

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
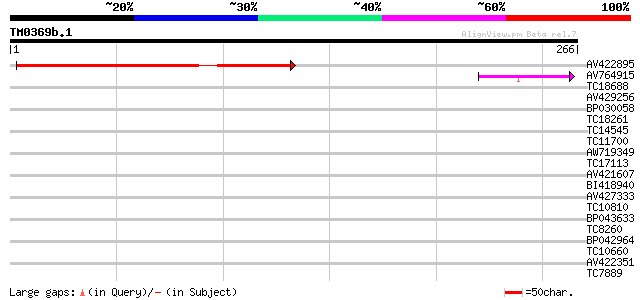
Query= TM0369b.1
(266 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV422895 185 8e-48
AV764915 46 6e-06
TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%) 39 0.001
AV429256 38 0.002
BP030058 36 0.006
TC18261 similar to UP|WSC2_YEAST (P53832) Cell wall integrity an... 36 0.006
TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial ... 35 0.010
TC11700 homologue to GB|AAL06971.1|15810091|AY056083 AT4g30260/F... 35 0.010
AW719349 35 0.010
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 34 0.022
AV421607 34 0.022
BI418940 34 0.022
AV427333 34 0.029
TC10810 similar to UP|PPF1_PEA (Q9FY06) Inner membrane protein P... 34 0.029
BP043633 33 0.038
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 33 0.038
BP042964 33 0.038
TC10660 similar to UP|Q8H5M9 (Q8H5M9) OJ1047_A06.31 protein, par... 33 0.038
AV422351 33 0.038
TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 33 0.038
>AV422895
Length = 477
Score = 185 bits (469), Expect = 8e-48
Identities = 96/131 (73%), Positives = 109/131 (82%)
Frame = +2
Query: 4 QRTSLLNCAYFFQGKSMEEVRQSLVYTTLELEQTRFAVQEELRKRDDQLLNLKDLLNNTI 63
QR LL AYF QGKSMEE++Q+L+YTTLELEQTR VQ+ELRKRD+Q+L LK+LLN +
Sbjct: 101 QRNPLLTWAYFCQGKSMEELKQTLMYTTLELEQTRTVVQDELRKRDEQVLTLKELLNRVM 280
Query: 64 RERDEAQEKCQRLLLEKFLFQHQQQQQQQQADPNSGVSSIEDEPRRGIDSNNGLSSSDCE 123
RERDEAQEKCQRLLLEK +FQ QQQ SG+SSIED+PRRGIDSNNGLSSSDCE
Sbjct: 281 RERDEAQEKCQRLLLEKLVFQKQQQL--------SGISSIEDDPRRGIDSNNGLSSSDCE 436
Query: 124 ESIVSSPVFDN 134
ESIVSSPV D+
Sbjct: 437 ESIVSSPVIDH 469
>AV764915
Length = 238
Score = 46.2 bits (108), Expect = 6e-06
Identities = 23/53 (43%), Positives = 32/53 (59%), Gaps = 8/53 (15%)
Frame = -2
Query: 221 LHQESFFNGSSSTSTHS--------QCGRVSRKRIFCDGSDSPSETKFQRVVL 265
L Q+SF++ S T CGR S+K+ C+G DSP+ETK+QR+VL
Sbjct: 237 LPQDSFYSNGSPGGTGPGTGPAPAPNCGRFSKKKELCEGFDSPAETKYQRIVL 79
>TC18688 weakly similar to UP|Q9LP77 (Q9LP77) T1N15.9, partial (13%)
Length = 514
Score = 38.9 bits (89), Expect = 0.001
Identities = 41/141 (29%), Positives = 54/141 (38%), Gaps = 9/141 (6%)
Frame = +3
Query: 117 LSSSDCEESIVSSPV-FDNMPQPQFP---LPLPSVAQSMIELTPEKPLPEKGKLLQAVMK 172
LSSS S SSP+ + + P P L P + TP P P G A
Sbjct: 66 LSSSSSSSSSSSSPLSLNQISPPNAPPS*LSAPPSPEERFSGTPPPPPPATGSASTATPT 245
Query: 173 AGPLLQTLLLAGPLPQWRH----PPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFN 228
++ P P H PP P + + V+ PSP PP PP
Sbjct: 246 PPTSSRSASPPSPSPANSHMASSPPSPTSALSVS-VSTPSPAHSPPTSPPA--------- 395
Query: 229 GSSSTSTHSQCGR-VSRKRIF 248
S+TST S+ VS +R+F
Sbjct: 396 PRSATSTSSRISSPVSSRRLF 458
>AV429256
Length = 337
Score = 38.1 bits (87), Expect = 0.002
Identities = 24/53 (45%), Positives = 26/53 (48%)
Frame = +3
Query: 173 AGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQES 225
A P TLL LP+ PPPP S + P PSPPL PP LLH S
