

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0335b.5
(1648 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
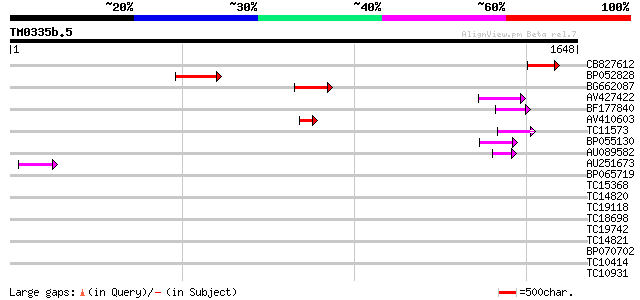
Score E
Sequences producing significant alignments: (bits) Value
CB827612 118 9e-27
BP052828 110 2e-24
BG662087 97 3e-20
AV427422 70 4e-12
BF177840 61 1e-09
AV410603 59 5e-09
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 59 8e-09
BP055130 54 3e-07
AU089582 46 4e-05
AU251673 45 1e-04
BP065719 36 0.056
TC15368 homologue to UP|Q84LL5 (Q84LL5) Cullin 4, partial (11%) 35 0.12
TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate p... 32 0.62
TC19118 weakly similar to UP|Q84YE8 (Q84YE8) Expressed protein-l... 32 0.81
TC18698 32 0.81
TC19742 weakly similar to UP|SDF2_ARATH (Q93ZE8) Stromal cell-de... 32 1.1
TC14821 homologue to UP|AAR12195 (AAR12195) Molecular chaperone ... 31 1.4
BP070702 31 1.8
TC10414 similar to GB|AAP13363.1|30023660|BT006255 At5g55710 {Ar... 31 1.8
TC10931 UP|O22322 (O22322) Ripening-associated protein (Fragment... 30 2.4
>CB827612
Length = 277
Score = 118 bits (295), Expect = 9e-27
Identities = 55/92 (59%), Positives = 70/92 (75%)
Frame = +1
Query: 1505 ACHLPVELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHD 1564
ACHLPVE+EH+AYWA++ N + + A +R LQL +L+E RL AYES+ IYKEKTK HD
Sbjct: 1 ACHLPVEIEHRAYWAVKSCNXEMQQAGIERKLQLQQLEELRLGAYESSRIYKEKTKHXHD 180
Query: 1565 RKILNREFVSGQLVLLFNSRLRLFPGKLKSRW 1596
+ I +EF GQ VLLFNSRL+L GKL+S+W
Sbjct: 181 KMISRKEFSVGQQVLLFNSRLKLMAGKLRSKW 276
>BP052828
Length = 523
Score = 110 bits (275), Expect = 2e-24
Identities = 59/138 (42%), Positives = 90/138 (64%), Gaps = 4/138 (2%)
Frame = +2
Query: 482 LCDLGASINLMSLRMFKRLNLGEVTPTMISLQMADRSLKTPYGIIEDVVVKVDKFVFPVD 541
+ DLGA IN+M ++ L+ G + T + +Q+A+RS P G++EDV+V+V++ +FP D
Sbjct: 107 MLDLGAGINVMPTSIYNILHPGPLQHTGLIVQLANRSNARPKGVVEDVLVQVNELIFPAD 286
Query: 542 FVILDME---EDSKVPLILGRPFLATGEAEIKVAKGTLTLKVGEDEVLFNIFDSLKH-RA 597
F ILDME + S+ P+ILGRPF+ T + +I V GT++++ G+ FNI D++KH
Sbjct: 287 FYILDMEGETKSSRAPIILGRPFMKTAKTKIDVDNGTMSMEFGDIIAKFNIDDAMKHPLE 466
Query: 598 DEEVFRCEVVDELVCEEF 615
D VF VV ELV + F
Sbjct: 467 DHSVFHIGVVSELVDDTF 520
>BG662087
Length = 373
Score = 96.7 bits (239), Expect = 3e-20
Identities = 44/110 (40%), Positives = 69/110 (62%)
Frame = +1
Query: 827 KWRVCIDYRRLNSVTRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKT 886
KWR+ +DY LN KD +PLP ID+++D + ++ +D YSGY+QI + P D++KT
Sbjct: 19 KWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDKT 198
Query: 887 AFTCPYGVFAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSV 936
AF + Y+ +PFGL NA AT+Q M +F D + +E+++D+ V
Sbjct: 199 AFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIV 348
>AV427422
Length = 417
Score = 69.7 bits (169), Expect = 4e-12
