

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0367.7
(172 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
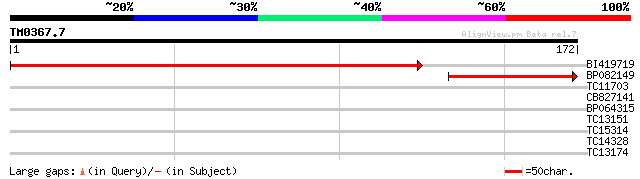
BI419719 206 2e-54
BP082149 85 8e-18
TC11703 homologue to UP|Q9ZSQ6 (Q9ZSQ6) T-complex protein 1 epsi... 29 0.38
CB827141 27 1.4
BP064315 26 4.2
TC13151 similar to UP|Q9SKS8 (Q9SKS8) At2g20920 protein (At2g209... 25 5.5
TC15314 25 7.2
TC14328 homologue to GB|AAG21976.1|10716947|AF263377 SKP1 intera... 25 9.4
TC13174 similar to UP|Q8GYL6 (Q8GYL6) Regulator of chromosome co... 25 9.4
>BI419719
Length = 531
Score = 206 bits (523), Expect = 2e-54
Identities = 96/125 (76%), Positives = 113/125 (89%)
Frame = +2
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
NV ++KSSKDTRY +DS+V+HDG K PCWAL +L SFK+K+G +AY++LDVIGIDEAQFF
Sbjct: 155 NVVMLKSSKDTRYAIDSVVSHDGVKFPCWALPDLMSFKEKYGHEAYQKLDVIGIDEAQFF 334
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
+DLYDFC +AAD DGK VVVAGLDG+YLRR+FGSVL IIPLAD+VTKLTARCE+CGK AF
Sbjct: 335 EDLYDFCCKAADVDGKIVVVAGLDGDYLRRSFGSVLHIIPLADTVTKLTARCELCGKRAF 514
Query: 121 FTLRK 125
FTLRK
Sbjct: 515 FTLRK 529
>BP082149
Length = 449
Score = 84.7 bits (208), Expect = 8e-18
Identities = 39/39 (100%), Positives = 39/39 (100%)
Frame = -3
Query: 134 IGGVDVYMPVCRQHYVNGQVAMETTRLVLESQKVECGSH 172
IGGVDVYMPVCRQHYVNGQVAMETTRLVLESQKVECGSH
Sbjct: 447 IGGVDVYMPVCRQHYVNGQVAMETTRLVLESQKVECGSH 331
>TC11703 homologue to UP|Q9ZSQ6 (Q9ZSQ6) T-complex protein 1 epsilon subunit
(Fragment), partial (53%)
Length = 665
Score = 29.3 bits (64), Expect = 0.38
Identities = 13/43 (30%), Positives = 23/43 (53%)
Frame = +1
Query: 38 KQKFGVDAYEQLDVIGIDEAQFFDDLYDFCREAADHDGKTVVV 80
K K +D E+ + E ++FDD+ C++A G T+V+
Sbjct: 259 KHKVDIDTVEKFQTLRKQEVKYFDDMVQQCKDA----GATLVI 375
>CB827141
Length = 513
Score = 27.3 bits (59), Expect = 1.4
Identities = 18/51 (35%), Positives = 22/51 (42%)
Frame = +3
Query: 91 NFGSVLDIIPLADSVTKLTARCEICGKHAFFTLRKTQETQVELIGGVDVYM 141
+ G L P+A +L E CGKH FF L + ELI YM
Sbjct: 342 DLGGFLSGDPIAKEAARLVG--EACGKHGFF-LVANHGIEAELISHAHSYM 485
>BP064315
Length = 350
Score = 25.8 bits (55), Expect = 4.2
Identities = 11/36 (30%), Positives = 21/36 (57%), Gaps = 1/36 (2%)
Frame = +1
Query: 12 RYGLDSIVTHDGTKLPCWALANLSSFKQKF-GVDAY 46
+Y ++ + HD +L CW N+S+ + F GV ++
Sbjct: 235 KYKMEELNDHDSLELFCWHAFNMSTPAENFEGVSSH 342
>TC13151 similar to UP|Q9SKS8 (Q9SKS8) At2g20920 protein
(At2g20920/F5H14.11), partial (21%)
Length = 562
Score = 25.4 bits (54), Expect = 5.5
Identities = 15/52 (28%), Positives = 26/52 (49%), Gaps = 6/52 (11%)
Frame = +2
Query: 70 AADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVT------KLTARCEIC 115
A ++ + +V+ LD +Y ++ + PL S+ KLT +CEIC
Sbjct: 326 AVENSLRVAIVSYLDIHYPASVLSEIIVVSPLLVSLYLRWRDGKLTLKCEIC 481
>TC15314
Length = 770
Score = 25.0 bits (53), Expect = 7.2
Identities = 8/22 (36%), Positives = 15/22 (67%)
Frame = +2
Query: 137 VDVYMPVCRQHYVNGQVAMETT 158
++V+ VC+ H++NG V + T
Sbjct: 11 MEVFQEVCQAHHINGTVRVHYT 76
>TC14328 homologue to GB|AAG21976.1|10716947|AF263377 SKP1 interacting
partner 1 {Arabidopsis thaliana;}, partial (5%)
Length = 620
Score = 24.6 bits (52), Expect = 9.4
Identities = 8/19 (42%), Positives = 12/19 (63%)
Frame = +1
Query: 100 PLADSVTKLTARCEICGKH 118
P VTK+ RC++C +H
Sbjct: 283 PNLSIVTKINPRCKLCNRH 339
>TC13174 similar to UP|Q8GYL6 (Q8GYL6) Regulator of chromosome condensation
like (Cell cycle regulatory protein), partial (25%)
Length = 519
Score = 24.6 bits (52), Expect = 9.4
Identities = 12/28 (42%), Positives = 18/28 (63%), Gaps = 3/28 (10%)
Frame = -3
Query: 74 DGKTVVVAGLDGNYL---RRNFGSVLDI 98
DG+ VVV G+D N + RRN+G + +
Sbjct: 226 DGEGVVVTGVDANNVGR*RRNWGRFVSV 143
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,612,697
Number of Sequences: 28460
Number of extensions: 29496
Number of successful extensions: 117
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 117
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 117
length of query: 172
length of database: 4,897,600
effective HSP length: 84
effective length of query: 88
effective length of database: 2,506,960
effective search space: 220612480
effective search space used: 220612480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0367.7


