

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0377b.5
(372 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
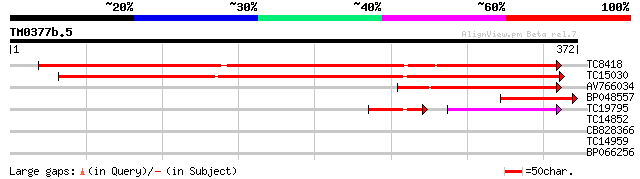
Score E
Sequences producing significant alignments: (bits) Value
TC8418 weakly similar to UP|Q8LFH1 (Q8LFH1) Aldose 1-epimerase-l... 380 e-106
TC15030 similar to UP|Q9LVH6 (Q9LVH6) Aldose 1-epimerase-like pr... 316 3e-87
AV766034 142 1e-34
BP048557 103 4e-23
TC19795 similar to UP|Q9LVH6 (Q9LVH6) Aldose 1-epimerase-like pr... 58 2e-15
TC14852 similar to UP|Q9LZI2 (Q9LZI2) DTDP-glucose 4-6-dehydrata... 30 0.65
CB828366 28 2.5
TC14959 similar to GB|AAN64529.1|24797034|BT001138 At5g58299/At5... 28 3.2
BP066256 26 9.3
>TC8418 weakly similar to UP|Q8LFH1 (Q8LFH1) Aldose 1-epimerase-like
protein, partial (87%)
Length = 1344
Score = 380 bits (977), Expect = e-106
Identities = 186/343 (54%), Positives = 246/343 (71%)
Frame = +2
Query: 20 VNGSAKKEKDHIRLFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTN 79
V A+ + I +ELKKG+ L ++N+GAT+VS+++PDKHG L D+ LGYD++ +Y N
Sbjct: 92 VLSQAQANDNKIGFYELKKGNFRLNLTNYGATVVSVIVPDKHGNLADIALGYDTVDEYKN 271
Query: 80 DSTYFGTTAGRVANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEG 139
D YFG GRVANRIG AQFTL+G YKL AN+ +NTLHGGP GF DV+W V+ + ++
Sbjct: 272 DGVYFGALVGRVANRIGNAQFTLDGKTYKLPANDKDNTLHGGPTGFGDVVWTVKTHKEDS 451
Query: 140 DKPRITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWN 199
ITF+YHSFD E+GFPG L V+V+Y++ N L + M A ++KPTPVNL H YWN
Sbjct: 452 ---HITFTYHSFDNEQGFPGQLEVSVTYMIIGTNKLGVRMIATPVDKPTPVNLAQHTYWN 622
Query: 200 LGNHNSGNILGEVVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKT 259
L H+SG+IL VQIFGS+IT +D LIPTGE VKGTPYDFL+P+ VG++I+ LP
Sbjct: 623 LRGHDSGDILSHTVQIFGSKITPVDDELIPTGEFQEVKGTPYDFLEPKEVGSQIHDLPGL 802
Query: 260 NGYDINYVLDGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFV 319
YD+NYVLD + L K+ AIV D SGR ++L++N PG+Q+YT+ +K+ KGKGG
Sbjct: 803 --YDVNYVLDPSSPQHLRKV-AIVKDNVSGRKLELWSNQPGVQYYTSGMLKDTKGKGGAT 973
Query: 320 YQPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFS 362
Y +G+ LE+Q FPDSVNHPNFPS IV P + YKH M+ +F+
Sbjct: 974 YHQYSGIALETQGFPDSVNHPNFPSMIVRPGQTYKHYMVYRFT 1102
>TC15030 similar to UP|Q9LVH6 (Q9LVH6) Aldose 1-epimerase-like protein,
partial (94%)
Length = 1212
Score = 316 bits (810), Expect = 3e-87
Identities = 161/333 (48%), Positives = 216/333 (64%), Gaps = 1/333 (0%)
Frame = +3
