

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
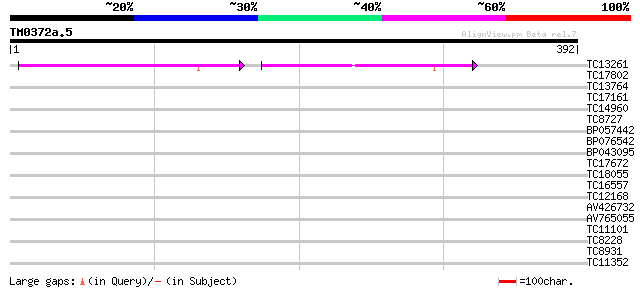
Query= TM0372a.5
(392 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC13261 weakly similar to UP|Q8S1K4 (Q8S1K4) PDI-like protein, p... 114 3e-26
TC17802 similar to UP|Q9XII2 (Q9XII2) At2g16050 protein, partial... 39 0.002
TC13764 similar to UP|Q9C9Y6 (Q9C9Y6) Thioredoxin-like protein, ... 37 0.007
TC17161 homologue to UP|THIF_PEA (P29450) Thioredoxin F-type, ch... 35 0.028
TC14960 similar to UP|Q8GUR9 (Q8GUR9) Thioredoxin h, partial (98%) 34 0.048
TC8727 33 0.063
BP057442 33 0.082
BP076542 33 0.082
BP043095 33 0.082
TC17672 similar to UP|Q93X83 (Q93X83) Hsp70 interacting protein/... 33 0.11
TC18055 similar to UP|CAE46765 (CAE46765) NADPH thioredoxin redu... 29 1.2
TC16557 28 2.6
TC12168 similar to UP|Q8L7S9 (Q8L7S9) At1g43560/T10P12_4, partia... 28 2.6
AV426732 28 3.4
AV765055 28 3.4
TC11101 27 4.5
TC8228 similar to UP|GPX4_CITSI (Q06652) Probable phospholipid h... 27 5.9
TC8931 similar to GB|AAL09731.1|15982767|AY057490 At2g32240/F22D... 27 7.7
TC11352 26 10.0
>TC13261 weakly similar to UP|Q8S1K4 (Q8S1K4) PDI-like protein, partial
(17%)
Length = 692
Score = 114 bits (285), Expect = 3e-26
Identities = 63/156 (40%), Positives = 89/156 (56%), Gaps = 7/156 (4%)
Frame = +1
Query: 175 LEELLAHEGRDFLISGDDRKVPVSELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKAT 234
L LLA + RDFL++ +V VS+L GK +G F A W PP R F L VY +LK++
Sbjct: 226 LSHLLASKDRDFLLAPSGAQVKVSDLEGKIVGFLFAANWYPPCRGFIQLLAGVYEDLKSS 405
Query: 235 KGNCFEIVLISTDRDHKEFNVNRSSTPWLAIPYED-RTRHDLCRIYDIKGIPALVLIGP- 292
+ FEIV +S+D D FN + PWLAIP+ D T+ L R YD++GIP L+++ P
Sbjct: 406 VPH-FEIVYVSSDEDMDAFNSFYETMPWLAIPFSDLETKKALTRKYDVEGIPCLIMLQPA 582
Query: 293 -----DGKVISLNGKFMVTSYGADAFPFTESRIRDL 323
+ NG +V YG A+PF++ R+ L
Sbjct: 583 KDHVKEDDATLRNGVELVHRYGTQAYPFSKERVEQL 690
Score = 110 bits (275), Expect = 4e-25
Identities = 58/164 (35%), Positives = 96/164 (58%), Gaps = 8/164 (4%)
Frame = +1
Query: 7 EAKHIDSHDVLKILAAEGAEYLLSCEG-KVPLSECDGKVICLFFSANWCRPCRAFVPRLV 65
E + + S + +LA++ ++LL+ G +V +S+ +GK++ F+ANW PCR F+ L
Sbjct: 199 EEEQVSSLKLSHLLASKDRDFLLAPSGAQVKVSDLEGKIVGFLFAANWYPPCRGFIQLLA 378
Query: 66 ELYETLRKRGINLEIIFVSFDREEDGFNEHLKSMPWLAVPF-DVNLHRRLIDRYQVDQIP 124
+YE L+ + EI++VS D + D FN ++MPWLA+PF D+ + L +Y V+ IP
Sbjct: 379 GVYEDLKSSVPHFEIVYVSSDEDMDAFNSFYETMPWLAIPFSDLETKKALTRKYDVEGIP 558
Query: 125 SFIPL------YSEDLIVEKNLIECIEDYGADAFPFTRKRHEEL 162
I L ED +N +E + YG A+PF+++R E+L
Sbjct: 559 CLIMLQPAKDHVKEDDATLRNGVELVHRYGTQAYPFSKERVEQL 690
>TC17802 similar to UP|Q9XII2 (Q9XII2) At2g16050 protein, partial (72%)
Length = 571
Score = 38.5 bits (88), Expect = 0.002
Identities = 20/63 (31%), Positives = 32/63 (50%), Gaps = 6/63 (9%)
Frame = +2
Query: 326 ALRKEGEALPQQVEDVKH----EHVLKLDMAK-AYVCDSCKEQGKFWAFSCD-VCDYDLH 379
+ RKE + ++ H +H L+ + ++ + CD CKE G + C +CDYDLH
Sbjct: 161 SFRKETSITNTKCSEISHFSHPKHKLRFEYSEFPFKCDGCKEVGIGSRYKCSSICDYDLH 340
Query: 380 PSC 382
C
Sbjct: 341 MHC 349
Score = 30.0 bits (66), Expect = 0.69
Identities = 9/26 (34%), Positives = 16/26 (60%)
Frame = +2
