

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.9
(434 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
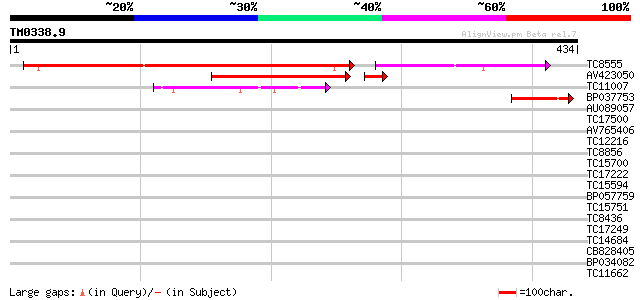
Score E
Sequences producing significant alignments: (bits) Value
TC8555 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin ... 409 e-131
AV423050 158 1e-39
TC11007 similar to GB|AAO64838.1|29028918|BT005903 At4g16120 {Ar... 74 6e-14
BP037753 57 8e-09
AU089057 37 0.008
TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase t... 32 0.20
AV765406 30 0.78
TC12216 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Ar... 29 1.3
TC8856 similar to UP|Q8W403 (Q8W403) Sec13p, partial (48%) 28 2.3
TC15700 27 5.0
TC17222 27 5.0
TC15594 similar to UP|CYTI_VIGUN (Q06445) Cysteine proteinase in... 27 6.6
BP057759 27 6.6
TC15751 similar to UP|Q8S4V8 (Q8S4V8) Holocarboxylase synthetase... 27 6.6
TC8436 homologue to UP|PSAD_LYCES (P12372) Photosystem I reactio... 27 6.6
TC17249 similar to UP|Q84LK2 (Q84LK2) Granule-bound starch synth... 27 8.6
TC14684 27 8.6
CB828405 27 8.6
BP034082 27 8.6
TC11662 similar to UP|O65748 (O65748) Acyl-CoA synthetase (Frag... 27 8.6
>TC8555 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (83%)
Length = 1776
Score = 409 bits (1052), Expect(2) = e-131
Identities = 183/262 (69%), Positives = 223/262 (84%), Gaps = 8/262 (3%)
Frame = +1
Query: 11 LPCTLLLLLP----SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQE 66
LP LLL+L +S+EAYD LDP GNITI+WD++SWT DGYVA VT+ NFQQYRHIQ
Sbjct: 118 LPFLLLLVLSCSTFTSTEAYDALDPTGNITIKWDVISWTPDGYVATVTMYNFQQYRHIQA 297
Query: 67 PGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQV 126
PGWSLGWTWAK E+IW M+GGQTTEQGDCSKFK + PHCCKKDP VDL+PG PY++QV
Sbjct: 298 PGWSLGWTWAKKEVIWNMMGGQTTEQGDCSKFKAGI-PHCCKKDPTVVDLLPGTPYNQQV 474
Query: 127 ANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIV 186
ANCCKGGVL S QDP NAV+SF++ VG AGTT K+V+LPKNFT K+PGPGYTCGPA+ V
Sbjct: 475 ANCCKGGVLNSWAQDPGNAVSSFQISVGSAGTTNKTVKLPKNFTLKAPGPGYTCGPAKNV 654
Query: 187 KPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ 246
+PT+++ DKRR+TQA+M+WNVTCTYSQFLA K P+CCVSLS+FYND+IV+CPTC CGC+
Sbjct: 655 RPTKYITSDKRRITQALMTWNVTCTYSQFLAQKTPSCCVSLSSFYNDSIVNCPTCTCGCK 834
Query: 247 S----GNCVDPNSPNLTSVISN 264
+ G+CV+P+SP+L+SV+S+
Sbjct: 835 NKTDPGSCVNPDSPHLSSVVSS 900
Score = 75.5 bits (184), Expect(2) = e-131
Identities = 57/136 (41%), Positives = 78/136 (56%), Gaps = 2/136 (1%)
Frame = +3
Query: 281 SSLACEAKLQAVLASEGDDY*FQLQDELF*LELSNRTPKLRQSDPAL*FQLRVINSLWGV 340
S+LA EA L VL + Y FQLQ+EL +E P+ S ++V N+L G+
Sbjct: 963 SALAREA*L*RVLEGQDHYYKFQLQNELLTMEPCCAAPQF**SYRDFQLSIQVTNTL*GL 1142