Query: 33 LFELKKGDLSLKVSNWGATLVSLVLPDKHGKLGDVVLGYDSLKQYTND-STYFGTTAGRV 91
LFEL G + + ++N G T+ SL +P K G L DVVLG DS++ Y S YFG GRV
Sbjct: 33 LFELNNGAMQVLITNLGCTITSLSVPAKDGVLSDVVLGLDSVESYQKGLSPYFGCIVGRV 212
Query: 92 ANRIGGAQFTLNGIHYKLIANEGNNTLHGGPRGFSDVLWKVERYVKEGDKPRITFSYHSF 151
ANRI +FTL+G+ Y L N N+LHGG GF +W+V Y ++G+ P ITF Y S
Sbjct: 213 ANRIKDGKFTLDGVEYSLPINNPPNSLHGGNVGFDKKIWEVIEY-RKGETPSITFKYQSH 389
Query: 152 DGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILGE 211
DGEEG+PG++ VT +Y L S +L + M+ +KPT +NL H YWNL H+SGNIL
Sbjct: 390 DGEEGYPGEITVTATYTLTSNTTLRLDMEGVPKDKPTIINLAQHTYWNLAGHSSGNILDH 569
Query: 212 VVQIFGSRITVLDSHLIPTGEIASVKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGG 271
+QI+ + IT +D + +PTGEI VKGTP+DF + +G I+Q+ GYD NYVLD G
Sbjct: 570 SIQIWANHITPVDQNTVPTGEIMPVKGTPFDFTSQKRIGDTISQVGM--GYDHNYVLDSG 743
Query: 272 KSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRAGLCLESQ 331
+ K ++ AA V + S RV+ L+TN PG+QFYTAN V GK G VY AGLCLE+Q
Sbjct: 744 EEKEGLRHAAKVKEPSSSRVLNLWTNVPGMQFYTANYVDGVAGKAGAVYGKHAGLCLETQ 923
Query: 332 VFPDSVNHPNFPSSIVTPEKPYKHVMLLKFSTK 364
FP+++N P+FPS +V P + Y+H M +FS +
Sbjct: 924 GFPNAINQPDFPSIVVRPGEKYQHSMSYEFSVE 1022
>AV766034
Length = 475
Score = 142 bits (357), Expect = 1e-34
Identities = 74/108 (68%), Positives = 83/108 (76%)
Frame = -2
Query: 255 QLPKTNGYDINYVLDGGKSKGLIKLAAIVMDKKSGRVMKLFTNAPGLQFYTANKVKNEKG 314
+L T GY +NYVLDG K K IKLAA V DKKSGR + L+ APG QF T N V++ KG
Sbjct: 456 KLAATKGYGLNYVLDGEKGKK-IKLAAKVFDKKSGRELVLYPKAPGGQF*TGNFVQDVKG 280
Query: 315 KGGFVYQPRAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFS 362
KGGFVYQ AGL LE+Q FPDSVN PNFPS+IVTPEKPYKH+ML KFS
Sbjct: 279 KGGFVYQAHAGLGLETQAFPDSVNPPNFPSTIVTPEKPYKHLMLFKFS 136
>BP048557
Length = 257
Score = 103 bits (258), Expect = 4e-23
Identities = 49/50 (98%), Positives = 49/50 (98%)
Frame = -3
Query: 323 RAGLCLESQVFPDSVNHPNFPSSIVTPEKPYKHVMLLKFSTKAPYAFSQS 372
RAGLCLESQV PDSVNHPNFPSSIVTPEKPYKHVMLLKFSTKAPYAFSQS
Sbjct: 255 RAGLCLESQVXPDSVNHPNFPSSIVTPEKPYKHVMLLKFSTKAPYAFSQS 106
>TC19795 similar to UP|Q9LVH6 (Q9LVH6) Aldose 1-epimerase-like protein,
partial (20%)
Length = 545
Score = 57.8 bits (138), Expect(2) = 2e-15
Identities = 31/76 (40%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Frame = +3
Query: 288 SGRVMKLFTNAPGLQFYTANKVKNEKGKGGFVYQPRA-GLCLESQVFPDSVNHPNFPSSI 346
S RV+ L+TN PG+ FYTAN V GK G VY + P PNFPS +
Sbjct: 156 SSRVLNLWTNVPGMPFYTANYVDGVAGKEGAVYAKACWAYVWRPRDSPMP*TTPNFPSIV 335
