

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0372b.7
(224 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
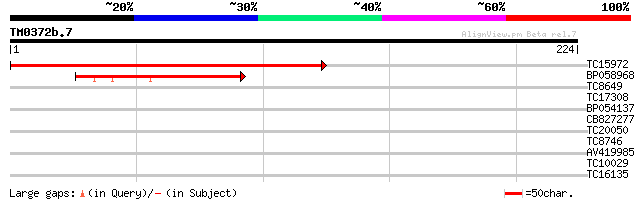
Score E
Sequences producing significant alignments: (bits) Value
TC15972 similar to GB|AAO63298.1|28950749|BT005234 At5g26220 {Ar... 273 1e-74
BP058968 90 3e-19
TC8649 similar to PIR|T52450|T52450 ribosomal protein S9 [import... 29 0.75
TC17308 weakly similar to UP|Q9LQV9 (Q9LQV9) F10B6.11, partial (... 28 1.7
BP054137 27 3.7
CB827277 26 6.3
TC20050 weakly similar to UP|Q9SFG3 (Q9SFG3) F2O10.5 protein (At... 26 6.3
TC8746 similar to PIR|T06396|T06396 isoprenylated protein - soyb... 26 6.3
AV419985 25 8.3
TC10029 25 8.3
TC16135 UP|PSAB_LOTJA (P58385) Photosystem I P700 chlorophyll A ... 25 8.3
>TC15972 similar to GB|AAO63298.1|28950749|BT005234 At5g26220 {Arabidopsis
thaliana;}, partial (58%)
Length = 570
Score = 273 bits (699), Expect = 1e-74
Identities = 125/125 (100%), Positives = 125/125 (100%)
Frame = +2
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE
Sbjct: 194 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 373
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD
Sbjct: 374 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 553
Query: 121 KVNNK 125
KVNNK
Sbjct: 554 KVNNK 568
>BP058968
Length = 508
Score = 90.1 bits (222), Expect = 3e-19
Identities = 48/70 (68%), Positives = 51/70 (72%), Gaps = 3/70 (4%)
Frame = +1
Query: 27 IKDYRR-VFDLACI-DHRGTPEHPARTCTL-EEKEGEICWGAVYCVRGGPEKEKLVMQYL 83
+KDY VFDLA EHP + +KEGEICWGAVYCVRGGPEKEKLVMQYL
Sbjct: 298 LKDYXTLVFDLAVH*SXEVHLEHPX*NLHIXRQKEGEICWGAVYCVRGGPEKEKLVMQYL 477
Query: 84 ERRECEYDHK 93
ERRECEYDHK
Sbjct: 478 ERRECEYDHK 507
>TC8649 similar to PIR|T52450|T52450 ribosomal protein S9 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(81%)
Length = 939
Score = 28.9 bits (63), Expect = 0.75
Identities = 14/28 (50%), Positives = 17/28 (60%), Gaps = 5/28 (17%)
Frame = +3
Query: 197 PVQIQHSPHAPIPALQL-----LPMPEP 219
P+ +QHS H+P P L LPMPEP
Sbjct: 60 PLSLQHSLHSPSPPTFLTTHKPLPMPEP 143
>TC17308 weakly similar to UP|Q9LQV9 (Q9LQV9) F10B6.11, partial (21%)
Length = 587
Score = 27.7 bits (60), Expect = 1.7
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = +1
Query: 95 LVDFYKEGDSSHP 107
+VDFY EGD SHP
Sbjct: 139 IVDFYNEGDHSHP 177
>BP054137
Length = 474
Score = 26.6 bits (57), Expect = 3.7
Identities = 10/23 (43%), Positives = 17/23 (73%)
Frame = +1
Query: 197 PVQIQHSPHAPIPALQLLPMPEP 219
P++ +H H+P+P+ LLP+P P
Sbjct: 346 PLRCRH--HSPLPSSPLLPLPPP 408
>CB827277
Length = 546
Score = 25.8 bits (55), Expect = 6.3
Identities = 21/83 (25%), Positives = 31/83 (37%)
Frame = +2
Query: 140 IATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRKVLGVGNVLPNDKKLSGPVQ 199
+ T + P G D+ + + G D + +A R + P Q
Sbjct: 257 MGTTYIPSGKQPDWKHIPTSSAMGAGEGDMNSMNMASSQRNPANM------------PSQ 400
Query: 200 IQHSPHAPIPALQLLPMPEPIAL 222
IQH P LLPMP P+A+
Sbjct: 401 IQHLA----PGSPLLPMPSPMAM 457
>TC20050 weakly similar to UP|Q9SFG3 (Q9SFG3) F2O10.5 protein (At3g05990),
partial (25%)
Length = 596
Score = 25.8 bits (55), Expect = 6.3
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = +3
Query: 204 PHAPIPALQLLPMPEP 219
PH P P L LLP+P P
Sbjct: 3 PHLPPPLLTLLPLPLP 50
>TC8746 similar to PIR|T06396|T06396 isoprenylated protein - soybean
(fragment) {Glycine max;}, partial (30%)
Length = 594
Score = 25.8 bits (55), Expect = 6.3
Identities = 9/26 (34%), Positives = 15/26 (57%)
Frame = -2
Query: 197 PVQIQHSPHAPIPALQLLPMPEPIAL 222
P ++ PH P P + + P P P+A+
Sbjct: 569 PKEMPPPPHVPPPPVPMCPGPPPVAV 492
>AV419985
Length = 408
Score = 25.4 bits (54), Expect = 8.3
Identities = 9/23 (39%), Positives = 13/23 (56%)
Frame = +3
Query: 202 HSPHAPIPALQLLPMPEPIALDS 224
HSP +P+P P P P+ D+
Sbjct: 156 HSPPSPLPPSPCAPAPPPMGADA 224
>TC10029
Length = 684
Score = 25.4 bits (54), Expect = 8.3
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = -2
Query: 62 CWGAVYCVRGGPEKEK 77
C GA CVRGG KE+
Sbjct: 98 CGGAYICVRGGGRKEE 51
>TC16135 UP|PSAB_LOTJA (P58385) Photosystem I P700 chlorophyll A apoprotein A2
(PsaB) (PSI-B), complete
Length = 2647
Score = 25.4 bits (54), Expect = 8.3
Identities = 13/42 (30%), Positives = 20/42 (46%)
Frame = +1
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTP 45
W+F +G LVW GF + GY ++ + + H TP
Sbjct: 1942 WMFLFGHLVWATGFMFLISWRGY---WQELIETLAWAHERTP 2058
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,577,252
Number of Sequences: 28460
Number of extensions: 66994
Number of successful extensions: 356
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 352
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 355
length of query: 224
length of database: 4,897,600
effective HSP length: 87
effective length of query: 137
effective length of database: 2,421,580
effective search space: 331756460
effective search space used: 331756460
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0372b.7


