

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
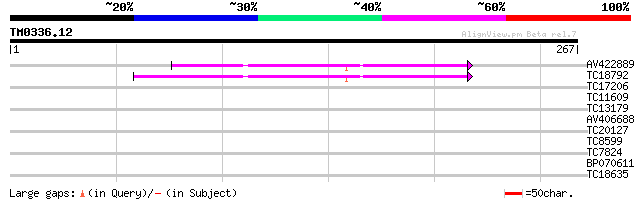
Query= TM0336.12
(267 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV422889 115 8e-27
TC18792 similar to UP|AAD03372 (AAD03372) Expressed protein, par... 108 1e-24
TC17206 similar to UP|Q8S531 (Q8S531) Cytosolic aldehyde dehydro... 29 0.95
TC11609 similar to UP|Q8S532 (Q8S532) Cytosolic aldehyde dehydro... 28 1.2
TC13179 similar to UP|Q9ZVA9 (Q9ZVA9) F9K20.4 protein, partial (5%) 27 2.8
AV406688 27 2.8
TC20127 27 3.6
TC8599 homologue to UP|Q40478 (Q40478) EREBP-4, partial (26%) 27 3.6
TC7824 similar to UP|O04919 (O04919) Lipoxygenase , partial (58%) 27 3.6
BP070611 26 6.2
TC18635 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (6%) 26 6.2
>AV422889
Length = 427
Score = 115 bits (288), Expect = 8e-27
Identities = 61/143 (42%), Positives = 85/143 (58%), Gaps = 1/143 (0%)
Frame = +2
Query: 77 VGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTE 136
+G + + LV KC FA+ KL+WE+LD K K E+PW I A++A E+
Sbjct: 2 IGTWEYKSRYEGDLVAKCYFAKHKLVWEVLDGCL--KNKIEIPWSDIMALKANYPEDAPG 175
Query: 137 ILEIELDEQPIFHKEINSQST-HTTWEISDHDFTGGHAMTYRIHHLEFASGDLGQHYENM 195
LE+ L +P+F +EIN Q HT W+ + DFTGG A R H ++ G LG+H+E +
Sbjct: 176 TLEVVLARRPLFFREINPQPRKHTLWQATS-DFTGGQASIQRRHFMQCPQGLLGKHFEKL 352
Query: 196 LKCDERLMKLSQQPFPRLDSPYF 218
++CD RL LSQQP L+SPYF
Sbjct: 353 IQCDPRLNFLSQQPELVLESPYF 421
>TC18792 similar to UP|AAD03372 (AAD03372) Expressed protein, partial (41%)
Length = 886
Score = 108 bits (269), Expect = 1e-24
Identities = 59/161 (36%), Positives = 86/161 (52%), Gaps = 1/161 (0%)
Frame = +1
Query: 59 SSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLIWEILDSSKNKKFKAEV 118
S K + F LL +G + + LV KC FA+ KL+WE+L+ K K E+
Sbjct: 238 SGAAEKLKASNFPGSLLRIGLWEYKSRYEGDLVAKCYFAKHKLVWEVLEGGL--KSKIEI 411
Query: 119 PWDRISAIRAFIEENKTEILEIELDEQPIFHKEINSQST-HTTWEISDHDFTGGHAMTYR 177
W I A++A EN L + L QP+F +E N Q HT W+ + DFT G +R
Sbjct: 412 QWADIMALKAQCPENGLSTLTVVLGRQPLFFRETNPQPRKHTLWQATA-DFTNGQCSKHR 588
Query: 178 IHHLEFASGDLGQHYENMLKCDERLMKLSQQPFPRLDSPYF 218
+H L+ G L +H+E +++CD RL LS+QP + SP+F
Sbjct: 589 LHFLQCPQGVLAKHFEKLIQCDLRLNFLSRQPEILMHSPHF 711
>TC17206 similar to UP|Q8S531 (Q8S531) Cytosolic aldehyde dehydrogenase
RF2C, partial (28%)
Length = 595
Score = 28.9 bits (63), Expect = 0.95
Identities = 13/44 (29%), Positives = 23/44 (51%)
Frame = +1
Query: 217 YFGPPVLFKNIRDYNTVQHDELMEQVMALDRTQPFLSELKSTIN 260
Y+ P +F N+++ + DE+ VMAL + + +KS N
Sbjct: 58 YYIEPTIFSNVKEDMLIVQDEIFGPVMALKKFKTIEEGIKSANN 189
>TC11609 similar to UP|Q8S532 (Q8S532) Cytosolic aldehyde dehydrogenase
RF2C, partial (25%)
Length = 600
Score = 28.5 bits (62), Expect = 1.2
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = +2
Query: 217 YFGPPVLFKNIRDYNTVQHDELMEQVMALDRTQPFLSELKSTINH 261
Y+ P +F N+++ + DE+ VMAL + + +KS NH
