

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369c.3
(975 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
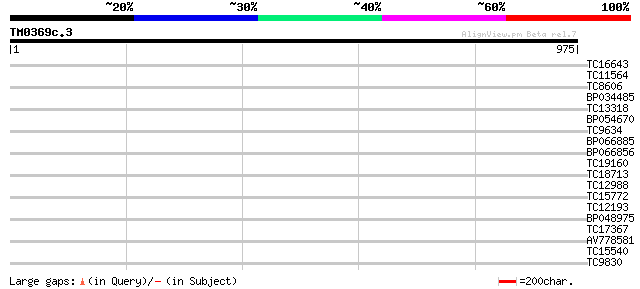
Score E
Sequences producing significant alignments: (bits) Value
TC16643 similar to UP|Q9FIH3 (Q9FIH3) Serine/threonine protein k... 33 0.22
TC11564 homologue to GB|AAC82338.1|3941324|AF042799 suppressor o... 32 0.49
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 31 0.83
BP034485 30 1.4
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 30 1.8
BP054670 30 1.8
TC9634 similar to UP|Q8W0Z3 (Q8W0Z3) At1g29350/F15D2_27, partial... 30 2.4
BP066885 29 3.1
BP066856 29 3.1
TC19160 similar to UP|Q9LPZ8 (Q9LPZ8) T23J18.3, partial (15%) 29 4.1
TC18713 similar to GB|AAL90989.1|19699246|AY090328 AT3g03000/F13... 29 4.1
TC12988 similar to UP|CYP1_BRUMA (Q27450) Peptidylprolyl isomera... 29 4.1
TC15772 UP|Q40218 (Q40218) RAB8D, complete 28 5.4
TC12193 28 7.0
BP048975 28 7.0
TC17367 similar to GB|BAB03049.1|9294683|AP001305 syringomycin b... 28 7.0
AV778581 28 9.2
TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fra... 28 9.2
TC9830 28 9.2
>TC16643 similar to UP|Q9FIH3 (Q9FIH3) Serine/threonine protein kinase-like
protein, partial (28%)
Length = 585
Score = 33.1 bits (74), Expect = 0.22
Identities = 25/68 (36%), Positives = 36/68 (52%), Gaps = 5/68 (7%)
Frame = +2
Query: 44 APFAKSQTMPPLPSRSHSSLSSR-----PTSSANNSSPGRGGGGNVSRNGSAMSLMSGLG 98
AP + S++ PP PS+ +S+SSR SSA +S G G +RNG + + S G
Sbjct: 344 APSSPSRSSPPTPSK--ASVSSRRRWKLSASSATATS*RSSGTGPPARNGYSSTSSSRKG 517
Query: 99 SYASLFPG 106
+ S F G
Sbjct: 518 TLTSGFTG 541
>TC11564 homologue to GB|AAC82338.1|3941324|AF042799 suppressor of white
apricot homolog 2 {Mus musculus;} , partial (3%)
Length = 428
Score = 32.0 bits (71), Expect = 0.49
Identities = 23/60 (38%), Positives = 35/60 (58%), Gaps = 5/60 (8%)
Frame = +2
Query: 269 SANKEDLISVKEFIYRQSDILR-----GRGGIVNSNSGSAAGVGMVAVAAAAAAASAASG 323
S +E L+SV++ + +QS+ GR G S G+ VG + AAAAAAAS+++G
Sbjct: 173 STMEEPLLSVEQSVEKQSEKWSSYQHMGRTGSA-SLGGTEVSVGEIRSAAAAAAASSSAG 349
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 31.2 bits (69), Expect = 0.83
Identities = 34/112 (30%), Positives = 51/112 (45%), Gaps = 8/112 (7%)
Frame = +1
Query: 57 SRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMS-GLGSYASLFPGQCIPV---- 111
S S SSLSS T S+++SSP + S + S+ S S S +S F P
Sbjct: 292 SSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSPFSSGPAPAPGPP 471
Query: 112 --TLFVFVDDFSNLS-NSSTNGEESSDVSSLSQSNSMSSAAKANLPAKVSGS 160
F + F L + ++ ++SS S S S SS+ ++LP+ S S
Sbjct: 472 SGAFFELLLLFELLPYPAPSSDDKSSSPSPNGPSPSSSSSPSSSLPSSPSAS 627
>BP034485
Length = 485
Score = 30.4 bits (67), Expect = 1.4
Identities = 15/34 (44%), Positives = 18/34 (52%)
Frame = +3
Query: 72 NNSSPGRGGGGNVSRNGSAMSLMSGLGSYASLFP 105
N G G GGN++ NG M MS LGS + P
Sbjct: 360 NGHDHGHGIGGNMNGNGGHMDPMSQLGSMGQMGP 461
