

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.8
(61 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
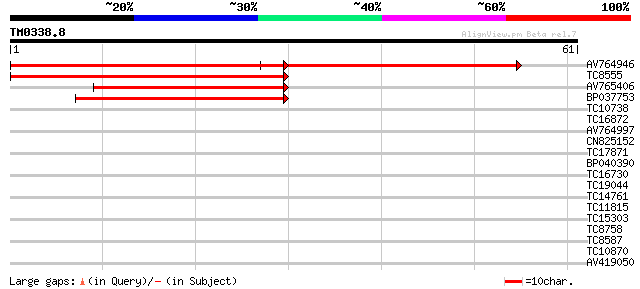
AV764946 73 3e-24
TC8555 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin ... 61 5e-11
AV765406 48 4e-07
BP037753 47 6e-07
TC10738 homologue to GB|AAO64838.1|29028918|BT005903 At4g16120 {... 33 0.014
TC16872 similar to UP|O64896 (O64896) Trehalose-6-phosphate phos... 26 1.3
AV764997 25 2.3
CN825152 25 2.3
TC17871 similar to UP|Q7UMK2 (Q7UMK2) Probable transcription ter... 24 5.1
BP040390 24 5.1
TC16730 similar to GB|AAO64817.1|29028876|BT005882 At3g04570 {Ar... 24 5.1
TC19044 similar to UP|Q9LJH9 (Q9LJH9) Gb|AAD25781.1, partial (16%) 24 6.7
TC14761 similar to UP|AAS07369 (AAS07369) Expressed protein, par... 24 6.7
TC11815 weakly similar to UP|Q8MLU0 (Q8MLU0) CG13503-PA, partial... 24 6.7
TC15303 similar to UP|Q9ZNZ5 (Q9ZNZ5) Peroxidase precursor (Fra... 23 8.7
TC8758 similar to GB|AAC77862.2|20197419|AC005623 Argonaute (AGO... 23 8.7
TC8587 similar to UP|Q84MD8 (Q84MD8) At4g21470, partial (97%) 23 8.7
TC10870 similar to GB|CAA77438.1|2924285|CHNTXX Ycf2 protein {Ni... 23 8.7
AV419050 23 8.7
>AV764946
Length = 421
Score = 72.8 bits (177), Expect(2) = 3e-24
Identities = 30/30 (100%), Positives = 30/30 (100%)
Frame = -2
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDKNTFSFKQGWAFPRKVYFNGDECMMPPP
Sbjct: 384 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 295
Score = 52.8 bits (125), Expect(2) = 3e-24
Identities = 25/28 (89%), Positives = 25/28 (89%)
Frame = -3
Query: 28 PPPTPTHFCQILLLQA*FPSRHSSSHYS 55
P TPTHFCQILLLQA*FPSRHSSSH S
Sbjct: 302 PHLTPTHFCQILLLQA*FPSRHSSSHCS 219
>TC8555 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} , partial
(83%)
Length = 1776
Score = 60.8 bits (146), Expect = 5e-11
Identities = 22/30 (73%), Positives = 28/30 (93%)
Frame = +1
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPP 30
KDK+TF+F +GWAFPR++YFNGD C+MPPP
Sbjct: 1246 KDKSTFTFDKGWAFPRRIYFNGDNCVMPPP 1335
>AV765406
Length = 323
Score = 47.8 bits (112), Expect = 4e-07
Identities = 16/21 (76%), Positives = 20/21 (95%)
Frame = -1
Query: 10 QGWAFPRKVYFNGDECMMPPP 30
+GWAFPR++YFNGD C+MPPP
Sbjct: 323 KGWAFPRRIYFNGDNCVMPPP 261
>BP037753
Length = 539
Score = 47.4 bits (111), Expect = 6e-07
Identities = 16/23 (69%), Positives = 21/23 (90%)
Frame = -2
Query: 8 FKQGWAFPRKVYFNGDECMMPPP 30
F++GWAFPR++YFNGD C+MP P
Sbjct: 538 FEKGWAFPRRIYFNGDNCVMPSP 470
>TC10738 homologue to GB|AAO64838.1|29028918|BT005903 At4g16120
{Arabidopsis thaliana;}, partial (5%)
Length = 596
Score = 32.7 bits (73), Expect = 0.014
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = +3
Query: 10 QGWAFPRKVYFNGDECMMPPPTPT 33
+G FP KV+FNG+EC +P P+
Sbjct: 21 RGDGFPSKVFFNGEECSLPSVIPS 92
>TC16872 similar to UP|O64896 (O64896) Trehalose-6-phosphate phosphatase ,
partial (15%)
Length = 670
Score = 26.2 bits (56), Expect = 1.3
Identities = 11/29 (37%), Positives = 16/29 (54%)
Frame = +2
Query: 25 CMMPPPTPTHFCQILLLQA*FPSRHSSSH 53
C +PP THF ++LQ + S + S H
Sbjct: 86 CPLPPRKATHFTLFMILQRLWNSSNHSRH 172
>AV764997
Length = 307
Score = 25.4 bits (54), Expect = 2.3
Identities = 9/13 (69%), Positives = 10/13 (76%)
Frame = -2
Query: 24 ECMMPPPTPTHFC 36
+C MPPP P HFC
Sbjct: 57 DCKMPPP-PVHFC 22
>CN825152
Length = 697
Score = 25.4 bits (54), Expect = 2.3
Identities = 14/27 (51%), Positives = 17/27 (62%), Gaps = 1/27 (3%)
Frame = +1
Query: 28 PPPTPTHFCQILLLQ-A*FPSRHSSSH 53
PPP PT +LLL A FPS+ + SH
Sbjct: 109 PPPQPTSSTLLLLLPLAMFPSQLNPSH 189
>TC17871 similar to UP|Q7UMK2 (Q7UMK2) Probable transcription termination
factor rho, partial (4%)
Length = 617
Score = 24.3 bits (51), Expect = 5.1