Identities = 42/135 (31%), Positives = 67/135 (49%), Gaps = 1/135 (0%)
Frame = +1
Query: 1364 PPSFGC*YILVAVDYVSKWVEASALS-TNDSKVVVAFLKKNIFTRFGVPRAIISDGGTHF 1422
P S G +LV VD +SK+ L +KV+ + + GVP +I+SD F
Sbjct: 13 PKSKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLF 192
Query: 1423 CNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSRKLDDAL 1482
+ ++ L + G K K+ST YHP++ GQ E+ NR L+ L + K W+ + A
Sbjct: 193 MSNFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAE 372
Query: 1483 WAYRTAFKTPIGTSP 1497
+ Y T++ G +P
Sbjct: 373 YWYNTSYHVSTGQTP 417
>BF177840
Length = 410
Score = 61.2 bits (147), Expect = 1e-09
Identities = 34/100 (34%), Positives = 50/100 (50%)
Frame = +2
Query: 1413 AIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRK 1472
+I+SD T F + + +L K G K ST HPQT GQ E+ N+ L +L V+ + K
Sbjct: 14 SIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLK 193
Query: 1473 DWSRKLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVEL 1512
W L +AY + SPF +V+G P++L
Sbjct: 194 MWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>AV410603
Length = 162
Score = 59.3 bits (142), Expect = 5e-09
Identities = 27/53 (50%), Positives = 36/53 (66%)
Frame = +1
Query: 841 TRKDHFPLPFIDQMLDRLAGHQYYCFLDGYSGYNQICVAPEDQEKTAFTCPYG 893
T KD FP+P +D++LD L G Q++ LD SGY+QI V PED+ KT F +G
Sbjct: 4 TVKDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 58.5 bits (140), Expect = 8e-09
Identities = 30/111 (27%), Positives = 56/111 (50%)
Frame = +2
Query: 1417 DGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDWSR 1476
+ G F ++ + + G++ + S+ HPQT+GQ E +N+ + + +++ + + W
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 1477 KLDDALWAYRTAFKTPIGTSPFHLVFGKACHLPVELEHKAYWAIRKLNFDW 1527
+L LW+Y T ++ I +PF L +G LPVE+ NF W
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEIS----------NFSW 310
>BP055130
Length = 567
Score = 53.5 bits (127), Expect = 3e-07
Identities = 36/112 (32%), Positives = 56/112 (49%), Gaps = 2/112 (1%)
Frame = +2
Query: 1365 PSFGC*YILVAVDYVSKWVEASALSTN--DSKVVVAFLKKNIFTRFGVPRAIISDGGTHF 1422
P G I+V VD +SK+ + N S+V F+ I G+PRAI+SD F
Sbjct: 170 PVCGFTVIIVVVDRLSKYGHFAPHRANYTSSQVAETFVS-TIVKLHGMPRAIVSDRDKAF 346
Query: 1423 CNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRILEKVVNSSRKDW 1474
+ ++ + +G +S+ YHPQT GQ E N+ L+ L V+ + + W
Sbjct: 347 TSAFWKHFFKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLW 502
>AU089582
Length = 383
Score = 46.2 bits (108), Expect = 4e-05
Identities = 26/68 (38%), Positives = 37/68 (54%)
Frame = +2
Query: 1404 IFTRFGVPRAIISDGGTHFCNRAFESLLEKYGVKHKVSTPYHPQTSGQVEISNRELKRIL 1463
I + GVP +IISD G F + + S G + K+ST +HPQT GQ E + + L+ +L
Sbjct: 44 IVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSERTIQILEDML 223