Query: 357 CDSCKEQGKFWAFSCDVCDYDLHPSC 382
C++C++ + + C C +DLHP C
Sbjct: 440 CNACEKGVTGFVYHCMTCGFDLHPCC 517
>TC13764 similar to UP|Q9C9Y6 (Q9C9Y6) Thioredoxin-like protein, partial
(73%)
Length = 585
Score = 36.6 bits (83), Expect = 0.007
Identities = 15/36 (41%), Positives = 21/36 (57%)
Frame = +3
Query: 34 KVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYE 69
K+ ++ DGK++ FSA WC PC+ P EL E
Sbjct: 6 KLEEAKRDGKIVIANFSATWCGPCKIIAPYYCELSE 113
>TC17161 homologue to UP|THIF_PEA (P29450) Thioredoxin F-type, chloroplast
precursor (TRX-F), partial (72%)
Length = 850
Score = 34.7 bits (78), Expect = 0.028
Identities = 15/48 (31%), Positives = 23/48 (47%)
Frame = +2
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREED 90
K + L WC PC+ P+ EL E NL+++F+ D +D
Sbjct: 311 KTVVLDMFTQWCGPCKVIAPKYQELAEK------NLDVVFLKLDCNQD 436
>TC14960 similar to UP|Q8GUR9 (Q8GUR9) Thioredoxin h, partial (98%)
Length = 688
Score = 33.9 bits (76), Expect = 0.048
Identities = 16/47 (34%), Positives = 26/47 (55%)
Frame = +1
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
K+I + F+A+WC PCR P L E+ + L ++F+ D +E
Sbjct: 163 KLIVVDFTASWCGPCRFIAPILAEIAKKLP------HVVFLKVDVDE 285
>TC8727
Length = 1382
Score = 33.5 bits (75), Expect = 0.063
Identities = 17/52 (32%), Positives = 27/52 (51%)
Frame = +1
Query: 279 YDIKGIPALVLIGPDGKVISLNGKFMVTSYGADAFPFTESRIRDLEAALRKE 330
+ K P +V++ P GKV N M+ +G FPFT+ D+E + K+
Sbjct: 337 WQFKKQPMVVVLSPQGKVQHTNAFHMIQVWGIKGFPFTQ----DIEVNIGKQ 480
>BP057442
Length = 429
Score = 33.1 bits (74), Expect = 0.082
Identities = 12/15 (80%), Positives = 14/15 (93%)
Frame = -1
Query: 378 LHPSCLEKVNKDQDW 392
LHPSCLEKV++D DW
Sbjct: 429 LHPSCLEKVHQDPDW 385
>BP076542
Length = 376
Score = 33.1 bits (74), Expect = 0.082
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = -1
Query: 367 WAFSCDVCDYDLHPSC 382
W++ C+ CD+DLHP C
Sbjct: 373 WSYYCEECDFDLHPKC 326
>BP043095
Length = 396
Score = 33.1 bits (74), Expect = 0.082
Identities = 19/47 (40%), Positives = 27/47 (57%)
Frame = +1
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
K+I + F+A+WC PCR P L E L K+ N +IF+ D +E
Sbjct: 145 KLIVVDFTASWCGPCRFIAPYLGE----LAKKYTN--VIFLKVDVDE 267
>TC17672 similar to UP|Q93X83 (Q93X83) Hsp70 interacting protein/thioredoxin
chimera, partial (33%)
Length = 709
Score = 32.7 bits (73), Expect = 0.11
Identities = 17/58 (29%), Positives = 27/58 (46%)
Frame = +1
Query: 32 EGKVPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREE 89
E K+ + ++ L+F+A WC PCR P L E K ++F+ D +E
Sbjct: 121 ETKLGAASKTSRLAILYFTATWCGPCRFISPIYTSLAEKYPK------VVFLKVDIDE 276
>TC18055 similar to UP|CAE46765 (CAE46765) NADPH thioredoxin reductase,
partial (20%)
Length = 748
Score = 29.3 bits (64), Expect = 1.2
Identities = 14/48 (29%), Positives = 25/48 (51%)
Frame = +1
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKRGINLEIIFVSFDREED 90
++IC+ +++ C PCR P L ++ + + + FV D EED
Sbjct: 253 RLICVLYTSPTCGPCRTLKPILSKVIDE-----FDQNVHFVEIDIEED 381
>TC16557
Length = 577
Score = 28.1 bits (61), Expect = 2.6
Identities = 18/51 (35%), Positives = 21/51 (40%)
Frame = +2
