADVSKGSTVVIF-GLGTVGLSVAQGAKLKGASRIIGVDNNPQKCE 232
+ G+ A V +G T+++ G VG + Q A GA+ +IG +N +K
Sbjct: 479 MLKGMTAQF-LLRRCFKVERGHTILVHAAAGGVGSLLCQWANALGAT-VIGTVSNEEKAA 652
Query: 233 NAKAFGITEVVDPNSYEEPIAQVINRITDG-GADFSFECVGDTDMITTALQSCCDGWGLT 291
AK G V+ EE +N IT G G + ++ VG D +L +C G
Sbjct: 653 QAKEDGCHHVIIYK--EEDFVARVNEITSGNGVEVVYDSVG-KDTFEGSL-ACLKQRGYM 820
Query: 292 VTLGVPKVKP 301
V+ G P
Sbjct: 821 VSFGQSSGSP 850
>BP028705
Length = 474
Score = 36.2 bits (82), Expect = 0.008
Identities = 15/29 (51%), Positives = 23/29 (78%)
Frame = -1
Query: 311 LMGRTLKGSMFGGWKPKSDLPSLVDKYVN 339
L G+TL G+ FG +KPK+ LPS+VD +++
Sbjct: 444 LDGKTLTGTAFGDYKPKTGLPSVVDLFMS 358
>TC9889 similar to UP|Q9AYU1 (Q9AYU1) Quinone-oxidoreductase QR1
(Fragment), partial (49%)
Length = 528
Score = 30.8 bits (68), Expect = 0.34
Identities = 28/83 (33%), Positives = 40/83 (47%), Gaps = 8/83 (9%)
Frame = +1
Query: 18 AWGAGEP-LVIEEVQVSPPQPMEIRVKVVCTSLCRSD--ISAWESHPIF-PRIFGH---- 69
A+G G L EV V P+ E+ +++ S+ D I P+F PR F H
Sbjct: 46 AYGGGAAGLKHVEVPVPSPKKNEVLLRLEAASINPVDWKIQKGVLRPLFLPRSFPHIPCT 225
Query: 70 EASGIVESVGLGVTEFKEGDHVL 92
+ +G V VG V +FK GD V+
Sbjct: 226 DVAGEVAEVGPQVKDFKVGDKVI 294
>TC7848 homologue to UP|Q9AVR8 (Q9AVR8) Sucrose synthase isoform 3 ,
complete
Length = 2779
Score = 30.4 bits (67), Expect = 0.44
Identities = 11/21 (52%), Positives = 17/21 (80%)
Frame = -3
Query: 160 VKVSPNAPLEKICLLSCGVAA 180
+K+SP APL+ CLLS G+++
Sbjct: 290 LKISPKAPLKSFCLLSSGISS 228
>BP047640
Length = 511
Score = 29.3 bits (64), Expect = 0.99
Identities = 12/16 (75%), Positives = 13/16 (81%)
Frame = -1
Query: 58 ESHPIFPRIFGHEASG 73
ES +PRIFGHEASG
Sbjct: 232 ESQRAYPRIFGHEASG 185
>TC16387 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, partial (28%)
Length = 530
Score = 27.7 bits (60), Expect = 2.9
Identities = 20/68 (29%), Positives = 28/68 (40%)
Frame = -2
Query: 112 VCQVLGLERNGLMHSDQKTRFSIKGKPVYHYCGVSSFSEYTVVHSGCAVKVSPNAPLEKI 171
+C +G+ L S S KPV G + T V G +KVSP L +
Sbjct: 247 LCLAIGIF--SLTESTNDKSISATAKPVDERSG----NRLTTVPHGSIIKVSP*LSLFSL 86
Query: 172 CLLSCGVA 179
C+ +C A
Sbjct: 85 CMPACAAA 62
>BF177503
Length = 355
Score = 26.9 bits (58), Expect = 4.9
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +1
Query: 14 KAAVAWGAGEPLVIEEVQVSPPQPMEIRVK 43
+ AV W +PL IEE + P+ E+ +K
Sbjct: 223 RGAVFWEPNKPLTIEEFHMPRPEAGEVLIK 312
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,635,478
Number of Sequences: 28460
Number of extensions: 96995
Number of successful extensions: 455
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 444
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 445
length of query: 339
length of database: 4,897,600
effective HSP length: 91
effective length of query: 248
effective length of database: 2,307,740
effective search space: 572319520
effective search space used: 572319520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0383.12