Query: 341 NE*YSNALGN*VLQ*FAHASW--PSWLCTVRPAFWKG*VNFYFS*RLGFSSEDLFQW*QL 398
+*Y A+GN VLQ*F+ S+ SW C VR +G +N +F * +G +DLFQ L
Sbjct: 1143 -K*YRYAVGNEVLQ*FSFFSF*KKSW*CAVRNLI*EGQINLHF**GMGLPKKDLFQRR*L 1319
Query: 399 CDATT*CLSKVTECRF 414
C A+T CLS T+C+F
Sbjct: 1320 CYASTRCLSLATKCKF 1367
>AV423050
Length = 424
Score = 158 bits (399), Expect(2) = 1e-39
Identities = 72/108 (66%), Positives = 88/108 (80%), Gaps = 1/108 (0%)
Frame = +3
Query: 155 RAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQ 214
+AGT+ K+V+LPKNFT +PGPGYTCGPA++V T FL DKRR TQA+M+WNVTCTYSQ
Sbjct: 3 QAGTSNKTVKLPKNFTLLAPGPGYTCGPAKVVPSTNFLTPDKRRKTQALMTWNVTCTYSQ 182
Query: 215 FLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS-GNCVDPNSPNLTSV 261
FLA K P+CCVSLS+FYN+TI CP+CACGCQ+ NCV +S L+ V
Sbjct: 183 FLARKNPSCCVSLSSFYNETITPCPSCACGCQNKKNCVKRDSKILSMV 326
Score = 22.3 bits (46), Expect(2) = 1e-39
Identities = 8/18 (44%), Positives = 12/18 (66%)
Frame = +2
Query: 272 MH*PFMPNPSSLACEAKL 289
+H ++PN LACE +L
Sbjct: 368 VHSSYVPNQGPLACEGQL 421
>TC11007 similar to GB|AAO64838.1|29028918|BT005903 At4g16120 {Arabidopsis
thaliana;}, partial (46%)
Length = 732
Score = 73.6 bits (179), Expect = 6e-14
Identities = 45/145 (31%), Positives = 68/145 (46%), Gaps = 10/145 (6%)
Frame = +2
Query: 111 PVAVDLIPGAPYSE----QVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLP 166
P VDL P +++ ++ +CC+ G + DP+ +V+ F++ V + +L
Sbjct: 5 PTIVDL-PPTKFNDTVLGKIPSCCRNGTILPPSMDPSMSVSRFQMEVFKMPPALNRSQLS 181
Query: 167 KNFTFKSPG---PGYTCGPARIVKPTRFLRYDKRRVTQ---AMMSWNVTCTYSQFLAAKA 220
+K G P Y CGP V PT D V M SW V C + +
Sbjct: 182 PPQNWKISGTLNPDYKCGPPMRVSPTE--NPDPSGVPSNKTVMASWQVVCNITTTKRTSS 355
Query: 221 PTCCVSLSTFYNDTIVSCPTCACGC 245
CCVS S +YN++++ C TCACGC
Sbjct: 356 K-CCVSFSAYYNESVIPCKTCACGC 427
>BP037753
Length = 539
Score = 56.6 bits (135), Expect = 8e-09
Identities = 29/47 (61%), Positives = 33/47 (69%)
Frame = -1
Query: 385 LGFSSEDLFQW*QLCDATT*CLSKVTECRFAAKGFPLTCFDVNILGN 431
LGFSSEDL QW*QLC A CLS VT C F+A+GF L CF + G+
Sbjct: 527 LGFSSEDLLQW*QLCYALARCLSMVT*CWFSARGF-LVCFSDGLFGS 390
>AU089057
Length = 154
Score = 36.6 bits (83), Expect = 0.008
Identities = 15/20 (75%), Positives = 16/20 (80%)
Frame = +2
Query: 351 *VLQ*FAHASWPSWLCTVRP 370
*VLQ F HASWPSWLC+ P
Sbjct: 95 *VLQRFTHASWPSWLCSFPP 154
>TC17500 UP|Q9ZPL9 (Q9ZPL9) Nodule-enhanced protein phosphatase type 2C,
complete
Length = 1325
Score = 32.0 bits (71), Expect = 0.20
Identities = 19/69 (27%), Positives = 29/69 (41%), Gaps = 9/69 (13%)
Frame = -1
Query: 199 VTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDT-IVSCPTCACGCQSGNC------- 250
