

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0370.4
(170 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
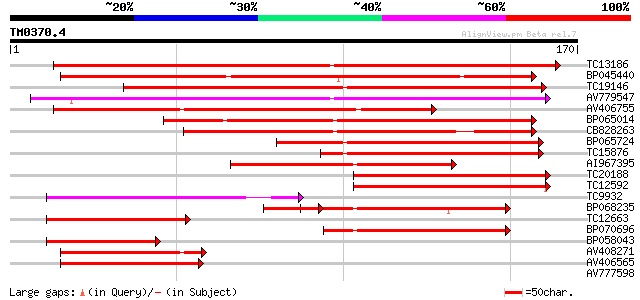
Sequences producing significant alignments: (bits) Value
TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidat... 139 3e-34
BP045440 134 1e-32
TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-contain... 129 3e-31
AV779547 125 3e-30
AV406755 120 1e-28
BP065014 116 2e-27
CB828263 104 9e-24
BP065724 100 1e-22
TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 78 9e-16
AI967395 71 1e-13
TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-... 63 3e-11
TC12592 62 5e-11
TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like... 60 2e-10
BP068235 50 5e-10
TC12663 weakly similar to UP|Q84ZV8 (Q84ZV8) R 3 protein, partia... 49 3e-07
BP070696 49 3e-07
BP058043 43 2e-05
AV408271 42 6e-05
AV406565 40 2e-04
AV777598 30 0.29
>TC13186 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate
resistance protein KR1, partial (6%)
Length = 586
Score = 139 bits (349), Expect = 3e-34
Identities = 71/152 (46%), Positives = 101/152 (65%)
Frame = +1
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
F YDVFIS DTR F L+ L KGI TF+DD + + +++S SKAIE S +
Sbjct: 97 FIYDVFISLITSDTRFGFSGDLYTALSDKGILTFIDDGLPRPDDIMSTVNSKAIESSRIA 276
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHE 133
I+V+S+NYASS++CLDEL+ IL+ R L++P+FY VEPS VR+QKG +G+A A+HE
Sbjct: 277 IVVISENYASSSFCLDELMYILK-RREEKGLLIWPVFYQVEPSDVRYQKGGFGKAFASHE 453
Query: 134 KKYGINSDRIRTWRSTLNEVCQLVGHHISTTG 165
+++G +S ++R WR+ L +V L H G
Sbjct: 454 ERFGSDSAKVRKWRNALKQVADLKAFHFFKHG 549
>BP045440
Length = 535
Score = 134 bits (336), Expect = 1e-32
Identities = 75/144 (52%), Positives = 90/144 (62%), Gaps = 1/144 (0%)
Frame = -3
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLII 75
Y +FISFRG DTR F K L++ L R+G TF DDE L+ G+ I L IEES II
Sbjct: 527 YQIFISFRGMDTRDDFTKPLYQGLCREGFVTFKDDESLEGGSPIEKLLDD-IEESRFAII 351
Query: 76 VLSKNYASSTWCLDELVKILEC-SRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEK 134
+LSKNYA S WCL EL KILEC + G QLV P V P+ VRH +Y +AMA HEK
Sbjct: 350 ILSKNYAESEWCLKELSKILECKDQEGKNQLVLPSSTEVTPTEVRHVINRYADAMAKHEK 171
Query: 135 KYGINSDRIRTWRSTLNEVCQLVG 158
GI+S I+ W+ L +VC L G
Sbjct: 170 N-GIDSKTIQAWKKALFDVCTLTG 102
>TC19146 similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing protein
TSDC, partial (12%)
Length = 701
Score = 129 bits (324), Expect = 3e-31
Identities = 64/127 (50%), Positives = 87/127 (68%)
Frame = +1
Query: 35 LHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVLIIVLSKNYASSTWCLDELVKI 94
L+ L+R+G +TF+DDE L GN I+ +L A E+S + IIV S+NY S+WCLDELVKI
Sbjct: 1 LYHALRREGFKTFMDDEGLVGGNEIAKSLLDASEKSRLSIIVFSENYGYSSWCLDELVKI 180