Sbjct: 6 APPSTSTLLRPVSLPESSLPPPPTCSLNLLPGLPPSPPLTPP-----LLHLTS 149
>BP030058
Length = 455
Score = 36.2 bits (82), Expect = 0.006
Identities = 15/27 (55%), Positives = 16/27 (58%)
Frame = -1
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQPP 218
PPPP S PP+ PSPP PP PP
Sbjct: 404 PPPPSPSPPPPPLPCPSPPPPPPSPPP 324
Score = 33.9 bits (76), Expect = 0.029
Identities = 16/34 (47%), Positives = 16/34 (47%)
Frame = -1
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P PPPP PP PSPP PP PP
Sbjct: 404 PPPPSPSPPPPPLPCPSPPPPPPSPPPPPPPSPP 303
Score = 31.6 bits (70), Expect = 0.15
Identities = 15/34 (44%), Positives = 15/34 (44%)
Frame = -1
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P PPPPL PP PP PP PP
Sbjct: 401 PPPSPSPPPPPLPCPSPPPPPPSPPPPPPPSPPP 300
Score = 26.2 bits (56), Expect = 6.1
Identities = 18/47 (38%), Positives = 22/47 (46%), Gaps = 9/47 (19%)
Frame = -1
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQP--PQQ-------PPQLLH 222
PLP PPPP S PP P PP P PQ PP++++
Sbjct: 371 PLP-CPSPPPPPPSPPPPPPPSPPPPKYPSCPQNCNPLSPPPPRIIY 234
>TC18261 similar to UP|WSC2_YEAST (P53832) Cell wall integrity and stress
response component 2 precursor, partial (5%)
Length = 538
Score = 36.2 bits (82), Expect = 0.006
Identities = 24/67 (35%), Positives = 31/67 (45%)
Frame = +3
Query: 193 PPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHSQCGRVSRKRIFCDGS 252
P PL + + PP SPP PP QPP L + +SS+S +S+ S S
Sbjct: 168 PNPLNAAQSPPT---SPPQPPPLQPPTSLSLMTSTTSTSSSSPNSRSSAASAS----SSS 326
Query: 253 DSPSETK 259
SPS K
Sbjct: 327 PSPSSVK 347
>TC14545 similar to UP|Q9FT78 (Q9FT78) P23 co-chaperone, partial (46%)
Length = 1127
Score = 35.4 bits (80), Expect = 0.010
Identities = 38/152 (25%), Positives = 55/152 (36%), Gaps = 1/152 (0%)
Frame = -2
Query: 110 GIDSNNGLSSSDCEESIVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQA 169
G D++ SS + SI SSP+ N+ + P P + P P+P
Sbjct: 697 GFDTSCSSSSLPSKSSIASSPIPPNLEKSIPPNP---------SIPPNPPIP-------- 569
Query: 170 VMKAGPLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNG 229
P+P P PP+ P + P PP P PP +G
Sbjct: 568 ---------------PIP----PIPPI------PPSPPRPPSPPIPIPPNFEKSIPPISG 464
Query: 230 SSSTSTHSQCGRVSRKRIF-CDGSDSPSETKF 260
S++TS+ S +S+ C G SP F
Sbjct: 463 SATTSSSSSSTHLSQSTFT*CGGLPSPRNNLF 368
>TC11700 homologue to GB|AAL06971.1|15810091|AY056083 AT4g30260/F9N11_110
{Arabidopsis thaliana;}, partial (45%)
Length = 561
Score = 35.4 bits (80), Expect = 0.010
Identities = 34/119 (28%), Positives = 49/119 (40%), Gaps = 9/119 (7%)
Frame = +3
Query: 123 EESIVSSPVFDNMPQPQFPLPLPSVAQSMIELT---PEKPLPEKGKLLQAVMKAGPLLQT 179
EES+V +P P LP P+ +S I T P P+ L + PLL +
Sbjct: 12 EESVVRK-CHTAIPYPFTHLPNPTSTRSRISSTHLRPPSSPPDHPVLHAPPSPSPPLLPS 188
Query: 180 LLLAGPLPQWRHPPPPLESFEIPPVTI------PSPPLQPPQQPPQLLHQESFFNGSSS 232
+ L +PP P +F+ P +++ P P PP P L+ S F SS
Sbjct: 189 SIPTSLLLFLPNPPSPNLNFQNPSLSLLLLLPPPDPKSPPPASDPLLIPSPSQFGTPSS 365
>AW719349
Length = 143
