

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0381.8
(460 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
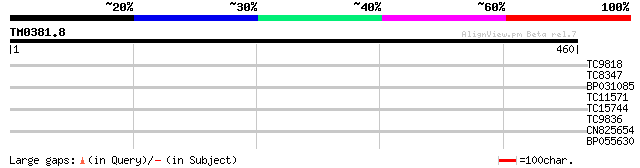
TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, ... 30 1.1
TC8347 UP|ATPH_LOTJA (P12231) ATP synthase C chain (Lipid-bindi... 29 1.9
BP031085 28 4.2
TC11571 similar to UP|Q940U9 (Q940U9) AT5g17440/K3M16_10, partia... 27 5.4
TC15744 homologue to UP|Q00508 (Q00508) Ribosomal protein L41 (R... 27 7.1
TC9836 weakly similar to UP|Q9LXV0 (Q9LXV0) Glucosyltransferase-... 27 9.2
CN825654 27 9.2
BP055630 27 9.2
>TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 1162
Score = 29.6 bits (65), Expect = 1.1
Identities = 15/64 (23%), Positives = 30/64 (46%)
Frame = +2
Query: 119 GISFDFWSITICRMLVGVGEASFISLAAPFIDDNAPVAQKTAWLATFYMCIPAGTALGYV 178
G S ++W + R + G+G + +A + + +P + + + + I G LGY+
Sbjct: 491 GFSPNYWFLMFGRFIAGIGIGYALMIAPVYTAEVSPASSRGFLTSFPEVFINGGILLGYI 670
Query: 179 YGFA 182
FA
Sbjct: 671 SNFA 682
>TC8347 UP|ATPH_LOTJA (P12231) ATP synthase C chain (Lipid-binding
protein) (Subunit III) , complete
Length = 1267
Score = 28.9 bits (63), Expect = 1.9
Identities = 11/43 (25%), Positives = 25/43 (57%)
Frame = +3
Query: 240 QVYIINVLGYISYNFVIGAYSYWGPKAGYSIYHMSNPDLLFGG 282
+++II +G+I Y+F + Y Y+ P + ++ + ++L G
Sbjct: 756 KIHIIFPVGFICYSFFLKFYYYFNPTIIFFLFEQEHKEILI*G 884
>BP031085
Length = 409
Score = 27.7 bits (60), Expect = 4.2
Identities = 12/28 (42%), Positives = 18/28 (63%)
Frame = +2
Query: 323 FCLVAFLFKNLSGFIVFFSIGELLIFAT 350
FCLV L + S +VFF +G L +F++
Sbjct: 56 FCLVLHLILSSSSSLVFFPLGLLFLFSS 139
>TC11571 similar to UP|Q940U9 (Q940U9) AT5g17440/K3M16_10, partial (67%)
Length = 1011
Score = 27.3 bits (59), Expect = 5.4
Identities = 19/54 (35%), Positives = 25/54 (46%)
Frame = -2
Query: 295 GGLILDRMTSTISNAFKLLSGATLLGAIFCLVAFLFKNLSGFIVFFSIGELLIF 348
G IL R+ + KLL G +L FCL+ N S F S+ LLI+
Sbjct: 914 GSCIL*RIQYWLLPGRKLLKGFSLFHCFFCLI-----NQSLLAKFLSLSHLLIY 768
>TC15744 homologue to UP|Q00508 (Q00508) Ribosomal protein L41 (Ribosomal
protein L44), complete
Length = 575
Score = 26.9 bits (58), Expect = 7.1
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = +2
Query: 110 VWTFAVAGCGISFDFW 125
V+ F V+GC +S DFW
Sbjct: 422 VYVFEVSGCFVSLDFW 469
>TC9836 weakly similar to UP|Q9LXV0 (Q9LXV0) Glucosyltransferase-like
protein, partial (55%)
Length = 1286
Score = 26.6 bits (57), Expect = 9.2
Identities = 10/29 (34%), Positives = 17/29 (58%)
Frame = +2
Query: 319 LGAIFCLVAFLFKNLSGFIVFFSIGELLI 347
+ A+FC V F N+S ++ S+G + I
Sbjct: 1145 ISALFCFVLFFE*NISALLILMSVGFMYI 1231
>CN825654
Length = 580
Score = 26.6 bits (57), Expect = 9.2
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = +2
Query: 428 DLSTEDDEDQSATSLRGKMKPLLEGN 453
+L+ ED+ED+ S GK KP G+
Sbjct: 101 ELTGEDEEDEQVVSFAGKKKPSKGGS 178
>BP055630
Length = 546
Score = 26.6 bits (57), Expect = 9.2
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = +1
Query: 336 FIVFFSIGELLIFATQAPVNYVSLRCVKPSLRPLSMAISTVS 377
F + FS+ LLI T AP+ L V S LSM+ +++S
Sbjct: 268 FFLTFSLVSLLIAVTWAPLVERGLGSVSKSFTTLSMSATSLS 393
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,145,747
Number of Sequences: 28460
Number of extensions: 115768
Number of successful extensions: 535
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 531
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 535
length of query: 460
length of database: 4,897,600
effective HSP length: 93
effective length of query: 367
effective length of database: 2,250,820
effective search space: 826050940
effective search space used: 826050940
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0381.8