Query: 95 LECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGINSDRIRTWRSTLNEVC 154
+EC +QLV+PIFY VE S V Q+ Y AM HEK YG +SD ++ WRS L+ +
Sbjct: 181 VECMET-KKQLVWPIFYKVEKSDVTSQENSYEHAMIGHEKNYGKDSDEVKKWRSALSAMA 357
Query: 155 QLVGHHI 161
+L GH++
Sbjct: 358 KLEGHNV 378
>AV779547
Length = 556
Score = 125 bits (315), Expect = 3e-30
Identities = 72/161 (44%), Positives = 92/161 (56%), Gaps = 5/161 (3%)
Frame = +1
Query: 7 DGEELHIFNYD-----VFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISP 61
+G +FN D +SFRGEDTR+ F L L KG TF DD ML+ G IS
Sbjct: 55 NGNPKQLFNSDDMEI*CVVSFRGEDTRNNFTDHLLGALYGKGFVTFKDDTMLRKGKNIST 234
Query: 62 ALSKAIEESMVLIIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQ 121
L +AIE S +LI+V SK YASSTWCL EL KI + G +Q V P+ V PS VR Q
Sbjct: 235 ELIQAIEGSQILIVVFSKYYASSTWCLQELAKIAD-GIVGKRQTVLPVVCDVTPSEVRKQ 411
Query: 122 KGKYGEAMAAHEKKYGINSDRIRTWRSTLNEVCQLVGHHIS 162
G YGEA HE+++ + + ++ WR L EV L G ++
Sbjct: 412 SGNYGEAFLKHEERFKEDLEMVQRWRKALAEVADLSGWDVT 534
>AV406755
Length = 425
Score = 120 bits (301), Expect = 1e-28
Identities = 60/115 (52%), Positives = 82/115 (71%)
Frame = +3
Query: 14 FNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESMVL 73
++YDVF+SFRGE++R +F L+ LK GI+ F+D+E L+ G IS +L KAIE+S +
Sbjct: 78 WSYDVFLSFRGEESRRSFTSHLYTALKNAGIKVFMDNE-LQRGEDISSSLLKAIEDSRIA 254
Query: 74 IIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEA 128
II+ S NY S WCLDEL KI+EC R Q+ V P+FY+V+PS +R Q+G GEA
Sbjct: 255 IIIFSTNYTGSKWCLDELEKIIECQRTIGQE-VMPVFYNVDPSDIRKQRGTVGEA 416
>BP065014
Length = 498
Score = 116 bits (291), Expect = 2e-27
Identities = 61/112 (54%), Positives = 77/112 (68%)
Frame = -2
Query: 47 FVDDEMLKAGNVISPALSKAIEESMVLIIVLSKNYASSTWCLDELVKILECSRRGNQQLV 106
F DD L+ G I L AIEES V I+VLS+N+A S WCL ELVKIL+C R+ + LV
Sbjct: 497 FKDDLSLEDGAPIEELLG-AIEESRVAIVVLSENFADSKWCLKELVKILDC-RKKKKPLV 324
Query: 107 YPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGINSDRIRTWRSTLNEVCQLVG 158
PIFY VEP+++RH KG+YGEAMA HEKK G +S +R W L+++ L G
Sbjct: 323 LPIFYKVEPTNIRHLKGRYGEAMAEHEKKLGKDSQTVREWTQALSDISYLKG 168
>CB828263
Length = 570
Score = 104 bits (259), Expect = 9e-24
Identities = 52/106 (49%), Positives = 70/106 (65%)
Frame = +2
Query: 53 LKAGNVISPALSKAIEESMVLIIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYH 112
L+ G+ IS +L +AIEE+ + +IV SKNY +S WCLDELVKILEC +R Q+V P+FY
Sbjct: 8 LQRGDEISSSLLRAIEEAKLSVIVFSKNYGNSKWCLDELVKILEC-KRTRGQIVLPVFYD 184
Query: 113 VEPSHVRHQKGKYGEAMAAHEKKYGINSDRIRTWRSTLNEVCQLVG 158
++PSHVR+Q G Y EA H + D+++ WR L E L G
Sbjct: 185 IDPSHVRNQTGTYAEAFVKHGQ-----VDKVQKWREALREAANLSG 307
>BP065724
Length = 495
Score = 100 bits (249), Expect = 1e-22
Identities = 44/80 (55%), Positives = 61/80 (76%)
Frame = -1
Query: 81 YASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGINS 140
YA S+WCL+E+VKILEC ++ N QLV+PIFY VEPS VRHQ+ Y +AM E +YG +S
