

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
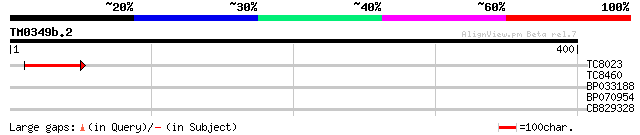
Query= TM0349b.2
(400 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8023 53 1e-07
TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F... 32 0.24
BP033188 30 0.71
BP070954 28 3.5
CB829328 27 6.0
>TC8023
Length = 1089
Score = 52.8 bits (125), Expect = 1e-07
Identities = 28/44 (63%), Positives = 32/44 (72%), Gaps = 1/44 (2%)
Frame = +3
Query: 11 FDLEAYLEKSEREDTYILNRFRDRRNLILEDSAPR-GRKHLSRD 53
FD+EAY +SE ED YI N FR RRNLILE PR RK+L+RD
Sbjct: 57 FDVEAYKRQSEIEDRYIFNLFRKRRNLILEGYTPRVKRKYLNRD 188
>TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;}, partial (36%)
Length = 880
Score = 31.6 bits (70), Expect = 0.24
Identities = 24/105 (22%), Positives = 44/105 (41%)
Frame = +3
Query: 68 NEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRML 127
+ P + + FRR +RM K F I +L S+ + + +E I + + L
Sbjct: 357 SHPDFPEEEFRRSFRMSKATFEMICRELDSA----VTKKNTMLREAIPVRQRVAVCIWRL 524
Query: 128 ANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQ 172
A G V + +G +T + + C I + ++LR P +
Sbjct: 525 ATGDPLRLVAKRFGLGISTCHKLVLEVCSAIKTVLMPKFLRWPDE 659
>BP033188
Length = 527
Score = 30.0 bits (66), Expect = 0.71
Identities = 13/35 (37%), Positives = 21/35 (59%), Gaps = 1/35 (2%)
Frame = -1
Query: 365 VDLVHRNHTRPRCYPLL-RIMCVLDPSCVIQKVHH 398
V ++ R H RP C L+ RI C+L+ + ++HH
Sbjct: 194 VSVLSRGHLRPNCKILVSRIFCMLERMSRVAEIHH 90
>BP070954
Length = 152
Score = 27.7 bits (60), Expect = 3.5
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = +2
Query: 371 NHTRPRCYPLLRIMCVLDPSCVIQKVHH 398
+HT C+ L C L P C+IQ V H
Sbjct: 65 HHTCDPCFALYL*ACALXPRCIIQCVLH 148
>CB829328
Length = 556
Score = 26.9 bits (58), Expect = 6.0
Identities = 13/47 (27%), Positives = 24/47 (50%)
Frame = +2
Query: 352 ILNAGPILSKLREVDLVHRNHTRPRCYPLLRIMCVLDPSCVIQKVHH 398
+L PIL+ L +N+ P+C +++C + SC Q++ H
Sbjct: 56 VLAYSPILNGTSIFQL*DKNNRSPKCRDHEKLICKVVSSCRKQQIKH 196
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.345 0.152 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,510,417
Number of Sequences: 28460
Number of extensions: 109403
Number of successful extensions: 728
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 714
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 726
length of query: 400
length of database: 4,897,600
effective HSP length: 92
effective length of query: 308
effective length of database: 2,279,280
effective search space: 702018240
effective search space used: 702018240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0349b.2


