

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0348.12
(180 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
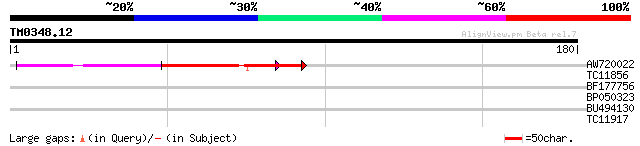
Score E
Sequences producing significant alignments: (bits) Value
AW720022 55 7e-09
TC11856 similar to GB|AAO63369.1|28950891|BT005305 At4g11960 {Ar... 27 2.1
BF177756 26 3.5
BP050323 26 4.6
BU494130 25 6.0
TC11917 25 7.8
>AW720022
Length = 530
Score = 55.1 bits (131), Expect = 7e-09
Identities = 29/46 (63%), Positives = 32/46 (69%)
Frame = +1
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFG 94
W GS+S V YPRCSIRRQS YADAE I + F WA+FLAFG
Sbjct: 394 WYGSKSVVKYPRCSIRRQSAYADAE-EDISMVFALASTWAVFLAFG 528
Score = 51.6 bits (122), Expect = 8e-08
Identities = 39/93 (41%), Positives = 52/93 (54%), Gaps = 9/93 (9%)
Frame = +2
Query: 3 SSRKSRTPSNRTPRLKLFVCNRHEMLIVVVSV**LMTCLIEWS*SIWLGSRSDV*YPRCS 62
SS S+ P+ + +L + +HEMLI+VV++**LM CLIEWS*S +S +
Sbjct: 263 SSVPSKPPTKKPSKLSIA---KHEMLIIVVNL**LMICLIEWS*S*DGMVQSQL------ 415
Query: 63 IRRQSTYADAEG---------RRIYLWFLH*QA 86
S+ DA R+IYLWFLH*QA
Sbjct: 416 ----SSILDAVSGGNPHMLMLRKIYLWFLH*QA 502
>TC11856 similar to GB|AAO63369.1|28950891|BT005305 At4g11960 {Arabidopsis
thaliana;}, partial (34%)
Length = 613
Score = 26.9 bits (58), Expect = 2.1
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = +3
Query: 164 DLVALRGACPNCGEE 178
D + L+G CPNCG E
Sbjct: 156 DFLILKGPCPNCGTE 200
>BF177756
Length = 455
Score = 26.2 bits (56), Expect = 3.5
Identities = 12/37 (32%), Positives = 20/37 (53%)
Frame = +1
Query: 135 NIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRGA 171
N++ +V+ F SAP+ +Q LW N ++ GA
Sbjct: 19 NVVALVINFFSACITGSAPLTAVQLLWVNLIMDTLGA 129
>BP050323
Length = 482
Score = 25.8 bits (55), Expect = 4.6
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = +2
Query: 119 SHGSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAP 153
+ GS N LGF ++ + IIF+ F YP P
Sbjct: 92 NQGSLNRELGFWVLFHKIIFVEKKFR*NYPFFYVP 196
>BU494130
Length = 433
Score = 25.4 bits (54), Expect = 6.0
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = +2
Query: 122 SHNPGLGFSIIVNNIIFIVLGF 143
S N GLGF + NNI I+L F
Sbjct: 50 SDNNGLGFHLTTNNIKAILLDF 115
>TC11917
Length = 820
Score = 25.0 bits (53), Expect = 7.8
Identities = 16/50 (32%), Positives = 21/50 (42%)
Frame = +3
Query: 95 SLACLGPLSYAVGIAYQNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGFV 144
+L CL +SY + + Y HGS N F I + I LG V
Sbjct: 474 TLCCLPMMSYFM*LCYNKLLMPSWLHGSLNQINAFPIKTLSFSIIKLGIV 623
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.340 0.149 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,729,524
Number of Sequences: 28460
Number of extensions: 54775
Number of successful extensions: 313
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 309
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 312
length of query: 180
length of database: 4,897,600
effective HSP length: 84
effective length of query: 96
effective length of database: 2,506,960
effective search space: 240668160
effective search space used: 240668160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0348.12


