

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
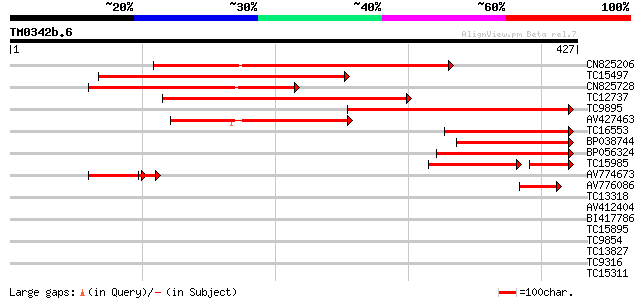
Query= TM0342b.6
(427 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CN825206 260 3e-70
TC15497 similar to UP|Q41695 (Q41695) Pectinacetylesterase precu... 213 6e-56
CN825728 207 3e-54
TC12737 similar to UP|Q41695 (Q41695) Pectinacetylesterase precu... 186 6e-48
TC9895 similar to UP|Q41695 (Q41695) Pectinacetylesterase precur... 179 6e-46
AV427463 169 8e-43
TC16553 weakly similar to UP|Q41695 (Q41695) Pectinacetylesteras... 109 1e-24
BP038744 107 3e-24
BP056324 106 8e-24
TC15985 weakly similar to UP|Q9M1R8 (Q9M1R8) Pectinacetylesteras... 72 2e-22
AV774673 49 5e-09
AV776086 46 1e-05
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 27 0.28
AV412404 31 0.45
BI417786 25 0.46
TC15895 26 1.2
TC9854 weakly similar to UP|Q9LSN7 (Q9LSN7) Gb|AAC24081.1 (AT3g1... 29 1.3
TC13827 29 1.7
TC9316 similar to UP|Q94F86 (Q94F86) Early light inducible prote... 29 1.7
TC15311 similar to UP|Q9LL85 (Q9LL85) DNA-binding protein p24, p... 29 1.7
>CN825206
Length = 679
Score = 260 bits (665), Expect = 3e-70
Identities = 118/226 (52%), Positives = 162/226 (71%)
Frame = +3
Query: 109 SISSCSRRKFTALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPE 168
++ +C RK T GSSKYME + F+GILS+ +NPDF+NWN+V +RYCDGASF+G E
Sbjct: 3 TVRNCVYRKRTRRGSSKYMENQIQFTGILSNKAEENPDFYNWNRVKLRYCDGASFSGDSE 182
Query: 169 SEKGSGLFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKET 228
+E L FRGQ IW A ++EL+S GM +A QALLSGCSAGGLA+++HCD+F+ + P T
Sbjct: 183 NESAE-LQFRGQKIWLAAVEELMSKGMQEANQALLSGCSAGGLASIIHCDEFQSLFPNST 359
Query: 229 TVKCLADAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEIL 288
VKCL+DAGFF+D D+ G T+RS + VVQLQ V K+L K CL + + + C FP I+
Sbjct: 360 KVKCLSDAGFFIDAADVSGGRTLRSLFEGVVQLQEVQKNLPKSCLNQQDATSCFFPQNIV 539
Query: 289 KNIKTPVFLVHSAYDFWQIRNILVPEGSDPSGHWKSCKLNIHNCNA 334
++++TP+FL+++AYD WQ+ L P +DP G W C+ N NCN+
Sbjct: 540 QHLETPLFLLNAAYDAWQVLASLAPPSADPLGFWSDCRSNHANCNS 677
>TC15497 similar to UP|Q41695 (Q41695) Pectinacetylesterase precursor ,
partial (55%)
Length = 799
Score = 213 bits (541), Expect = 6e-56
Identities = 91/190 (47%), Positives = 136/190 (70%), Gaps = 1/190 (0%)
Frame = +2
Query: 68 NAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFTALGSSKYM 127
+A+ +GA+CLDGS P +HF KGFG+G +W++H EGG WCN++++C R+ T LGSSK M
Sbjct: 230 SAVAKGAVCLDGSPPAYHFHKGFGAGINNWIVHFEGGAWCNNVTTCLARRDTRLGSSKKM 409
Query: 128 ETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPES-EKGSGLFFRGQIIWEAI 186
+ FSG S+ + NPDF+NWN++ +RYCDG+SF G E+ + + L FRG I+ A+
