

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369c.1
(573 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
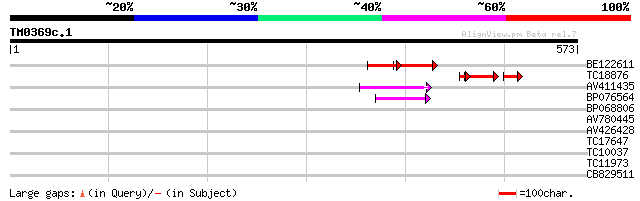
Sequences producing significant alignments: (bits) Value
BE122611 53 6e-16
TC18876 51 3e-09
AV411435 47 6e-06
BP076564 45 4e-05
BP068806 36 0.015
AV780445 32 0.21
AV426428 32 0.36
TC17647 weakly similar to UP|AAR06918 (AAR06918) UDP-glycosyltra... 29 1.8
TC10037 28 5.3
TC11973 weakly similar to UP|AAR34269 (AAR34269) Inositol-1-mono... 27 6.9
CB829511 27 6.9
>BE122611
Length = 357
Score = 52.8 bits (125), Expect(2) = 6e-16
Identities = 22/35 (62%), Positives = 28/35 (79%)
Frame = -3
Query: 362 LIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLV 396
L+WKA A EKV FFLWL GR+ALP+N +R+H L+
Sbjct: 229 LVWKANAPEKVRFFLWLVGRNALPSNARRHHITLL 125
Score = 48.1 bits (113), Expect(2) = 6e-16
Identities = 19/44 (43%), Positives = 27/44 (61%)
Frame = -2
Query: 389 KRYHSHLVKFDACFRCGASVEDVEHALRRCPGSVWVRSRFAKAL 432
K+ HL AC RCGA ED H R CPGS W+ ++F++++
Sbjct: 149 KKAPYHLASSAACIRCGAPSEDANHVFRWCPGSAWMWTQFSRSI 18
>TC18876
Length = 555
Score = 50.8 bits (120), Expect(3) = 3e-09
Identities = 23/35 (65%), Positives = 25/35 (70%)
Frame = +2
Query: 460 NPCKFSLFLAQMLERHGCWKFLHENSRKSSCISRN 494
N +LFL QMLE HGCW FLHENS KS +SRN
Sbjct: 377 NRANSALFLVQMLELHGCWNFLHENSCKSFPVSRN 481
Score = 23.9 bits (50), Expect(3) = 3e-09
Identities = 11/19 (57%), Positives = 12/19 (62%)
Frame = +3
Query: 500 FFSNFNAILCVF*RKCAKL 518
F NF+AI V R CAKL
Sbjct: 498 FCPNFHAIFSVLWRNCAKL 554
Score = 22.3 bits (46), Expect(3) = 3e-09
Identities = 9/11 (81%), Positives = 9/11 (81%)
Frame = +1
Query: 455 EILAFNPCKFS 465
EILA PCKFS
Sbjct: 361 EILASKPCKFS 393
>AV411435
Length = 426
Score = 47.4 bits (111), Expect = 6e-06
Identities = 24/73 (32%), Positives = 33/73 (44%)
Frame = +3
Query: 354 HPKFAVMKLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEH 413
HP + IW A +KV FLW A +A+P ++ + D C C ED+ H
Sbjct: 150 HPSSFLWHTIWGAQVPKKVRSFLWRAANNAIPVKRNLKRRNMGRDDFCPICNKGPEDINH 329
Query: 414 ALRRCPGSVWVRS 426
AL C W R+
Sbjct: 330 ALLACE---WTRA 359
>BP076564
Length = 371
Score = 44.7 bits (104), Expect = 4e-05
Identities = 23/58 (39%), Positives = 31/58 (52%), Gaps = 2/58 (3%)
Frame = +2
Query: 370 EKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPGS--VWVR 425
EK F + L ALP N +R+ + C RC A+V+D+ H L+ CP S VW R
Sbjct: 68 EKCKFMVGLGLHQALPINARRHACSMAATGGCGRCSAAVKDILHCLQDCPHSREVWNR 241
>BP068806
Length = 418
Score = 36.2 bits (82), Expect = 0.015
Identities = 20/58 (34%), Positives = 27/58 (46%), Gaps = 2/58 (3%)
Frame = +3
Query: 370 EKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC--PGSVWVR 425