Sbjct: 26 YYIEPTIFSNVKEDMLIVQDEIFGPVMALKKFKTVEEAIKSA-NH 157
>TC13179 similar to UP|Q9ZVA9 (Q9ZVA9) F9K20.4 protein, partial (5%)
Length = 533
Score = 27.3 bits (59), Expect = 2.8
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = +2
Query: 6 PFGLELSSSTMNEREPHLRNVC 27
PFGL L+ ++NE++P L C
Sbjct: 173 PFGLNLNKGSINEKQPRLHLHC 238
>AV406688
Length = 423
Score = 27.3 bits (59), Expect = 2.8
Identities = 12/23 (52%), Positives = 15/23 (65%)
Frame = +1
Query: 54 SKKQNSSTTLKFAPATFYAKLLN 76
S NSS+T+ F P+TFY LN
Sbjct: 253 SPPMNSSSTVSFFPSTFYPPQLN 321
>TC20127
Length = 582
Score = 26.9 bits (58), Expect = 3.6
Identities = 11/48 (22%), Positives = 22/48 (44%), Gaps = 3/48 (6%)
Frame = +1
Query: 146 PIFHKEIN---SQSTHTTWEISDHDFTGGHAMTYRIHHLEFASGDLGQ 190
P++H+ + ++H W + + T GH Y L A+G+ +
Sbjct: 301 PLYHRAVELAEHDNSHQNWRVKAKNRTSGHVEEYAGKFLVVATGETAE 444
>TC8599 homologue to UP|Q40478 (Q40478) EREBP-4, partial (26%)
Length = 632
Score = 26.9 bits (58), Expect = 3.6
Identities = 19/59 (32%), Positives = 24/59 (40%)
Frame = +3
Query: 3 SKPPFGLELSSSTMNEREPHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSST 61
SKPP SSS ++ R+P L P GLK S KE + S + T
Sbjct: 201 SKPP----RSSSNLSNRKPSLNISIPSINSGLKSNISQSKETSFSHQSSSSSSSSDHQT 365
>TC7824 similar to UP|O04919 (O04919) Lipoxygenase , partial (58%)
Length = 1675
Score = 26.9 bits (58), Expect = 3.6
Identities = 18/60 (30%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Frame = +3
Query: 42 KEREKMWRKMQ-KSKKQNSSTTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRK 100
K+ E W KMQ +++ SSTTL + + +A +N G+Y + +N+ + +F K
Sbjct: 966 KKNEPWWPKMQTRAELIESSTTLIWTASALHA-AVNFGQYPYGGLILNRPTLSRRFMPEK 1142
>BP070611
Length = 486
Score = 26.2 bits (56), Expect = 6.2
Identities = 13/36 (36%), Positives = 18/36 (49%)
Frame = -1
Query: 191 HYENMLKCDERLMKLSQQPFPRLDSPYFGPPVLFKN 226
HY+ L+ RL + SQ+PF R+ LF N
Sbjct: 411 HYQTTLRTGMRLTRHSQEPFMRVSLKLIISRSLFSN 304
>TC18635 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (6%)
Length = 610
Score = 26.2 bits (56), Expect = 6.2
Identities = 13/42 (30%), Positives = 22/42 (51%)
Frame = +3
Query: 223 LFKNIRDYNTVQHDELMEQVMALDRTQPFLSELKSTINHELS 264
+F IR+ Q DE+ ++ L RT+P ++ T E+S
Sbjct: 3 MFAKIRETTLAQDDEISRNIVGLARTRP---DIFGTTEEEVS 119
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.133 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,626,764
Number of Sequences: 28460
Number of extensions: 60282
Number of successful extensions: 320
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 317
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 318
length of query: 267
length of database: 4,897,600
effective HSP length: 89
effective length of query: 178
effective length of database: 2,364,660
effective search space: 420909480
effective search space used: 420909480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0336.12