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 30.0 bits (66), Expect = 1.8
Identities = 19/53 (35%), Positives = 27/53 (50%)
Frame = -3
Query: 43 MAPFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMS 95
+AP S + LPS S SS SS +SS ++SS + S + S+ S S
Sbjct: 415 LAPSFPSTSTSSLPSSSSSSSSSSSSSSPSSSSSSSSSSSSSSSSSSSSSSSS 257
>BP054670
Length = 367
Score = 30.0 bits (66), Expect = 1.8
Identities = 21/62 (33%), Positives = 30/62 (47%), Gaps = 1/62 (1%)
Frame = +2
Query: 42 AMAPFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPG-RGGGGNVSRNGSAMSLMSGLGSY 100
++ P P PS S SSL TS + + +PG + G + S S+ SL S GS
Sbjct: 182 SVPPGKSMMKSPSSPSSSSSSLERSSTSLSLSPNPGPKRGSTSESSESSSSSLSSSSGSE 361
Query: 101 AS 102
+S
Sbjct: 362 SS 367
>TC9634 similar to UP|Q8W0Z3 (Q8W0Z3) At1g29350/F15D2_27, partial (7%)
Length = 687
Score = 29.6 bits (65), Expect = 2.4
Identities = 18/58 (31%), Positives = 25/58 (43%)
Frame = -3
Query: 45 PFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSGLGSYAS 102
PF + ++ S + SRP ANN+S RGGGG V + G + S
Sbjct: 340 PFHEVKSKREKKKESKDTTDSRPRG-ANNNSSSRGGGGGVRGGADRYAGRGGAAQFNS 170
>BP066885
Length = 469
Score = 29.3 bits (64), Expect = 3.1
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 1/50 (2%)
Frame = -1
Query: 99 SYASLFPGQCIPVTL-FVFVDDFSNLSNSSTNGEESSDVSSLSQSNSMSS 147
S A FPG C+P +L +++ DF L N + + SL NS S
Sbjct: 355 SSALSFPGDCLPASLKSLYIRDFRELEFPKQNQQXHELLESLDIDNSCDS 206
>BP066856
Length = 485
Score = 29.3 bits (64), Expect = 3.1
Identities = 19/51 (37%), Positives = 25/51 (48%), Gaps = 2/51 (3%)
Frame = +2
Query: 51 TMPPLPSRSHSSLSSR--PTSSANNSSPGRGGGGNVSRNGSAMSLMSGLGS 99
T PP SRS SS S+ T SA+ +S R + S G ++ G GS
Sbjct: 65 TGPPTTSRSMSSASTGTISTGSASTTSASRSSSSSDSSKGYSLGTNEGTGS 217
>TC19160 similar to UP|Q9LPZ8 (Q9LPZ8) T23J18.3, partial (15%)
Length = 557
Score = 28.9 bits (63), Expect = 4.1
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = -1
Query: 294 GIVNSNSGSAAGVGMVAVAAAAAAASAASGKTFTTPDLPNFEIWS 338
GI +S SGSA + V V + +++S + +P P+ +WS
Sbjct: 404 GICSSTSGSARSLAFVEVKLSKSSSSFRRDSSEESPPSPSSALWS 270
>TC18713 similar to GB|AAL90989.1|19699246|AY090328 AT3g03000/F13E7_5
{Arabidopsis thaliana;}, partial (88%)
Length = 868
Score = 28.9 bits (63), Expect = 4.1
Identities = 16/46 (34%), Positives = 21/46 (44%)
Frame = -3
Query: 42 AMAPFAKSQTMPPLPSRSHSSLSSRPTSSANNSSPGRGGGGNVSRN 87
AM + + M P PSRS++ S R SS GR G + N
Sbjct: 521 AME*ASSAAVMNPFPSRSNTLKSCRSCSSVYGDLAGRSSGATTATN 384
>TC12988 similar to UP|CYP1_BRUMA (Q27450) Peptidylprolyl isomerase 1
(Peptidylprolyl cis-trans isomerase 1) (PPIase 1)
(Cyclophilin) (BmCYP-1) , partial (4%)
Length = 453
Score = 28.9 bits (63), Expect = 4.1
Identities = 32/90 (35%), Positives = 42/90 (46%)
Frame = -1
Query: 58 RSHSSLSSRPTSSANNSSPGRGGGGNVSRNGSAMSLMSGLGSYASLFPGQCIPVTLFVFV 117
+S SS SS S ++NSS S GSA L + ++ S F + + L
Sbjct: 264 QSSSSESSFAPS*SSNSS---------SYKGSATCLET---AWCSRFSSDDLLLEL---- 133