Sbjct: 410 SQTLSFSGFFSNGQKFNPDFYNWNRIKVRYCDGSSFTGDVEAVDPKTNLHFRGGRIFVAV 589
Query: 187 MDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGFFLDEKDIR 246
+++LL+ GM A+ A+LSGCSAGGL +++ CD FR ++P VKCL+DAG+F++ KD+
Sbjct: 590 VEDLLANGMKNAQNAILSGCSAGGLTSILQCDRFRSLIPAAAKVKCLSDAGYFINLKDVS 769
Query: 247 GNNTMRSFYN 256
G + Y+
Sbjct: 770 GAAHIEQLYS 799
>CN825728
Length = 711
Score = 207 bits (527), Expect = 3e-54
Identities = 94/159 (59%), Positives = 122/159 (76%)
Frame = +3
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISSCSRRKFT 119
++P T + A +GA+CLDG+ PG+HF GFGSG+ SWL+ LEGGGWCN+I +C RK T
Sbjct: 237 MVPLTLIQGAASKGAVCLDGTLPGYHFHPGFGSGANSWLVELEGGGWCNTIRNCVYRKTT 416
Query: 120 ALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGSGLFFRG 179
GSSKYME +PF+GILS+ NPDFFNWN+V +RYCDGASF+G ++E + L FRG
Sbjct: 417 RRGSSKYMEKQLPFTGILSNKAEYNPDFFNWNRVRVRYCDGASFSGDSQNE-AAQLQFRG 593
Query: 180 QIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCD 218
Q I+ A M+EL+S GM A QALL+GCSAGG+A+++HCD
Sbjct: 594 QKIYLAAMEELMSKGMKNANQALLTGCSAGGVASIIHCD 710
>TC12737 similar to UP|Q41695 (Q41695) Pectinacetylesterase precursor ,
partial (49%)
Length = 616
Score = 186 bits (472), Expect = 6e-48
Identities = 83/188 (44%), Positives = 130/188 (69%), Gaps = 1/188 (0%)
Frame = +1
Query: 116 RKFTALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKG-SG 174
RK LGSSK M + FSGIL + + NPDF+NWN++ +RYCDG+SF G E+ +
Sbjct: 52 RKNNRLGSSKKMANQIAFSGILHNRKQFNPDFYNWNRIKVRYCDGSSFTGDVEAVNPVTK 231
Query: 175 LFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLA 234
L FRG I+ A++++LL+ GM A+ A++SGCSAGGL +++HCD FR +LP+ VKCL+
Sbjct: 232 LHFRGARIFAAVIEDLLAKGMKNARNAIISGCSAGGLTSVLHCDRFRALLPRGAKVKCLS 411
Query: 235 DAGFFLDEKDIRGNNTMRSFYNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTP 294
DAG+F++ +D+ G++ ++ ++ VV G A++L + C +++ P C FP ++ I TP
Sbjct: 412 DAGYFINARDVSGSSYIKQYFTQVVITHGSARNLPQSCTSRISPGLCFFPQYLVSQITTP 591
Query: 295 VFLVHSAY 302
+F V++AY
Sbjct: 592 IFFVNAAY 615
Score = 31.2 bits (69), Expect = 0.34
Identities = 12/32 (37%), Positives = 19/32 (58%)
Frame = +2
Query: 102 EGGGWCNSISSCSRRKFTALGSSKYMETVVPF 133
EGGGWCN++++C + T + T +PF
Sbjct: 11 EGGGWCNNVTTCLVARTTV*VHPRKWPTKLPF 106
>TC9895 similar to UP|Q41695 (Q41695) Pectinacetylesterase precursor ,
partial (43%)
Length = 713
Score = 179 bits (455), Expect = 6e-46
Identities = 75/170 (44%), Positives = 110/170 (64%)
Frame = +2
Query: 255 YNDVVQLQGVAKSLHKECLAKMEPSKCLFPSEILKNIKTPVFLVHSAYDFWQIRNILVPE 314
Y+ VV+ G AK+L C +++ P C FP + IKTP+F V++AYD +Q++NIL P
Sbjct: 5 YSQVVETHGSAKNLPASCTSRLRPGLCFFPQNVAGQIKTPIFFVNAAYDSYQVKNILAPG 184