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +2
Query: 1 KDKNTFSFKQGWAFPRKVYFNGDECMMPPPTPTHFCQILLL 41
K ++ F GW PR++ + PPP+P+ FC ++ L
Sbjct: 41 KTLSSTRFVFGWP-PRRLLLARRQ---PPPSPSVFCPLIFL 151
>BP040390
Length = 461
Score = 24.3 bits (51), Expect = 5.1
Identities = 13/33 (39%), Positives = 16/33 (48%), Gaps = 6/33 (18%)
Frame = -1
Query: 28 PPPTPTHFCQILLLQ------A*FPSRHSSSHY 54
PPP P HF L+Q + +RH SHY
Sbjct: 344 PPPPPPHFFPQRLMQHILDQLQHYQARHP*SHY 246
>TC16730 similar to GB|AAO64817.1|29028876|BT005882 At3g04570 {Arabidopsis
thaliana;}, partial (5%)
Length = 562
Score = 24.3 bits (51), Expect = 5.1
Identities = 15/44 (34%), Positives = 18/44 (40%)
Frame = -2
Query: 10 QGWAFPRKVYFNGDECMMPPPTPTHFCQILLLQA*FPSRHSSSH 53
QG A NG+ PP+PT L A PSR S+
Sbjct: 198 QGSAVAHWEELNGEGDYRTPPSPTRLLPQTLRPAAVPSREEESN 67
>TC19044 similar to UP|Q9LJH9 (Q9LJH9) Gb|AAD25781.1, partial (16%)
Length = 620
Score = 23.9 bits (50), Expect = 6.7
Identities = 8/18 (44%), Positives = 12/18 (66%)
Frame = +3
Query: 26 MMPPPTPTHFCQILLLQA 43
++PPP P F +LL+ A
Sbjct: 423 LLPPPPPNDFTMMLLMMA 476
>TC14761 similar to UP|AAS07369 (AAS07369) Expressed protein, partial (14%)
Length = 628
Score = 23.9 bits (50), Expect = 6.7
Identities = 14/35 (40%), Positives = 17/35 (48%), Gaps = 8/35 (22%)
Frame = -2
Query: 24 ECMMP--------PPTPTHFCQILLLQA*FPSRHS 50
+C++P P T CQ LL * PSRHS
Sbjct: 378 KCLIPFQ*SPQKHPSVTTSVCQFLLGG**LPSRHS 274
>TC11815 weakly similar to UP|Q8MLU0 (Q8MLU0) CG13503-PA, partial (3%)
Length = 629
Score = 23.9 bits (50), Expect = 6.7
Identities = 10/30 (33%), Positives = 16/30 (53%)
Frame = +2
Query: 5 TFSFKQGWAFPRKVYFNGDECMMPPPTPTH 34
+F+F + P K+ + PPP+PTH
Sbjct: 5 SFNFPTKNSNPNKIPNQHPTNLQPPPSPTH 94
>TC15303 similar to UP|Q9ZNZ5 (Q9ZNZ5) Peroxidase precursor (Fragment) ,
partial (36%)
Length = 710
Score = 23.5 bits (49), Expect = 8.7
Identities = 8/16 (50%), Positives = 13/16 (81%)
Frame = -3
Query: 31 TPTHFCQILLLQA*FP 46
+P+HFC I+ ++ *FP
Sbjct: 630 SPSHFCFIIPIRT*FP 583
>TC8758 similar to GB|AAC77862.2|20197419|AC005623 Argonaute (AGO1)-like
protein {Arabidopsis thaliana;} , partial (32%)
Length = 1142
Score = 23.5 bits (49), Expect = 8.7
Identities = 8/12 (66%), Positives = 10/12 (82%)
Frame = +3
Query: 21 NGDECMMPPPTP 32
NG+E +MPPP P
Sbjct: 93 NGNEDLMPPPPP 128
>TC8587 similar to UP|Q84MD8 (Q84MD8) At4g21470, partial (97%)
Length = 1519
Score = 23.5 bits (49), Expect = 8.7
Identities = 11/31 (35%), Positives = 13/31 (41%)
Frame = +2
Query: 7 SFKQGWAFPRKVYFNGDECMMPPPTPTHFCQ 37
SF GW V GDE P+P F +
Sbjct: 470 SFHDGWKDSFAVIIGGDEVRTGKPSPDIFIE 562
>TC10870 similar to GB|CAA77438.1|2924285|CHNTXX Ycf2 protein {Nicotiana
tabacum;}, complete
Length = 6897
Score = 23.5 bits (49), Expect = 8.7
Identities = 8/13 (61%), Positives = 11/13 (84%)
Frame = -3
Query: 49 HSSSHYSSFLFYL 61
H S+H+S FLF+L
Sbjct: 4507 HQSNHHSRFLFFL 4469
>AV419050
Length = 409
Score = 23.5 bits (49), Expect = 8.7
Identities = 8/27 (29%), Positives = 13/27 (47%)
Frame = +3
Query: 6 FSFKQGWAFPRKVYFNGDECMMPPPTP 32
F+F + W P + + PPP+P
Sbjct: 120 FAFNRPWPLPPRALSLSFRALPPPPSP 200
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.335 0.145 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,902,110
Number of Sequences: 28460
Number of extensions: 34180
Number of successful extensions: 314
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 313
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 314
length of query: 61
length of database: 4,897,600
effective HSP length: 37
effective length of query: 24
effective length of database: 3,844,580
effective search space: 92269920
effective search space used: 92269920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0338.8