Query: 1464 EKVVNSSR 1471
V R
Sbjct: 224 RACVXDLR 247
>AU251673
Length = 413
Score = 44.7 bits (104), Expect = 1e-04
Identities = 36/122 (29%), Positives = 55/122 (44%), Gaps = 8/122 (6%)
Frame = +2
Query: 25 SEAIRLSLFPFSLKDAAEEWLN----SQPQGSLT-TWEDLAEKFTTRFFPRSLLRKLKND 79
S+ + L F L+ A +W N ++P GS TW D + +F RF P+S+ D
Sbjct: 8 SDTRAVELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVRD 187
Query: 80 IMTFTQSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFYDGL---LYSSRFGLDAASS 136
Q+ + E HF L R P + L + E V +F GL L+ S G +++
Sbjct: 188 FERLEQAEGMTVSEYSAHFTHLSRYVP-YPLLEEERVKRFVRGLKEYLFKSVVGSKSSTL 364
Query: 137 GE 138
E
Sbjct: 365 SE 370
>BP065719
Length = 567
Score = 35.8 bits (81), Expect = 0.056
Identities = 25/99 (25%), Positives = 44/99 (44%)
Frame = +3
Query: 25 SEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTWEDLAEKFTTRFFPRSLLRKLKNDIMTFT 84
+E +++ FP SL A W + S+ TW L F +FF R + D+ +
Sbjct: 273 NENLKMKYFPSSLTKNAFTWFTTLAPRSVHTWAQLERIFHEQFF-RGECKVSXKDLASVK 449
Query: 85 QSTDENLYEAWEHFKKLLRKCPQHNLTQAECVAKFYDGL 123
+ E++ + F+ L +C H +++ E V GL
Sbjct: 450 RKPAESIDDYLNRFRMLKSRCFTH-VSEHELVVLXAGGL 563
>TC15368 homologue to UP|Q84LL5 (Q84LL5) Cullin 4, partial (11%)
Length = 517
Score = 34.7 bits (78), Expect = 0.12
Identities = 19/83 (22%), Positives = 38/83 (44%)
Frame = +1
Query: 628 IMEGLEIEDQNLERELDSLYHEVNATLSQLESVSSLATKSIWKEELTRDEEIPIEEKSEL 687
+ E +E+ ER ++V+A + ++ + + ++ EL + + PI+
Sbjct: 1 LKETVEVNTSTTERVFQDRQYQVDAAIVRIMKTRKVLSHTLLITELFQQLKFPIKPADLK 180
Query: 688 KSLPSSLKYAYLEEGENKPVILN 710
K + S + YLE +N P I N
Sbjct: 181 KRIESLIDREYLERDKNNPQIYN 249
>TC14820 homologue to UP|HS80_LYCES (P36181) Heat shock cognate protein 80,
partial (53%)
Length = 1180
Score = 32.3 bits (72), Expect = 0.62
Identities = 16/67 (23%), Positives = 33/67 (48%)
Frame = +1
Query: 1511 ELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNR 1570
E E K I++++ +W + ++++ + + + +E Y A+ YK T W + +
Sbjct: 799 EKEEKKKKKIKEVSHEWSLVNKQKPIWMRKPEEINKEEY--AAFYKSLTNDWEEHLAVKH 972
Query: 1571 EFVSGQL 1577
V GQL
Sbjct: 973 FSVEGQL 993
>TC19118 weakly similar to UP|Q84YE8 (Q84YE8) Expressed protein-like protein,
partial (11%)
Length = 932
Score = 32.0 bits (71), Expect = 0.81
Identities = 24/74 (32%), Positives = 34/74 (45%), Gaps = 5/74 (6%)
Frame = +3
Query: 1437 KHKVSTPYHPQT-----SGQVEISNRELKRILEKVVNSSRKDWSRKLDDALWAYRTAFKT 1491
+HK+S P HP T S +++ + L R + V ++ S LD+AL R
Sbjct: 3 EHKISDPQHPSTYKSEQSNRLKEDSMLLMRGFDSVAHTLSL-MSSNLDNALQGARELANP 179
Query: 1492 PIGTSPFHLVFGKA 1505
P T FH F KA
Sbjct: 180 PTLTDIFHSKFDKA 221
>TC18698
Length = 808
Score = 32.0 bits (71), Expect = 0.81
Identities = 16/42 (38%), Positives = 24/42 (57%)
Frame = -2
Query: 895 FAYKRMPFGLCNAPATFQRCMFAIFSDLIETCIEIFMDDFSV 936