Query: 315 GSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTW 374
+DP G W+ CKL+I C++N + ++ FR+ LKA++ + GMF++ C+ HCQT
Sbjct: 185 VADPHGTWRDCKLDIKKCSSNQLTVMQGFRTEFLKAISVVENSPSKGMFIDGCYSHCQTG 364
Query: 375 RGETWYSPNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDF 424
ETW +SP + N TIA++V DWF++R + IDC YPCNPTC N F
Sbjct: 365 MQETWMRSDSPVLANTTIAKAVGDWFYERRSFSQIDCSYPCNPTCHNRVF 514
>AV427463
Length = 417
Score = 169 bits (428), Expect = 8e-43
Identities = 81/140 (57%), Positives = 103/140 (72%), Gaps = 3/140 (2%)
Frame = +2
Query: 122 GSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAG---HPESEKGSGLFFR 178
GSS +ME +PF+GILS+ +NPDFFNWN+V +RYCDGASFAG HP ++ L FR
Sbjct: 8 GSSAFMEKEIPFTGILSNKAEENPDFFNWNRVKLRYCDGASFAGDAAHPTAQ----LQFR 175
Query: 179 GQIIWEAIMDELLSIGMSKAKQALLSGCSAGGLATLVHCDDFRQILPKETTVKCLADAGF 238
GQ IW A M++L+S GM A QALLSGCSAGGLAT++HCD+FR P+ VKCL+DAG
Sbjct: 176 GQRIWAAAMEDLMSKGMRFANQALLSGCSAGGLATIIHCDEFRGYFPRTAKVKCLSDAGL 355
Query: 239 FLDEKDIRGNNTMRSFYNDV 258
FLD D+ G ++R+ Y+ V
Sbjct: 356 FLDAIDVSGGRSLRNLYSGV 415
>TC16553 weakly similar to UP|Q41695 (Q41695) Pectinacetylesterase precursor
, partial (24%)
Length = 571
Score = 109 bits (272), Expect = 1e-24
Identities = 44/97 (45%), Positives = 66/97 (67%)
Frame = +2
Query: 328 NIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKI 387
+I+NC+ + +D++ FR+ L+AL GMF++SC+ HC T ETW++ +SP +
Sbjct: 2 DINNCSPDPLDLMQGFRTEFLRALTVLGNSPSKGMFIDSCYAHCPTEMQETWFTSDSPVV 181
Query: 388 QNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDF 424
TIA++VADWF++R + + IDCPYPCNPTC N F
Sbjct: 182 NKTTIAKAVADWFYERRLFHQIDCPYPCNPTCHNRVF 292
>BP038744
Length = 531
Score = 107 bits (268), Expect = 3e-24
Identities = 44/88 (50%), Positives = 60/88 (68%)
Frame = -2
Query: 337 IDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYSPNSPKIQNKTIAESV 396
+ L FR+ +L + +F + + G+F+NSCF HCQ+ R +TW+S NSP I NK IA +V
Sbjct: 521 VQFLQGFRNRMLNVIKDFSRSNRNGLFINSCFAHCQSERQDTWFSDNSPVIGNKAIAVAV 342
Query: 397 ADWFFDREVINLIDCPYPCNPTCRNLDF 424
DW+FDR + IDCPYPC+ TC NL F
Sbjct: 341 GDWYFDRAGVKAIDCPYPCDKTCHNLIF 258
>BP056324
Length = 421
Score = 106 bits (264), Expect = 8e-24
Identities = 47/103 (45%), Positives = 64/103 (61%)
Frame = -2
Query: 322 WKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWRGETWYS 381
W C+ N C + I L FRS +L+ F + ++ G+F+NSCF C + R +TW++
Sbjct: 402 WMECRKNYARCRSPPIPYLPGFRSPMLRVTRGFSRARKNGLFINSCFSPCPSERPDTWFA 223
Query: 382 PNSPKIQNKTIAESVADWFFDREVINLIDCPYPCNPTCRNLDF 424
+SP+I NK IAESV DWFFDR + I C YPC+ TC NL F