Sbjct: 495 YADSSWCLEEVVKILECKQK-NDQLVWPIFYKVEPSDVRHQRNSYEKAMIKQETRYGKDS 319
Query: 141 DRIRTWRSTLNEVCQLVGHH 160
++++ WRS+L++VC L H
Sbjct: 318 EKVKKWRSSLSQVCDLKAFH 259
>TC15876 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (13%)
Length = 558
Score = 77.8 bits (190), Expect = 9e-16
Identities = 35/67 (52%), Positives = 49/67 (72%)
Frame = +1
Query: 94 ILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGINSDRIRTWRSTLNEV 153
ILEC ++ N QLV+PIFY VEPS VRHQ+ Y +AM E +YG +S++++ WRS+L++V
Sbjct: 16 ILECXQK-NDQLVWPIFYKVEPSDVRHQRNSYEKAMIKPETRYGKDSEKVKKWRSSLSQV 192
Query: 154 CQLVGHH 160
C L H
Sbjct: 193 CDLKAFH 213
>AI967395
Length = 205
Score = 70.9 bits (172), Expect = 1e-13
Identities = 34/68 (50%), Positives = 50/68 (73%)
Frame = +3
Query: 67 IEESMVLIIVLSKNYASSTWCLDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYG 126
I+ES ++I+V S++YA S CL+ELV I++ NQ LV+P++Y VE +RHQ+G+YG
Sbjct: 3 IQESRLIIVVFSEHYAVSASCLNELVNIVDVMIMNNQ-LVWPVYYGVEAPEIRHQRGRYG 179
Query: 127 EAMAAHEK 134
EAMA E+
Sbjct: 180 EAMAKFEE 203
>TC20188 weakly similar to UP|Q8S4X3 (Q8S4X3) TIR-similar-domain-containing
protein TSDC, partial (11%)
Length = 494
Score = 62.8 bits (151), Expect = 3e-11
Identities = 27/59 (45%), Positives = 40/59 (67%)
Frame = +1
Query: 104 QLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGINSDRIRTWRSTLNEVCQLVGHHIS 162
QLV+PIFY + S V +Q YG AM AHE +G +S++++ W+S L+E+ L G HI+
Sbjct: 4 QLVWPIFYEISXSDVSNQTKSYGHAMLAHENSFGKDSEKVQKWKSALSEMAFLQGEHIT 180
>TC12592
Length = 534
Score = 62.0 bits (149), Expect = 5e-11
Identities = 29/61 (47%), Positives = 41/61 (66%), Gaps = 2/61 (3%)
Frame = +3
Query: 104 QLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGINSDRIRTWRSTLNEVCQLVGH--HI 161
QLV P+FY VEP+ +R+ KGK+GEAMA E K+G +S+ I W+ L ++ L G H+
Sbjct: 3 QLVLPVFYGVEPTDIRNLKGKFGEAMAEIENKFGKDSNTIHEWKQALRDISYLKGEPCHV 182
Query: 162 S 162
S
Sbjct: 183 S 185
>TC9932 weakly similar to UP|Q93YA7 (Q93YA7) Resistance gene-like, partial
(5%)
Length = 689
Score = 60.5 bits (145), Expect = 2e-10
Identities = 32/77 (41%), Positives = 46/77 (59%)
Frame = -2
Query: 12 HIFNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISPALSKAIEESM 71
H + YDVFI+FRG+DTRH+FV L+ L G+ F+DDE L+ G + P L K I ++
Sbjct: 214 HQWIYDVFINFRGKDTRHSFVSHLYTALSNAGVNVFLDDENLERGLELEPELQKLINLNL 35
Query: 72 VLIIVLSKNYASSTWCL 88
+++ SST L
Sbjct: 34 -------QHFMSSTQAL 5
>BP068235
Length = 511
Score = 50.1 bits (118), Expect(2) = 5e-10
Identities = 26/64 (40%), Positives = 40/64 (61%), Gaps = 1/64 (1%)
Frame = -1
Query: 88 LDELVKILECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMA-AHEKKYGINSDRIRTW 146
L L + LEC + NQ L++P Y VEPS +R++ YG A+A +G +S++I+TW
Sbjct: 475 LMNLSRFLECKKMKNQ-LLWPCPYKVEPSDIRYRPSSYGTALA*TRTDAFGNDSEKIQTW 299
Query: 147 RSTL 150
+S L