Score = 35.4 bits (80), Expect = 0.010
Identities = 18/38 (47%), Positives = 21/38 (54%), Gaps = 5/38 (13%)
Frame = -2
Query: 187 PQWRHPPPPLESFEIPP-----VTIPSPPLQPPQQPPQ 219
PQ HPPP + PP + +P PP QPPQ PPQ
Sbjct: 109 PQPPHPPP-----QPPPQPPLFIILPQPPPQPPQPPPQ 11
Score = 32.3 bits (72), Expect = 0.085
Identities = 22/45 (48%), Positives = 24/45 (52%), Gaps = 1/45 (2%)
Frame = -2
Query: 175 PLLQTLLLAGPLPQWRHPP-PPLESFEIPPVTIPSPPLQPPQQPP 218
P L T+ P P + PP PPL F I P P PP QPP QPP
Sbjct: 130 PPLSTIFPQPPHPPPQPPPQPPL--FIILPQPPPQPP-QPPPQPP 5
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 34.3 bits (77), Expect = 0.022
Identities = 26/95 (27%), Positives = 35/95 (36%), Gaps = 2/95 (2%)
Frame = +1
Query: 126 IVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
++ P P P FP P + + P +P P + + P Q
Sbjct: 535 LLPDPFQPPSPPPLFPNPFQPPSPPPLIPNPFQPPPSPPPFIPNPFQPPPSKQ------- 693
Query: 186 LPQWRHPPPPLESFEIPPVTIP--SPPLQPPQQPP 218
PP PL F PP+ IP +PP PP PP
Sbjct: 694 ------PPSPL--FPFPPIVIPGLTPPPPPPPPPP 774
Score = 27.3 bits (59), Expect = 2.7
Identities = 17/55 (30%), Positives = 22/55 (39%), Gaps = 11/55 (20%)
Frame = +1
Query: 175 PLLQTLLLAGPLPQWRHPPPPLESFEIPPVTIPSP-----------PLQPPQQPP 218
PL+ +P PPP + + PP IP+P P QPP PP
Sbjct: 409 PLIPNPFQPPLVPNPFQPPPLIPNPFQPPPLIPNPFQPSPPPLLPDPFQPPSPPP 573
Score = 26.6 bits (57), Expect = 4.7
Identities = 13/27 (48%), Positives = 15/27 (55%), Gaps = 1/27 (3%)
Frame = +1
Query: 193 PPPLESFEIPPVTIPSPPLQP-PQQPP 218
P P + F PP+ PPL P P QPP
Sbjct: 358 PDPEDFFFFPPIPFFPPPLIPNPFQPP 438
>AV421607
Length = 245
Score = 34.3 bits (77), Expect = 0.022
Identities = 18/47 (38%), Positives = 25/47 (52%), Gaps = 2/47 (4%)
Frame = +1
Query: 174 GPLLQTLLLAGPLPQWRHPPPPLE--SFEIPPVTIPSPPLQPPQQPP 218
GP++ L P+P PPPL SF +PP + P PP + P+ P
Sbjct: 82 GPVIALPPLPIPIPTLTLTPPPLPILSFLLPPPSNPDPPSRNPKNSP 222
>BI418940
Length = 550
Score = 34.3 bits (77), Expect = 0.022
Identities = 24/81 (29%), Positives = 33/81 (40%)
Frame = +3
Query: 140 FPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPPLESF 199
+P P P+ A + TP PL E +G L L + +H P P
Sbjct: 327 YPPPTPAPATTQTPTTPPLPLSEN---------SGKLSNHSQQVIAL*RAKHQPIPPSPS 479
Query: 200 EIPPVTIPSPPLQPPQQPPQL 220
+PP P PP+ P PP+L
Sbjct: 480 SLPPTPPPPPPISTP--PPRL 536
>AV427333
Length = 387
Score = 33.9 bits (76), Expect = 0.029
Identities = 17/38 (44%), Positives = 19/38 (49%)
Frame = +3
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPPQLLH 222
P ++ PPPP SF PP PSPP P PP H
Sbjct: 150 PSHYFKSPPPPSHSFAPPPRGHPSPP--PSSPPPPSSH 257
Score = 29.6 bits (65), Expect = 0.55
Identities = 14/34 (41%), Positives = 14/34 (41%)
Frame = +3
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P R PP P PP P PP PP P
Sbjct: 270 PPPHVRPPPSPHHPITPPPHVRPPPPPLPPSPSP 371
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/33 (39%), Positives = 15/33 (45%)
Frame = +3
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQP 217