Sbjct: 222 RHSPRIGNKGIAESVGDWFFDRSSVPAIGCAYPCDQTCPNLVF 94
>TC15985 weakly similar to UP|Q9M1R8 (Q9M1R8) Pectinacetylesterase
precursor-like protein, partial (22%)
Length = 763
Score = 72.0 bits (175), Expect(2) = 2e-22
Identities = 27/70 (38%), Positives = 45/70 (63%)
Frame = +3
Query: 316 SDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCFIHCQTWR 375
+DP G W +CKLN CN++ I FR++++ + +F + G+F+NSCF HC + R
Sbjct: 9 ADPHGSWTACKLNHAKCNSSQIQFFQDFRNAMVNDVRDFSRTPPTGLFINSCFSHCPSER 188
Query: 376 GETWYSPNSP 385
+TW++ +SP
Sbjct: 189 QDTWFADDSP 218
Score = 50.4 bits (119), Expect(2) = 2e-22
Identities = 18/33 (54%), Positives = 25/33 (75%)
Frame = +1
Query: 392 IAESVADWFFDREVINLIDCPYPCNPTCRNLDF 424
+A +V +W+FDR+ + IDCPYPC+ TC NL F
Sbjct: 238 VAVAVGNWYFDRQAVKEIDCPYPCDNTCHNLVF 336
>AV774673
Length = 208
Score = 48.9 bits (115), Expect(2) = 5e-09
Identities = 19/44 (43%), Positives = 28/44 (63%)
Frame = +3
Query: 60 LIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEG 103
L P T + +A+ GA+ +DG P +H KGFG+G +WL+H G
Sbjct: 42 LYPIT*VQSAVANGAVSIDGRPPAYHNHKGFGAGINNWLVHFAG 173
Score = 28.1 bits (61), Expect(2) = 5e-09
Identities = 8/16 (50%), Positives = 13/16 (81%)
Frame = +1
Query: 98 LLHLEGGGWCNSISSC 113
L L+GG WCN++++C
Sbjct: 157 LFTLQGGAWCNNVTTC 204
>AV776086
Length = 366
Score = 45.8 bits (107), Expect = 1e-05
Identities = 17/31 (54%), Positives = 24/31 (76%)
Frame = -1
Query: 385 PKIQNKTIAESVADWFFDREVINLIDCPYPC 415
P + TIA++VADWF+ R++ + IDCPYPC
Sbjct: 366 PLVYKTTIAKAVADWFYVRKLFHQIDCPYPC 274
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 26.9 bits (58), Expect(2) = 0.28
Identities = 12/21 (57%), Positives = 12/21 (57%)
Frame = -1
Query: 37 FLLLSLAFFSQLHHHHHSHSL 57
FL L L F L HHHH H L
Sbjct: 402 FLRLLLLHFPLLPHHHHHHLL 340
Score = 23.9 bits (50), Expect(2) = 2.6
Identities = 7/9 (77%), Positives = 7/9 (77%)
Frame = -1
Query: 49 HHHHHSHSL 57
HHHHH H L
Sbjct: 321 HHHHHHHHL 295
Score = 23.1 bits (48), Expect(2) = 0.28
Identities = 9/23 (39%), Positives = 12/23 (52%)
Frame = -1
Query: 49 HHHHHSHSLSNLIPFTPLPNAIL 71
HHHHH H + PL + +L
Sbjct: 327 HHHHHHHHHHLHLHHLPLHHHLL 259
Score = 22.7 bits (47), Expect(2) = 2.6
Identities = 10/18 (55%), Positives = 10/18 (55%)
Frame = -1
Query: 36 FFLLLSLAFFSQLHHHHH 53
F LL L F HHHHH
Sbjct: 402 FLRLLLLHFPLLPHHHHH 349
>AV412404
Length = 386
Score = 30.8 bits (68), Expect = 0.45
Identities = 16/33 (48%), Positives = 20/33 (60%), Gaps = 2/33 (6%)
Frame = +1
Query: 35 AFFLL--LSLAFFSQLHHHHHSHSLSNLIPFTP 65
+FF+L L L+F SQ+HHHH L PF P
Sbjct: 61 SFFILFFLFLSFHSQIHHHHPFPPLLPPQPFNP 159
>BI417786
Length = 513