Sbjct: 298 KSAL 287
Score = 28.5 bits (62), Expect(2) = 5e-10
Identities = 13/18 (72%), Positives = 15/18 (83%)
Frame = -3
Query: 77 LSKNYASSTWCLDELVKI 94
LS+NYA S CLDELV+I
Sbjct: 509 LSENYAYSARCLDELVQI 456
>TC12663 weakly similar to UP|Q84ZV8 (Q84ZV8) R 3 protein, partial (4%)
Length = 494
Score = 49.3 bits (116), Expect = 3e-07
Identities = 21/43 (48%), Positives = 30/43 (68%)
Frame = +3
Query: 12 HIFNYDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLK 54
H+ ++VF++FRG DT H F+ L K L KGI TF++DE L+
Sbjct: 15 HVLTFNVFLNFRGSDTCHGFISFLCKSLLDKGIYTFINDEELQ 143
>BP070696
Length = 429
Score = 49.3 bits (116), Expect = 3e-07
Identities = 22/56 (39%), Positives = 35/56 (62%)
Frame = -1
Query: 95 LECSRRGNQQLVYPIFYHVEPSHVRHQKGKYGEAMAAHEKKYGINSDRIRTWRSTL 150
LEC ++ N ++ PIF ++E + +Q Y + M AHE+K G SDR++ W+S L
Sbjct: 429 LECKKKKNN-IIRPIFQNMELLKIIYQINSYDDVMVAHERKNGKESDRVKKWKSAL 265
>BP058043
Length = 375
Score = 43.1 bits (100), Expect = 2e-05
Identities = 20/34 (58%), Positives = 24/34 (69%)
Frame = -3
Query: 12 HIFNYDVFISFRGEDTRHTFVKILHKELKRKGIR 45
H F YDVFIS RGE R TF++ KEL +KGI+
Sbjct: 166 HNFTYDVFISLRGEVLRDTFIECFLKELMQKGIK 65
Score = 32.7 bits (73), Expect = 0.034
Identities = 15/35 (42%), Positives = 25/35 (70%)
Frame = -2
Query: 27 TRHTFVKILHKELKRKGIRTFVDDEMLKAGNVISP 61
TRH + ++ +G +TF+DDE+++AG+VISP
Sbjct: 119 TRHLY-RMFS*RADAEGDQTFIDDEIMEAGDVISP 18
>AV408271
Length = 409
Score = 42.0 bits (97), Expect = 6e-05
Identities = 22/44 (50%), Positives = 28/44 (63%)
Frame = +1
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNVI 59
YDVF+SFRGED R F+ L ++K I F+DD+ LK G I
Sbjct: 280 YDVFVSFRGEDIRDGFLSHLADAFRQKKINAFMDDK-LKRGQEI 408
>AV406565
Length = 207
Score = 40.4 bits (93), Expect = 2e-04
Identities = 17/43 (39%), Positives = 28/43 (64%)
Frame = +2
Query: 16 YDVFISFRGEDTRHTFVKILHKELKRKGIRTFVDDEMLKAGNV 58
+DVF+SFRGEDTR+T L L R ++T++D ++ + +
Sbjct: 77 HDVFLSFRGEDTRYTCTGHLQATLTRLQVKTYIDYDLQRGDEI 205
>AV777598
Length = 371
Score = 29.6 bits (65), Expect = 0.29
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = +3
Query: 14 FNYDVFISFRGEDTRHTFVKILH 36
F YDVF+SF G DTR +F L+
Sbjct: 303 FTYDVFLSFCGVDTRFSFTGNLY 371
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.326 0.140 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,952,079
Number of Sequences: 28460
Number of extensions: 37570
Number of successful extensions: 240
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 228
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 228
length of query: 170
length of database: 4,897,600
effective HSP length: 84
effective length of query: 86
effective length of database: 2,506,960
effective search space: 215598560
effective search space used: 215598560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0370.4