Query: 118 DDFSNLSNSSTNGEESSDVSSLSQSNSMSS 147
+LS SS + ESS SS S SNS SS
Sbjct: 132 ----SLSGSSNSNSESSTASSSSSSNSSSS 55
>TC15772 UP|Q40218 (Q40218) RAB8D, complete
Length = 1139
Score = 28.5 bits (62), Expect = 5.4
Identities = 27/102 (26%), Positives = 44/102 (42%), Gaps = 16/102 (15%)
Frame = +2
Query: 685 VVAGSAAVDSGFPPLQQRKLPASGSEKGMKL---SKPSNQNGERVNAAVDYEISQKSQD- 740
+V A +D + K A E G+K S +N N E V ++ +I Q+ D
Sbjct: 611 LVGNKADMDESKRAVPTSKGQALADEYGIKFFETSAKTNLNVEEVFFSIARDIKQRLADT 790
Query: 741 --------ILLTEGSLDGSGN----NSCRDDDPFLRIGSNVV 770
+ + + S G+G +SC D L+IGSN++
Sbjct: 791 DHKAEPTTLKINQDSAAGAGEAANKSSCCG*DHILQIGSNII 916
>TC12193
Length = 395
Score = 28.1 bits (61), Expect = 7.0
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = +2
Query: 581 SKGLLQSGFCATEKNLLKWTVYL 603
+KGL+QSGFC LL TV+L
Sbjct: 251 NKGLIQSGFCTDWSILLTLTVWL 319
>BP048975
Length = 530
Score = 28.1 bits (61), Expect = 7.0
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = +3
Query: 693 DSGFPPLQQRKLPASGSEKGMKLSKPSNQNGERVN 727
+S P LQ P+S +KGM +SKP+ + +R N
Sbjct: 264 ESNLPHLQHS**PSSS*KKGMTISKPTPKKPKREN 368
>TC17367 similar to GB|BAB03049.1|9294683|AP001305 syringomycin biosynthesis
enzyme-like protein {Arabidopsis thaliana;}, partial
(17%)
Length = 341
Score = 28.1 bits (61), Expect = 7.0
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +3
Query: 45 PFAKSQTMPPLPSRSHSSLSSRPTSSANNSSP 76
P + + P PS S +S S+RPT+S +S P
Sbjct: 195 PTSNHSSTNPAPSFSETSHSTRPTTSTTSSRP 290
>AV778581
Length = 525
Score = 27.7 bits (60), Expect = 9.2
Identities = 12/28 (42%), Positives = 17/28 (59%)
Frame = -2
Query: 173 GGFRKKLQASLEAQIRFLIKKCHTLSGS 200
GG + +Q SL +R +KKCH L+ S
Sbjct: 278 GGVMEMIQESLPESLRKSVKKCHQLAKS 195
>TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fragment),
partial (6%)
Length = 806
Score = 27.7 bits (60), Expect = 9.2
Identities = 13/26 (50%), Positives = 17/26 (65%)
Frame = -1
Query: 294 GIVNSNSGSAAGVGMVAVAAAAAAAS 319
G+V + SG AA VG+V VAAA +
Sbjct: 140 GVVAAASGKAAAVGLVEVAAAVVVGA 63
>TC9830
Length = 574
Score = 27.7 bits (60), Expect = 9.2
Identities = 12/30 (40%), Positives = 20/30 (66%)
Frame = +2
Query: 60 HSSLSSRPTSSANNSSPGRGGGGNVSRNGS 89
H +++ R +SS+ ++ G GGGG + NGS
Sbjct: 485 HKAVARRSSSSSAAAAEGDGGGGWLRSNGS 574
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.130 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,780,739
Number of Sequences: 28460
Number of extensions: 246832
Number of successful extensions: 1344
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 1312
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1340
length of query: 975
length of database: 4,897,600
effective HSP length: 99
effective length of query: 876
effective length of database: 2,080,060
effective search space: 1822132560
effective search space used: 1822132560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0369c.3