Score = 24.6 bits (52), Expect(2) = 0.46
Identities = 8/12 (66%), Positives = 9/12 (74%)
Frame = +2
Query: 49 HHHHHSHSLSNL 60
HHHHH H L +L
Sbjct: 134 HHHHHHHVLLHL 169
Score = 24.6 bits (52), Expect(2) = 0.46
Identities = 9/20 (45%), Positives = 10/20 (50%)
Frame = +2
Query: 36 FFLLLSLAFFSQLHHHHHSH 55
F L L + HHHHH H
Sbjct: 74 FCTLSHLTSYHPSHHHHHHH 133
>TC15895
Length = 622
Score = 25.8 bits (55), Expect(2) = 1.2
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = -3
Query: 23 KLTTKDYAIAAFAFFLLLSLAFFSQLHHHHHSH 55
++ KD A F L+ L+ S LHHHH+ H
Sbjct: 410 EIKQKDIFALAHTVFTLIQLSS-SILHHHHNLH 315
Score = 21.9 bits (45), Expect(2) = 1.2
Identities = 7/13 (53%), Positives = 8/13 (60%)
Frame = -1
Query: 50 HHHHSHSLSNLIP 62
HHHH H +L P
Sbjct: 211 HHHHHHQPGSLPP 173
>TC9854 weakly similar to UP|Q9LSN7 (Q9LSN7) Gb|AAC24081.1
(AT3g17100/K14A17_22_), partial (43%)
Length = 1108
Score = 29.3 bits (64), Expect = 1.3
Identities = 11/16 (68%), Positives = 11/16 (68%)
Frame = +2
Query: 49 HHHHHSHSLSNLIPFT 64
HHHHH H S LIP T
Sbjct: 23 HHHHHHHHPSPLIPVT 70
>TC13827
Length = 517
Score = 28.9 bits (63), Expect = 1.7
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +2
Query: 49 HHHHHSHSLSNLIPFT 64
HHHHH L NL+P+T
Sbjct: 158 HHHHHVLLLRNLLPYT 205
>TC9316 similar to UP|Q94F86 (Q94F86) Early light inducible protein,
partial (84%)
Length = 831
Score = 28.9 bits (63), Expect = 1.7
Identities = 13/18 (72%), Positives = 14/18 (77%)
Frame = +2
Query: 38 LLLSLAFFSQLHHHHHSH 55
LLL+LA QLHHHHH H
Sbjct: 281 LLLALA---QLHHHHHHH 325
>TC15311 similar to UP|Q9LL85 (Q9LL85) DNA-binding protein p24, partial
(69%)
Length = 966
Score = 28.9 bits (63), Expect = 1.7
Identities = 21/67 (31%), Positives = 27/67 (39%)
Frame = +2
Query: 53 HSHSLSNLIPFTPLPNAILRGAICLDGSAPGFHFQKGFGSGSRSWLLHLEGGGWCNSISS 112
HSH + L P P +L C P F F + W LHL+ G +S S
Sbjct: 191 HSHQMPPLRPVRPKNLLLLHSTTC----KPRCCFSWSFAT*GLCWSLHLQREGCSHSHS* 358
Query: 113 CSRRKFT 119
SR + T
Sbjct: 359 TSRIRTT 379
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.138 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,066,583
Number of Sequences: 28460
Number of extensions: 175892
Number of successful extensions: 3338
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 2337
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2841
length of query: 427
length of database: 4,897,600
effective HSP length: 93
effective length of query: 334
effective length of database: 2,250,820
effective search space: 751773880
effective search space used: 751773880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0342b.6