+ Q +SW++T Y L ++ C YNDT ++ C C +C
Sbjct: 1265 IIQHNLSWHLTRCYMSLLIMRSQLCYSPSQLNYNDTGVIPASLCQCYLSQKSCSLRSTTL 1086
Query: 251 -VDPNSPNL 258
+ NSPNL
Sbjct: 1085 ILITNSPNL 1059
>AV765406
Length = 323
Score = 30.0 bits (66), Expect = 0.78
Identities = 14/30 (46%), Positives = 19/30 (62%)
Frame = -3
Query: 385 LGFSSEDLFQW*QLCDATT*CLSKVTECRF 414
+G +DLFQ LC A+ CLS T+C+F
Sbjct: 318 MGLPKKDLFQRR*LCYASPKCLSLATKCKF 229
>TC12216 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;}, partial (30%)
Length = 570
Score = 29.3 bits (64), Expect = 1.3
Identities = 18/64 (28%), Positives = 25/64 (38%)
Frame = +2
Query: 77 KNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLT 136
+NE +W + G + DCSK+ C + D CK GVL
Sbjct: 47 QNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCD------------------SCKAGVLA 172
Query: 137 SLKQ 140
SLK+
Sbjct: 173 SLKK 184
>TC8856 similar to UP|Q8W403 (Q8W403) Sec13p, partial (48%)
Length = 568
Score = 28.5 bits (62), Expect = 2.3
Identities = 16/45 (35%), Positives = 21/45 (46%), Gaps = 1/45 (2%)
Frame = -1
Query: 243 CGCQSGNCVDPNSPNLTSVISNP-GNNGHYMH*PFMPNPSSLACE 286
C Q P P L S SNP ++G Y+ PF P+ + CE
Sbjct: 241 CSPQLS*LCGPKKP*LASYHSNPLQHHGQYLGDPFQPSEQACFCE 107
>TC15700
Length = 549
Score = 27.3 bits (59), Expect = 5.0
Identities = 12/35 (34%), Positives = 19/35 (54%)
Frame = +2
Query: 74 TWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCK 108
+W +EI W IG + E+ S KG++ CC+
Sbjct: 269 SWTNSEIFWDSIGERRGERFVNSGLKGEI---CCR 364
>TC17222
Length = 534
Score = 27.3 bits (59), Expect = 5.0
Identities = 16/52 (30%), Positives = 20/52 (37%)
Frame = -3
Query: 200 TQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQSGNCV 251
T ++ WN +C LA A L D IVS C C +CV
Sbjct: 331 THSVGMWNASCKALNCLAITAAISSSHLILPSKDEIVS*SVCRTSCTMQSCV 176
>TC15594 similar to UP|CYTI_VIGUN (Q06445) Cysteine proteinase inhibitor
(Cystatin), complete
Length = 703
Score = 26.9 bits (58), Expect = 6.6
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = -2
Query: 220 APTCCVSLSTFYNDTIVSC 238
A TCCV+L+TF N V C
Sbjct: 318 ATTCCVALTTFPNSRSVFC 262
>BP057759
Length = 563
Score = 26.9 bits (58), Expect = 6.6
Identities = 12/49 (24%), Positives = 19/49 (38%), Gaps = 3/49 (6%)
Frame = -3
Query: 73 WTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSP---HCCKKDPVAVDLIP 118
W+W + + W+ + G + K SP HCC+ V P
Sbjct: 240 WSWERKILAWERVCGLSRRSRRDRKAAYAASPDCLHCCRNSSTVVGEAP 94
>TC15751 similar to UP|Q8S4V8 (Q8S4V8) Holocarboxylase synthetase 1, partial
(80%)
Length = 1470
Score = 26.9 bits (58), Expect = 6.6
