

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0344.6
(242 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
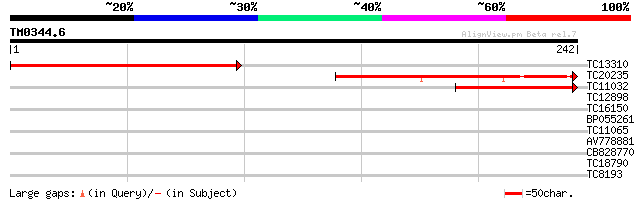
Sequences producing significant alignments: (bits) Value
TC13310 201 9e-53
TC20235 137 1e-33
TC11032 109 4e-25
TC12898 similar to UP|Q7PVN0 (Q7PVN0) ENSANGP00000010532 (Fragme... 32 0.075
TC16150 similar to GB|AAO63328.1|28950809|BT005264 At4g25840 {Ar... 28 1.9
BP055261 27 3.2
TC11065 weakly similar to UP|Q8RDP8 (Q8RDP8) Histidinol-phosphat... 26 5.4
AV778881 26 7.0
CB828770 26 7.0
TC18790 26 7.0
TC8193 similar to UP|Q9M5J4 (Q9M5J4) Beta-galactosidase (Lactas... 25 9.2
>TC13310
Length = 544
Score = 201 bits (511), Expect = 9e-53
Identities = 99/99 (100%), Positives = 99/99 (100%)
Frame = +1
Query: 1 MANLTLNQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAV 60
MANLTLNQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAV
Sbjct: 247 MANLTLNQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAV 426
Query: 61 QGPYQSISILDSRIANQHVRKYGYLHIGLVQVAVKPLTR 99
QGPYQSISILDSRIANQHVRKYGYLHIGLVQVAVKPLTR
Sbjct: 427 QGPYQSISILDSRIANQHVRKYGYLHIGLVQVAVKPLTR 543
>TC20235
Length = 543
Score = 137 bits (346), Expect = 1e-33
Identities = 73/105 (69%), Positives = 81/105 (76%), Gaps = 2/105 (1%)
Frame = +1
Query: 140 CFSSYPVCLRDPHVTTCVDLDIKIPNEMFIDGIPP-VSIMYRFCYKVTDNLFGIKAVRDY 198
C+ SYPV LRDPHVT C+DLD+KIPNEMF+DG PP VSIMYRFCYKVTD L G+KA R
Sbjct: 1 CYPSYPVSLRDPHVTKCLDLDVKIPNEMFVDGAPPPVSIMYRFCYKVTDYLRGVKAQRSN 180
Query: 199 TVNETEVTEIN-AENSSRITRKILKHDEITYPQEWLQGCSSQPAP 242
VNET+V E + A SS ITR ILKHD I YP+EWL G SS P P
Sbjct: 181 PVNETDVIETHIALYSSEITR-ILKHDRIAYPEEWLPGRSS-PIP 309
>TC11032
Length = 610
Score = 109 bits (273), Expect = 4e-25
Identities = 52/52 (100%), Positives = 52/52 (100%)
Frame = +1
Query: 191 GIKAVRDYTVNETEVTEINAENSSRITRKILKHDEITYPQEWLQGCSSQPAP 242
GIKAVRDYTVNETEVTEINAENSSRITRKILKHDEITYPQEWLQGCSSQPAP
Sbjct: 1 GIKAVRDYTVNETEVTEINAENSSRITRKILKHDEITYPQEWLQGCSSQPAP 156
>TC12898 similar to UP|Q7PVN0 (Q7PVN0) ENSANGP00000010532 (Fragment),
partial (4%)
Length = 784
Score = 32.3 bits (72), Expect = 0.075
Identities = 23/79 (29%), Positives = 33/79 (41%), Gaps = 4/79 (5%)
Frame = +1
Query: 37 VYKTSIFEFNPDYEIKTVEKSFAVQGPYQSISILDSRIANQHV----RKYGYLHIGLVQV 92
V+ T F+ +YE ++F LD IA +H ++Y L L
Sbjct: 481 VWSTRARVFHSEYEFLIPYQAFCPPHGVHCSCALDLCIATEHTDPYEQRYHVLEDPLPSP 660
Query: 93 AVKPLTRLRYCDLPILICL 111
A PLT L Y + I +CL
Sbjct: 661 AAPPLTHLPYLSISISVCL 717
>TC16150 similar to GB|AAO63328.1|28950809|BT005264 At4g25840 {Arabidopsis
thaliana;}, partial (90%)
Length = 1065
Score = 27.7 bits (60), Expect = 1.9
