

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
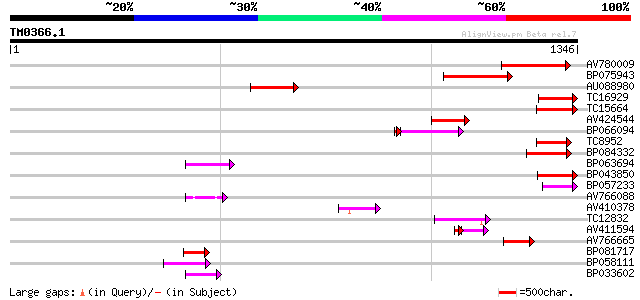
Query= TM0366.1
(1346 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV780009 228 5e-60
BP075943 164 1e-40
AU088980 142 5e-34
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 119 3e-27
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 115 4e-26
AV424544 109 3e-24
BP066094 93 4e-21
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 98 1e-20
BP084332 82 7e-16
BP063694 80 2e-15
BP043850 76 4e-14
BP057233 67 1e-11
AV766088 61 1e-09
AV410378 61 1e-09
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 61 1e-09
AV411594 56 2e-09
AV766665 59 5e-09
BP081717 55 1e-07
BP058111 55 1e-07
BP033602 54 2e-07
>AV780009
Length = 529
Score = 228 bits (581), Expect = 5e-60
Identities = 106/163 (65%), Positives = 129/163 (79%)
Frame = -1
Query: 1168 KAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSW 1227
+AA RVLRY+K +P GL F +S + +Q YSD+DWAGCPDTRRS++GY ++G SL+SW
Sbjct: 529 QAATRVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISW 350
Query: 1228 KAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAAN 1287
+ KKQTTVSRSS+EAEYRALA CE+QWL YL Q LK+ LFCDNQSALHIA N
Sbjct: 349 RTKKQTTVSRSSSEAEYRALAATVCEVQWLSYLFQFLKLNVPLPVPLFCDNQSALHIAHN 170
Query: 1288 PVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTK 1330
P FHERTKH+++DCHVVR KLQAG++ LLPI T Q+AD+FTK
Sbjct: 169 PTFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTK 41
>BP075943
Length = 547
Score = 164 bits (414), Expect = 1e-40
Identities = 90/165 (54%), Positives = 109/165 (65%), Gaps = 1/165 (0%)
Frame = -3
Query: 1029 SFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQ 1088
SFT LL+YVDDV+L GN + E QLVK SLH F IKDLG KYFLGLE+A STSGI L Q
Sbjct: 533 SFTALLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQ 354
Query: 1089 RKYCLDLLQETGTLGSKP-VATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPD 1147
RKY L L+ ++G G +P + S L G P D +YRR+VGRLLYL TTRPD
Sbjct: 353 RKYALQLISDSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTTRPD 174
Query: 1148 ISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNST 1192
I+ A QLSQ++S P D H + L+YL SPG GL +P +S+
Sbjct: 173 ITFAVNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFYPASSS 39
>AU088980
Length = 360
Score = 142 bits (357), Expect = 5e-34
Identities = 64/115 (55%), Positives = 87/115 (75%)
Frame = +2
Query: 571 NGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYRE 630
N VERKH+HILN+ RAL+F S LPK +W ++V HA +L+N +PS LLK + PY+ L +
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 631 AVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEI 685
+ L +KVFG+LC+++TLT+N KLDPRARKC+FLG+K G GY++MDL T I
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMDLPTTXI 346
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 119 bits (298), Expect = 3e-27
Identities = 53/92 (57%), Positives = 70/92 (75%)
Frame = +1
Query: 1255 QWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILK 1314
QWL YLLQDLKV +++CDN SA HIAANPVFHERTKH++IDCH+VRE++Q G++