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = -3
Query: 222 TCCVSLSTFYNDTIVSCPTC 241
TC ++LS+ ++ TI CP C
Sbjct: 997 TCSITLSSLFSCTITLCPLC 938
>TC8436 homologue to UP|PSAD_LYCES (P12372) Photosystem I reaction center
subunit II, chloroplast precursor (Photosystem I 20 kDa
subunit) (PSI-D), partial (77%)
Length = 777
Score = 26.9 bits (58), Expect = 6.6
Identities = 11/41 (26%), Positives = 17/41 (40%)
Frame = -2
Query: 210 CTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQSGNC 250
C + +F + CC + ++SC C C C G C
Sbjct: 263 CVWVKFWWGEPNGCCFLCCYWCLCFLISCSDCCCCCSWGCC 141
>TC17249 similar to UP|Q84LK2 (Q84LK2) Granule-bound starch synthase Ib
precursor , partial (20%)
Length = 610
Score = 26.6 bits (57), Expect = 8.6
Identities = 8/14 (57%), Positives = 10/14 (71%)
Frame = +1
Query: 237 SCPTCACGCQSGNC 250
SC +C CGC S +C
Sbjct: 124 SCGSCGCGCSSKDC 165
>TC14684
Length = 757
Score = 26.6 bits (57), Expect = 8.6
Identities = 9/27 (33%), Positives = 15/27 (55%)
Frame = -2
Query: 56 NNFQQYRHIQEPGWSLGWTWAKNEIIW 82
N+ Q + ++ W +GWT K +I W
Sbjct: 102 NHHQDPQSQEKEAWIVGWTMNKRKIFW 22
>CB828405
Length = 552
Score = 26.6 bits (57), Expect = 8.6
Identities = 17/53 (32%), Positives = 24/53 (45%), Gaps = 4/53 (7%)
Frame = -1
Query: 19 LPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAV----TLNNFQQYRHIQEP 67
LPSSS ++ L N T+ WD L T G+ TL F+ ++ P
Sbjct: 321 LPSSSFSF*TLGSRANSTV*WDSLPTTHIGHCGDFRN*PTLQKFRDSSKVRNP 163
>BP034082
Length = 412
Score = 26.6 bits (57), Expect = 8.6
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = +2
Query: 10 VLPCTLLLLLPSSSEAYDPLDP 31
V+PC L+++ SSSEA +DP
Sbjct: 227 VIPCLLVVVYDSSSEASSRVDP 292
>TC11662 similar to UP|O65748 (O65748) Acyl-CoA synthetase (Fragment) ,
partial (81%)
Length = 640
Score = 26.6 bits (57), Expect = 8.6
Identities = 13/32 (40%), Positives = 16/32 (49%)
Frame = -2
Query: 234 TIVSCPTCACGCQSGNCVDPNSPNLTSVISNP 265
TI+SCP C C C + N L +V NP
Sbjct: 138 TILSCPMCK--CLLSGCNNSNKEGLETVSINP 49
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.334 0.145 0.504
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,595,881
Number of Sequences: 28460
Number of extensions: 160765
Number of successful extensions: 1314
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 1297
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1308
length of query: 434
length of database: 4,897,600
effective HSP length: 93
effective length of query: 341
effective length of database: 2,250,820
effective search space: 767529620
effective search space used: 767529620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0338.9