+ Y+ MP GL N T+QR M IF I +E++++D V
Sbjct: 771 YYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIV 646
>TC19742 weakly similar to UP|SDF2_ARATH (Q93ZE8) Stromal cell-derived
factor 2-like protein precursor (SDF2-like protein),
partial (24%)
Length = 474
Score = 31.6 bits (70), Expect = 1.1
Identities = 16/36 (44%), Positives = 21/36 (57%)
Frame = -3
Query: 330 SGRVLHEPKEKVEGEKNEEAEVERDEGVMEERAREP 365
SG E + ++EGE+ EE+E E E EER EP
Sbjct: 352 SGGRRREGEFRIEGEEEEESESEESESHEEERCDEP 245
>TC14821 homologue to UP|AAR12195 (AAR12195) Molecular chaperone Hsp90-1,
partial (29%)
Length = 612
Score = 31.2 bits (69), Expect = 1.4
Identities = 16/67 (23%), Positives = 33/67 (48%)
Frame = +3
Query: 1511 ELEHKAYWAIRKLNFDWKVASEKRLLQLNELDEFRLRAYESASIYKEKTKKWHDRKILNR 1570
E E K I++++ +W + ++++ + + + +E Y A+ YK T W + +
Sbjct: 138 EKEEKKKKKIKEVSHEWSLVNKQKPIWMRKPEEITKDEY--AAFYKSLTNDWEEHLAVKH 311
Query: 1571 EFVSGQL 1577
V GQL
Sbjct: 312 FSVEGQL 332
>BP070702
Length = 511
Score = 30.8 bits (68), Expect = 1.8
Identities = 15/39 (38%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Frame = -2
Query: 712 VLTPLKEEKLLKVLRD-HKSALGWTIDDIKGISPAICMH 749
V PL +EKL++VLR +K WT+DD + + + H
Sbjct: 402 VYHPLIDEKLMRVLRRRNKRVYAWTVDDAESMQKMLFEH 286
>TC10414 similar to GB|AAP13363.1|30023660|BT006255 At5g55710 {Arabidopsis
thaliana;}, partial (34%)
Length = 504
Score = 30.8 bits (68), Expect = 1.8
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = -3
Query: 14 NCNTYRVVNVSSEAIRLSLFPFSLKDAAEEWLNSQPQGSLTTW 56
NC Y+ +N E + L + FS+ A EEW+ +G T W
Sbjct: 430 NCR-YQWLNEDLERVELGDYVFSVLYAVEEWVEVAERGDQTVW 305
>TC10931 UP|O22322 (O22322) Ripening-associated protein (Fragment), partial
(7%)
Length = 890
Score = 30.4 bits (67), Expect = 2.4
Identities = 34/120 (28%), Positives = 49/120 (40%), Gaps = 13/120 (10%)
Frame = +1
Query: 638 NLERELDS---LYHEVNATLSQLESVSSL-ATKSI----WKEELTRDEEIPIEEKSELKS 689
NL + D LY E +AT L S +SL +KS+ + + + EE+P+EEK
Sbjct: 214 NLSEDSDQTPLLYQEESATNGSLLSQTSLNLSKSMPHTEYDQSMLHSEEMPVEEKIVTVE 393
Query: 690 LPSSLKY-----AYLEEGENKPVILNSVLTPLKEEKLLKVLRDHKSALGWTIDDIKGISP 744
S KY L E E +EE L ++ GWT ++ SP
Sbjct: 394 RSYSKKYKAPFKGLLREEEK------------EEEAHLLLMPQPPDNHGWTKKEVSSTSP 537
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,049,414
Number of Sequences: 28460
Number of extensions: 392931
Number of successful extensions: 1702
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 1686
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1699
length of query: 1648
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1545
effective length of database: 1,966,220
effective search space: 3037809900
effective search space used: 3037809900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0335b.5