Sbjct: 1 QWLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIH 180
Query: 1315 LLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
LLPI ++ +AD++TKAL P+ F KL +
Sbjct: 181 LLPISSSEPLADIYTKALSPQNFHQICAKLGL 276
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 115 bits (289), Expect = 4e-26
Identities = 56/95 (58%), Positives = 68/95 (70%)
Frame = +1
Query: 1252 CELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAG 1311
CEL WL Y+LQDL+V + +L+CD+QSA HIA N VFHERTKHLDIDCHVVREKLQA
Sbjct: 1 CEL*WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAK 180
Query: 1312 ILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
+ LLPI + Q AD+ TK L+ F +KL +
Sbjct: 181 LFHLLPISSVDQTADILTKPLESGPFSHLVSKLGV 285
>AV424544
Length = 276
Score = 109 bits (273), Expect = 3e-24
Identities = 56/90 (62%), Positives = 68/90 (75%)
Frame = +3
Query: 1002 KLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAF 1061
KLS++L LG+ Q+A DHSLF KF+ +SFT +LVYVDD+IL GN + E Q VK+ L F
Sbjct: 6 KLSSYLHILGYIQSAHDHSLFTKFRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQF 185
Query: 1062 GIKDLGVLKYFLGLEVAHSTSGISLCQRKY 1091
IKDLG LKYFLGLEVA S+ G+ L QRKY
Sbjct: 186 RIKDLGTLKYFLGLEVARSSCGLFLSQRKY 275
>BP066094
Length = 532
Score = 93.2 bits (230), Expect(2) = 4e-21
Identities = 52/149 (34%), Positives = 85/149 (56%), Gaps = 1/149 (0%)
Frame = +2
Query: 929 AKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVPSVK-PNQVCKLLK 987
+K +R++++ + + N LHQ++V +AFL+G + E+VY+ P G K + + KL K
Sbjct: 86 SKTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKK 265
Query: 988 SLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTV 1047
SLYGLKQA R WYE+LS+ L + D +LF K + +YVDD+I
Sbjct: 266 SLYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANP 445
Query: 1048 TEFQLVKDSLHQAFGIKDLGVLKYFLGLE 1076
+ + + + F ++ +G LKYFLG++
Sbjct: 446 SLCKEFSEMMQAEFEMRMMGELKYFLGIQ 532
Score = 26.6 bits (57), Expect(2) = 4e-21
Identities = 10/18 (55%), Positives = 15/18 (82%)
Frame = +3
Query: 913 YNQVEGLDYFDTFSPVAK 930
Y+Q G+DY +TF+PVA+
Sbjct: 39 YSQQ*GIDYTETFAPVAR 92
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 97.8 bits (242), Expect = 1e-20
Identities = 44/81 (54%), Positives = 61/81 (74%)
Frame = +3
Query: 1252 CELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAG 1311
CE WL Y L DL++ V++CDN+SALH+AAN VFH+RT++++IDCH+V K+ G
Sbjct: 18 CEALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFG 197
Query: 1312 ILKLLPIPTTLQVADVFTKAL 1332
IL LL +P++ QVADVFTK +
Sbjct: 198 ILHLLHVPSSDQVADVFTKTI 260
>BP084332
Length = 368
Score = 81.6 bits (200), Expect = 7e-16
Identities = 38/107 (35%), Positives = 69/107 (63%), Gaps = 1/107 (0%)
Frame = +2
Query: 1227 WKAK-KQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIA 1285
W+A Q+T++ S+ EAEY + A + ++ W+ + L+D ++ + P+ +CDN +A+ ++
Sbjct: 14 WEAT*GQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPI-YCDNTAAISLS 190
Query: 1286 ANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKAL 1332
NP+ H R KH+++ H +R+ +Q G+L L + T Q AD+FTK L
Sbjct: 191 KNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPL 331