Identities = 16/56 (28%), Positives = 30/56 (53%), Gaps = 4/56 (7%)
Frame = -3
Query: 114 KRMLRFHESVVA---VLESNIQN-GPAYFNCFSSYPVCLRDPHVTTCVDLDIKIPN 165
+R+L F+E ++ + S++ N P+ F CF S+ +CL P + + K+ N
Sbjct: 412 QRVLSFYEKLLGAERITNSSLFNKDPSSFYCFISHHLCLEGPIKCFVISSEYKLLN 245
>BP055261
Length = 339
Score = 26.9 bits (58), Expect = 3.2
Identities = 22/76 (28%), Positives = 36/76 (46%), Gaps = 7/76 (9%)
Frame = +3
Query: 154 TTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIKAVRDYTVNETEVTEINAEN- 212
T + L+ + NE+FI + ++I RF + + F K DY V++ + E+ N
Sbjct: 45 TQSLRLNSTVYNELFIIQLKLLTIT*RFKSNIFLHFFLCKEEEDYIVHDMTLQEVPNNNQ 224
Query: 213 ------SSRITRKILK 222
S + T KILK
Sbjct: 225 EDLIHGSFKRTFKILK 272
>TC11065 weakly similar to UP|Q8RDP8 (Q8RDP8) Histidinol-phosphatase ,
partial (4%)
Length = 508
Score = 26.2 bits (56), Expect = 5.4
Identities = 10/37 (27%), Positives = 21/37 (56%)
Frame = -3
Query: 9 HANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEF 45
HA+ ++E ++ Q + + K++ + +YKTS F
Sbjct: 503 HAHATKEGIPSIHQNIYMEKVMKLKADMIYKTSKLSF 393
>AV778881
Length = 550
Score = 25.8 bits (55), Expect = 7.0
Identities = 17/40 (42%), Positives = 22/40 (54%), Gaps = 4/40 (10%)
Frame = +3
Query: 28 SIPKIRTNQVYKTSIFEFN----PDYEIKTVEKSFAVQGP 63
SIP + +N V TSI E N + E +V KSF+V P
Sbjct: 267 SIPYLASNSVMDTSIVEHNL*FANEEESDSVLKSFSV*NP 386
>CB828770
Length = 486
Score = 25.8 bits (55), Expect = 7.0
Identities = 10/38 (26%), Positives = 22/38 (57%)
Frame = -2
Query: 182 CYKVTDNLFGIKAVRDYTVNETEVTEINAENSSRITRK 219
CY +++ ++ +K++ + V ++ I NSSR R+
Sbjct: 215 CYTLSNQIYHVKSIMAFCVQN*KIALILKHNSSRWRRR 102
>TC18790
Length = 555
Score = 25.8 bits (55), Expect = 7.0
Identities = 8/20 (40%), Positives = 16/20 (80%)
Frame = -2
Query: 7 NQHANNSEENDSNLEQKLQN 26
NQH ++S +ND +++K++N
Sbjct: 419 NQHHSHSSDNDKQIKKKIRN 360
>TC8193 similar to UP|Q9M5J4 (Q9M5J4) Beta-galactosidase (Lactase) , partial
(80%)
Length = 2257
Score = 25.4 bits (54), Expect = 9.2
Identities = 11/28 (39%), Positives = 14/28 (49%)
Frame = -1
Query: 197 DYTVNETEVTEINAENSSRITRKILKHD 224
+Y E + NSSR+T K KHD
Sbjct: 1780 EYASYEHTSCSFHQRNSSRVTSKFFKHD 1697
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,490,962
Number of Sequences: 28460
Number of extensions: 61249
Number of successful extensions: 280
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 279
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 279
length of query: 242
length of database: 4,897,600
effective HSP length: 88
effective length of query: 154
effective length of database: 2,393,120
effective search space: 368540480
effective search space used: 368540480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0344.6