Query: 8 AKHIDSHDVLKILAAEGAEYLLSCEGKVPLSECDGKVICLFFSANWCRPCR 58
A H DSHD EG L SC + L E +G L +C CR
Sbjct: 122 ATHGDSHD-------EGLSSLCSCSLRRNLKELEGIFCLLHVFEGYCLECR 253
>TC12168 similar to UP|Q8L7S9 (Q8L7S9) At1g43560/T10P12_4, partial (50%)
Length = 408
Score = 28.1 bits (61), Expect = 2.6
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +1
Query: 43 KVICLFFSANWCRPCRAFVPRLVELYETLRKR 74
K + + F A WC PC+ VP L E+ L+ +
Sbjct: 262 KPVFVDFYATWCGPCQFMVPILDEVSTRLKDK 357
>AV426732
Length = 238
Score = 27.7 bits (60), Expect = 3.4
Identities = 10/26 (38%), Positives = 16/26 (61%)
Frame = +3
Query: 44 VICLFFSANWCRPCRAFVPRLVELYE 69
++ + FSA+WC PCR P + + E
Sbjct: 132 LLVIDFSASWCGPCRFIEPAIHAMAE 209
>AV765055
Length = 516
Score = 27.7 bits (60), Expect = 3.4
Identities = 10/32 (31%), Positives = 18/32 (56%)
Frame = -2
Query: 228 YNNLKATKGNCFEIVLISTDRDHKEFNVNRSS 259
Y N+K +G C I ++ R+H F + ++S
Sbjct: 464 YKNIKRPRGGCLNI*ILFPKREHSPFFLEKNS 369
>TC11101
Length = 416
Score = 27.3 bits (59), Expect = 4.5
Identities = 13/34 (38%), Positives = 17/34 (49%)
Frame = +2
Query: 45 ICLFFSANWCRPCRAFVPRLVELYETLRKRGINL 78
+CL WC R F P + TLRKR ++L
Sbjct: 26 LCLILKM-WCMSLRNFTPSTLPKRSTLRKRRVHL 124
>TC8228 similar to UP|GPX4_CITSI (Q06652) Probable phospholipid
hydroperoxide glutathione peroxidase (PHGPx)
(Salt-associated protein) , partial (95%)
Length = 883
Score = 26.9 bits (58), Expect = 5.9
Identities = 18/64 (28%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Frame = +2
Query: 35 VPLSECDGKVICLFFSANWCRPCRAFVPRLVELYETLRKRGIN-LEIIFVSFDREEDGFN 93
V L + GKV+ + A+ C + L +LYE + +G+ L F +E G N
Sbjct: 233 VNLGDYKGKVLLIVNVASQCGLTNSNYTELSQLYEKYKSKGLEILGFPCNQFGAQEPGDN 412
Query: 94 EHLK 97
E ++
Sbjct: 413 EQIQ 424
>TC8931 similar to GB|AAL09731.1|15982767|AY057490 At2g32240/F22D22.1
{Arabidopsis thaliana;} , partial (4%)
Length = 876
Score = 26.6 bits (57), Expect = 7.7
Identities = 26/91 (28%), Positives = 38/91 (41%), Gaps = 11/91 (12%)
Frame = -2
Query: 186 FLISGDDRKVPVS-ELAGKTIGLYFCAYWSPPSRAFTAQLTDVYNNLKATKGNCF----- 239
FL GD+ P+S +L ++WS P+ T L+ V N + CF
Sbjct: 344 FLFDGDEMLDPMSLDLTSNP------SFWSTPTSPLTDNLSLVSCNFCSKSVTCFCKRVI 183
Query: 240 EIVLISTD-----RDHKEFNVNRSSTPWLAI 265
LIS+ +D N S+T WLA+
Sbjct: 182 SFFLISSSSREFFKDASSSNF*DSATFWLAM 90
>TC11352
Length = 738
Score = 26.2 bits (56), Expect = 10.0
Identities = 13/23 (56%), Positives = 16/23 (69%), Gaps = 2/23 (8%)
Frame = -1
Query: 54 CRPCRAF--VPRLVELYETLRKR 74
C+P R F P +V LY+TLRKR
Sbjct: 159 CKPQRKFDQFPPIVILYQTLRKR 91
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.140 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,255,998
Number of Sequences: 28460
Number of extensions: 102243
Number of successful extensions: 543
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 534
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 538
length of query: 392
length of database: 4,897,600
effective HSP length: 92
effective length of query: 300
effective length of database: 2,279,280
effective search space: 683784000
effective search space used: 683784000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0372a.5