P P PPPP P T P P ++PP P
Sbjct: 216 PSPPPSSPPPPSSH----PFTPPPPHVRPPPSP 302
Score = 25.8 bits (55), Expect = 8.0
Identities = 13/34 (38%), Positives = 14/34 (40%)
Frame = +3
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P PPP PP + P PP P PP
Sbjct: 174 PPPSHSFAPPP-RGHPSPPPSSPPPPSSHPFTPP 272
>TC10810 similar to UP|PPF1_PEA (Q9FY06) Inner membrane protein PPF-1,
chloroplast precursor (Post-floral-specific protein 1),
partial (30%)
Length = 502
Score = 33.9 bits (76), Expect = 0.029
Identities = 33/109 (30%), Positives = 39/109 (35%)
Frame = +2
Query: 136 PQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPPP 195
P F L L +S+ L P G P Q LA PL P P
Sbjct: 59 PTHSFLLHLSPALRSLHSLAATSPTARPGYSPPRPSTKSP--QFTPLAIPLTSPESSPGP 232
Query: 196 LESFEIPPVTIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHSQCGRVSR 244
F P T PS PL P+ P LLH+ + +S S C R R
Sbjct: 233 KVFFTRSP-TPPSSPLILPRLPSMLLHRRTEAGSASFPKPWSTCSRCRR 376
>BP043633
Length = 542
Score = 33.5 bits (75), Expect = 0.038
Identities = 17/44 (38%), Positives = 20/44 (44%), Gaps = 4/44 (9%)
Frame = +1
Query: 185 PLPQWRHPPPP----LESFEIPPVTIPSPPLQPPQQPPQLLHQE 224
P PQW PPPP + P P PP PP +PP H +
Sbjct: 385 PPPQWGPPPPPPPPPSNPYSYHPKQFPPPP--PPPRPPHPPHPQ 510
Score = 26.2 bits (56), Expect = 6.1
Identities = 19/56 (33%), Positives = 26/56 (45%), Gaps = 4/56 (7%)
Frame = +3
Query: 186 LPQWRHPPPPLESFEIPPV----TIPSPPLQPPQQPPQLLHQESFFNGSSSTSTHS 237
LP PPP +SF V T S ++P PPQ + S + SSS ++ S
Sbjct: 342 LPPISVPPPIPDSFPSTAVGATTTAASSSVEPLFLPPQAVSSSSSSSSSSSPASSS 509
Score = 25.8 bits (55), Expect = 8.0
Identities = 16/45 (35%), Positives = 18/45 (39%), Gaps = 3/45 (6%)
Frame = +1
Query: 185 PLPQWRHP---PPPLESFEIPPVTIPSPPLQPPQQPPQLLHQESF 226
P Q+ HP P P PP P PP PP P H + F
Sbjct: 346 PQFQYHHPSQTPSP------PPQWGPPPPPPPPPSNPYSYHPKQF 462
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 33.5 bits (75), Expect = 0.038
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +3
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQPP 218
P P + PPPP+ ++ P SPP PP + P
Sbjct: 435 PHPVYHSPPPPVHTYPHPHPVYHSPPPPPPHEKP 536
Score = 29.6 bits (65), Expect = 0.55
Identities = 14/33 (42%), Positives = 17/33 (51%)
Frame = +3
Query: 185 PLPQWRHPPPPLESFEIPPVTIPSPPLQPPQQP 217
P P + PPPP +E P SPP PP +P
Sbjct: 201 PHPVYHSPPPP---YEKPHPVYHSPPPPPPHKP 290
Score = 28.1 bits (61), Expect = 1.6
Identities = 10/26 (38%), Positives = 14/26 (53%)
Frame = +3
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQP 217
PPPP+ ++ P P PP+ P P
Sbjct: 555 PPPPVHTYPHPVYHSPPPPVHSPPPP 632
Score = 27.7 bits (60), Expect = 2.1
Identities = 25/85 (29%), Positives = 33/85 (38%)
Frame = +3
Query: 130 PVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQW 189
PV+ + P P P P +P P P + K + P + T P P +
Sbjct: 441 PVYHSPPPPVHTYPHPHPVYH----SPPPPPPHE-KPYKYASPPPPPVHTY----PHPVY 593
Query: 190 RHPPPPLESFEIPPVTIPSPPLQPP 214
PPPP+ S P SPP PP