Query: 347 VTPEKPYKHVMLLKFS 362
+ P + Y+H +L +FS
Sbjct: 336 IRPGEKYQHTLLYEFS 383
Score = 40.8 bits (94), Expect(2) = 2e-15
Identities = 21/39 (53%), Positives = 26/39 (65%)
Frame = +1
Query: 236 VKGTPYDFLKPQVVGARINQLPKTNGYDINYVLDGGKSK 274
VKGTP+DF + +G I+QL GYD NYVLD G+ K
Sbjct: 4 VKGTPFDFTSQKRIGDTISQLGM--GYDHNYVLDCGEEK 114
>TC14852 similar to UP|Q9LZI2 (Q9LZI2) DTDP-glucose 4-6-dehydratase homolog
D18 (At3g62830/F26K9_260), partial (70%)
Length = 1470
Score = 30.0 bits (66), Expect = 0.65
Identities = 22/80 (27%), Positives = 40/80 (49%), Gaps = 10/80 (12%)
Frame = +3
Query: 159 GDLLVTVSY-ILGSKNSLSIIMKAKALNKPTPVNLINHAYWNLGNHNSGNILGEV---VQ 214
G + TV + ++ S +S S +LN+P P++ NH Y H+ + +G++ ++
Sbjct: 570 GVAIATVFFTVVPSSSSSSSSFTTTSLNEPVPISYFNHEYKQPAFHHRIHSVGKIPLGIK 749
Query: 215 IFGSRITV------LDSHLI 228
G RI V + SHL+
Sbjct: 750 RKGLRIVVTGGAGFVGSHLV 809
>CB828366
Length = 568
Score = 28.1 bits (61), Expect = 2.5
Identities = 19/63 (30%), Positives = 26/63 (41%)
Frame = -1
Query: 130 WKVERYVKEGDKPRITFSYHSFDGEEGFPGDLLVTVSYILGSKNSLSIIMKAKALNKPTP 189
W + K+ D PR H + G GD+ S G N L K + + PTP
Sbjct: 439 WTMYHNEKQRDVPRARMRPHKREPLAG-GGDIYFDKSMCRGQLNILLPSYKVRIIAPPTP 263
Query: 190 VNL 192
VN+
Sbjct: 262 VNV 254
>TC14959 similar to GB|AAN64529.1|24797034|BT001138 At5g58299/At5g58299
{Arabidopsis thaliana;}, partial (16%)
Length = 1015
Score = 27.7 bits (60), Expect = 3.2
Identities = 13/39 (33%), Positives = 19/39 (48%)
Frame = -1
Query: 84 FGTTAGRVANRIGGAQFTLNGIHYKLIANEGNNTLHGGP 122
+G T G IGG L G+ + + ++ LHGGP
Sbjct: 511 WGVTLGFFGRVIGGLSHALIGLTAVHVLDHAHHVLHGGP 395
>BP066256
Length = 335
Score = 26.2 bits (56), Expect = 9.3
Identities = 14/32 (43%), Positives = 17/32 (52%)
Frame = +2
Query: 305 TANKVKNEKGKGGFVYQPRAGLCLESQVFPDS 336
T NK N K K QP LC ESQ++P +
Sbjct: 227 TQNKTSNPK*KHAL--QPN*RLCKESQIYPSN 316
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,067,800
Number of Sequences: 28460
Number of extensions: 79015
Number of successful extensions: 360
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 350
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 351
length of query: 372
length of database: 4,897,600
effective HSP length: 92
effective length of query: 280
effective length of database: 2,279,280
effective search space: 638198400
effective search space used: 638198400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0377b.5