>BP063694
Length = 511
Score = 80.5 bits (197), Expect = 2e-15
Identities = 41/118 (34%), Positives = 61/118 (50%), Gaps = 3/118 (2%)
Frame = -2
Query: 418 LWHFRLGHLSHDRLLALHT---VQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFD 474
LWH RLGH++ D + LH + S + K I C C +AK KR +F + +A D
Sbjct: 360 LWHSRLGHVNFDIIKQLHKHGCLDVSSILPKPICCTSCQMAKSKRLVFHDNNKRASAVLD 181
Query: 475 LVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQF 532
L+H D+ GP AS+ YF+ +DD+SRF W L K + + F ++ +F
Sbjct: 180 LIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVLLRFKVFMENRF 7
>BP043850
Length = 515
Score = 75.9 bits (185), Expect = 4e-14
Identities = 37/94 (39%), Positives = 57/94 (60%)
Frame = -1
Query: 1253 ELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGI 1312
E+ WL LL +L + L DN SA+ IAANPV+HE T+H+++DCH VRE +
Sbjct: 512 EIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRRV 333
Query: 1313 LKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
+ L + T++Q+AD+ TK+L + +KL +
Sbjct: 332 ITLPHVSTSVQIADILTKSLTRQRHNFLVSKLML 231
>BP057233
Length = 473
Score = 67.4 bits (163), Expect = 1e-11
Identities = 32/82 (39%), Positives = 49/82 (59%)
Frame = -2
Query: 1265 KVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQV 1324
K+ P+L+CDN SA +A+NPV H R+KH++ID H +R+++ + + +PTT Q+
Sbjct: 472 KLPMPRKPILWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQI 293
Query: 1325 ADVFTKALQPRVFQGFATKLAM 1346
AD TK L F KL +
Sbjct: 292 ADCLTKPLSHTRFSQLRDKLGV 227
>AV766088
Length = 501
Score = 60.8 bits (146), Expect = 1e-09
Identities = 36/104 (34%), Positives = 55/104 (52%), Gaps = 3/104 (2%)
Frame = -3
Query: 417 ALWHFRLGHLSHDRLLALHTVQNSIS---VSKSIVCDVCHLAKQKRKMFTVSVSKAQKCF 473
+LWH RLGH S L L + IS ++ S VC+ C K R F S + F
Sbjct: 310 SLWHSRLGHPSSSALRYLRS-NKFISYELLNYSPVCESCVFGKHVRLPFVSSNNVTVMPF 134
Query: 474 DLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEV 517
D++H D+W +S H++++ LDD++ F+W L+NK +V
Sbjct: 133 DILHSDLWTSPVLSSA-GHRFYVLFLDDFTDFLWTFPLSNKSQV 5
>AV410378
Length = 358
Score = 60.8 bits (146), Expect = 1e-09
Identities = 37/106 (34%), Positives = 59/106 (54%), Gaps = 8/106 (7%)
Frame = +1
Query: 782 EGEVEVRRSTRPLKRPVHLANFQ--------FSALPIRSSTAHSIVHHYSDERLSTAHRA 833
EG V+ R S R + P +L ++ S +P + ++ + S +L+ ++RA
Sbjct: 34 EGNVQ-RISNRVRRPPSYLQDYHCTLAASTTVSKVPSSAGISYPLSKVISYHKLTPSYRA 210
Query: 834 YALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLP 879
+ +NI EP+ ++EA K WR+AM EI+ALE+N TW LVD P
Sbjct: 211 FIMNITTVSEPTRYSEAVKHECWRKAMDQEIEALERNHTWILVDKP 348
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 60.8 bits (146), Expect = 1e-09
Identities = 42/141 (29%), Positives = 67/141 (46%), Gaps = 9/141 (6%)
Frame = +2
Query: 1009 TLGFKQTASDHSLFVK-FQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLG 1067
+LG+ + +SDH + K F + F LL+YVDD+++ G Q +K L + F +KDLG
Sbjct: 20 SLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREFDMKDLG 199
Query: 1068 VLKYFLGLEVAHSTSG--ISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRL------SQ 1119
LG+++ I L Q+ Y +L+ P++TPL + +L S