Sbjct: 594 HSPPPPVHSPPPPHYYYKSPP--PP 662
Score = 26.9 bits (58), Expect = 3.6
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = +3
Query: 192 PPPPLESFEIPPVTIPSPPLQPPQQPP 218
PPP S+ PP + SPP PP + P
Sbjct: 129 PPPKPYSYHSPPPPVHSPP--PPYEKP 203
>BP042964
Length = 575
Score = 33.5 bits (75), Expect = 0.038
Identities = 16/30 (53%), Positives = 16/30 (53%)
Frame = +3
Query: 191 HPPPPLESFEIPPVTIPSPPLQPPQQPPQL 220
H PPP P P PPLQPPQ PP L
Sbjct: 81 HSPPPQT-----PTQTPHPPLQPPQIPPLL 155
>TC10660 similar to UP|Q8H5M9 (Q8H5M9) OJ1047_A06.31 protein, partial (12%)
Length = 417
Score = 33.5 bits (75), Expect = 0.038
Identities = 24/63 (38%), Positives = 27/63 (42%), Gaps = 3/63 (4%)
Frame = -3
Query: 160 LPEKGKLLQAVMKAGPLLQTLLLAGPLPQWR---HPPPPLESFEIPPVTIPSPPLQPPQQ 216
LP L A A P L L ++ P P+WR H PP S PV P PP
Sbjct: 196 LPHAPPPLAAPSYAPPPL--LRISPPAPRWRTPSHAPPSCPSLLPSPVRQERGPSPPPPP 23
Query: 217 PPQ 219
PPQ
Sbjct: 22 PPQ 14
>AV422351
Length = 489
Score = 33.5 bits (75), Expect = 0.038
Identities = 23/68 (33%), Positives = 30/68 (43%), Gaps = 11/68 (16%)
Frame = +2
Query: 187 PQWRHPPPPLESFEIPPVTIP-SPPLQPPQQ----PPQLLHQESF------FNGSSSTST 235
P PPP + PP T SPP PP + PP+ Q F GS+++S
Sbjct: 152 PTITSPPPTAATERYPPTTTSISPPTPPPARKQSTPPRSSAQTMFSALTASSTGSTASSC 331
Query: 236 HSQCGRVS 243
H C R+S
Sbjct: 332 HVPCRRIS 355
>TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (66%)
Length = 1158
Score = 33.5 bits (75), Expect = 0.038
Identities = 28/95 (29%), Positives = 38/95 (39%), Gaps = 2/95 (2%)
Frame = +1
Query: 126 IVSSPVFDNMPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGP 185
IV P+ P P P P P V ++ P PLP V+ + P++ P
Sbjct: 268 IVKPPIV--YPPPYVPTP-PIVKPPIVYPPPYVPLPP-------VVPSPPIVTPPTPTPP 417
Query: 186 LPQWRHPPPPLESFEIPPVTIPSPPLQP--PQQPP 218
+ P PP+ + PP P PP P P PP
Sbjct: 418 IVTPPTPTPPVVTPPSPPTETPCPPPPPVVPSPPP 522
Score = 28.9 bits (63), Expect = 0.94
Identities = 21/87 (24%), Positives = 34/87 (38%), Gaps = 7/87 (8%)
Frame = +1
Query: 135 MPQPQFPLPLPSVAQSMIELTPEKPLPEKGKLLQAVMKAGPLLQTLLLAGPLPQWRHPPP 194
+P+P P P V + + P P P + + P++ P P +PPP
Sbjct: 70 VPKPPVVKPPPYVPKPPVVSPPYVPKPPVVPVTPPYVPKPPVVPVTPPYVPKPPIVYPPP 249
Query: 195 PLESFEI-------PPVTIPSPPLQPP 214
+ I PP +P+PP+ P
Sbjct: 250 YVPKPPIVKPPIVYPPPYVPTPPIVKP 330
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.132 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,375,678
Number of Sequences: 28460
Number of extensions: 119168
Number of successful extensions: 2701
Number of sequences better than 10.0: 389
Number of HSP's better than 10.0 without gapping: 1668
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2273
length of query: 266
length of database: 4,897,600
effective HSP length: 89
effective length of query: 177
effective length of database: 2,364,660
effective search space: 418544820
effective search space used: 418544820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0369b.1