EK+ ++W + A+ N R+ L A RC ED+ H LR C VWVR
Sbjct: 156 EKIRVWVWFIMQGAIQVNSFRFRCKLATSPAGPRCQHQFEDIWHCLRDCRIARDVWVR 329
>AV780445
Length = 504
Score = 32.3 bits (72), Expect = 0.21
Identities = 15/32 (46%), Positives = 20/32 (61%), Gaps = 2/32 (6%)
Frame = +2
Query: 396 VKFDACFRCGASVEDVEHALRRCPGS--VWVR 425
+ F + FRC A ED+ H LR CP S +W+R
Sbjct: 44 LSFFSFFRCPAPEEDI*HCLRDCPHSQEIWLR 139
>AV426428
Length = 413
Score = 31.6 bits (70), Expect = 0.36
Identities = 17/57 (29%), Positives = 25/57 (43%)
Frame = +3
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP 419
+WKA A + W A LP + L + C RC A+ + +HA+ CP
Sbjct: 54 LWKAEALPRCCETGWRACLGVLPVKASLHARGLAEDPICPRCLAAPKTADHAVLFCP 224
>TC17647 weakly similar to UP|AAR06918 (AAR06918) UDP-glycosyltransferase
91D1, partial (13%)
Length = 1059
Score = 29.3 bits (64), Expect = 1.8
Identities = 15/35 (42%), Positives = 18/35 (50%)
Frame = +3
Query: 459 FNPCKFSLFLAQMLERHGCWKFLHENSRKSSCISR 493
FNP + LFL + HG WK H N R IS+
Sbjct: 138 FNPVRVFLFLIIISYLHGWWKQPHRNHRHLVGISK 242
>TC10037
Length = 560
Score = 27.7 bits (60), Expect = 5.3
Identities = 13/28 (46%), Positives = 15/28 (53%), Gaps = 2/28 (7%)
Frame = -3
Query: 401 CFRCGASVEDVEHALRRCP--GSVWVRS 426
C RC E ++HALR CP VW S
Sbjct: 387 CLRCDRLEETMDHALRDCP*ARRVWFAS 304
>TC11973 weakly similar to UP|AAR34269 (AAR34269) Inositol-1-monophosphatase
, partial (26%)
Length = 669
Score = 27.3 bits (59), Expect = 6.9
Identities = 19/70 (27%), Positives = 27/70 (38%)
Frame = +1
Query: 454 HEILAFNPCKFSLFLAQMLERHGCWKFLHENSRKSSCISRNCHVVLFFSNFNAILCVF*R 513
H +F C+ L ++ +GCW F SR+ S F + LC F*
Sbjct: 28 HVSCSFGNCRSLLGISSKAMGYGCWCFDGGGSRRHS----------FSDGWRKFLC-F** 174
Query: 514 KCAKLRWGRP 523
C +W P
Sbjct: 175 ICIGFKWSAP 204
>CB829511
Length = 572
Score = 27.3 bits (59), Expect = 6.9
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = -1
Query: 462 CKFSLFLAQMLERHGCWKFLHENSRKSSCI 491
CK L + +LERH CW H + SC+
Sbjct: 323 CKTHLLIYPLLERHLCW***HLSQSLHSCL 234
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.339 0.147 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,689,668
Number of Sequences: 28460
Number of extensions: 182201
Number of successful extensions: 1350
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1339
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1349
length of query: 573
length of database: 4,897,600
effective HSP length: 95
effective length of query: 478
effective length of database: 2,193,900
effective search space: 1048684200
effective search space used: 1048684200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0369c.1