Sbjct: 200 PANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSSSMIPSS 379
Query: 1120 EQGKPYDDPAAYRRLVGRLLY 1140
E + Y VG L+Y
Sbjct: 380 EAERMEMSRVPYASAVGSLMY 442
>AV411594
Length = 244
Score = 56.2 bits (134), Expect(2) = 2e-09
Identities = 29/65 (44%), Positives = 37/65 (56%)
Frame = +2
Query: 1072 FLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAY 1131
FL +A S GI L +RKY L LL +T +G KP PLDPSI+L+ G+ D Y
Sbjct: 50 FLRT*IAKSKKGILLSRRKYALQLLDDTRNMGCKPPPFPLDPSIKLNSTDGELLSDVTLY 229
Query: 1132 RRLVG 1136
R +G
Sbjct: 230 IRFIG 244
Score = 24.3 bits (51), Expect(2) = 2e-09
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = +3
Query: 1057 LHQAFGIKDLGVLKYFLGLE 1076
L ++F +K L +KYFLGLE
Sbjct: 6 LAKSFKLKVLDDMKYFLGLE 65
>AV766665
Length = 601
Score = 58.9 bits (141), Expect = 5e-09
Identities = 29/75 (38%), Positives = 46/75 (60%)
Frame = +3
Query: 1172 RVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKK 1231
R+ YLKA+ G LF + ++ G++ AD+ G R S GY ++ +LV+W++K+
Sbjct: 372 RIF*YLKANSRRGPLFQKEGKSSMDGFTYADYLGSIVDRLSTMGYYMFLSGNLVTWRSKQ 551
Query: 1232 QTTVSRSSNEAEYRA 1246
Q ++RSS EAE RA
Sbjct: 552 QNIIARSSGEAELRA 596
>BP081717
Length = 291
Score = 54.7 bits (130), Expect = 1e-07
Identities = 24/60 (40%), Positives = 36/60 (60%)
Frame = -1
Query: 414 PPGALWHFRLGHLSHDRLLALHTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCF 473
P +WHFRLGH S L + V + + +I+CD CH+AKQ + F +S +K++ CF
Sbjct: 180 PQFDMWHFRLGHPSSKVLQHISKVFPYVKLHSNIICDFCHMAKQTKLPFPISDTKSKACF 1
>BP058111
Length = 570
Score = 54.7 bits (130), Expect = 1e-07
Identities = 36/113 (31%), Positives = 56/113 (48%), Gaps = 1/113 (0%)
Frame = -1
Query: 366 EVQTQRRIGLGSLQPDELYHLVVSSSSSTCFSIQHSSTSEINDFGHIIPPGALWHFRLGH 425
E T + IG + D LY L+ + ++ + ++++ F LWH RLGH
Sbjct: 339 ERPTLKMIGAAEERGD-LYALIDPAVKVKHPPVEDNIKTQVSHFNFCQFDCNLWHLRLGH 163
Query: 426 LSHDRLLALHTVQNSISVSK-SIVCDVCHLAKQKRKMFTVSVSKAQKCFDLVH 477
S +L L I+ K + CD CH+AKQK+ F SV+ + + FDL+H
Sbjct: 162 TSSKKLAILQKNFPFITCKKITSPCDTCHMAKQKKLSFPNSVTLSSEIFDLIH 4
>BP033602
Length = 533
Score = 53.9 bits (128), Expect = 2e-07
Identities = 28/86 (32%), Positives = 43/86 (49%)
Frame = +3
Query: 418 LWHFRLGHLSHDRLLALHTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFDLVH 477
LWH RLGH + L L S+ C+ C LAK +R + A + F L+H
Sbjct: 276 LWHNRLGHPNFQYLRHLFPDLFKNVNCSSLECESCVLAKNQRAPYYSQPYHASRPFYLIH 455
Query: 478 MDIWGPLAQASVHNHKYFLTVLDDYS 503
D+WGP + ++F+T +DD++
Sbjct: 456 SDVWGPSKITTQFGKRWFVTFIDDHT 533
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.335 0.144 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,095,259
Number of Sequences: 28460
Number of extensions: 303053
Number of successful extensions: 2288
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 2250
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2279
length of query: 1346
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1245
effective length of database: 2,023,140
effective search space: 2518809300
effective search space used: 2518809300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0366.1


