

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0334a.1
(1167 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
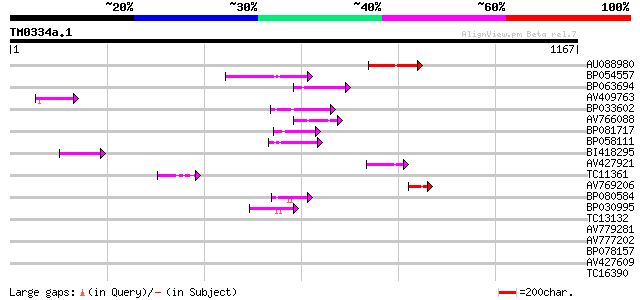
Score E
Sequences producing significant alignments: (bits) Value
AU088980 121 6e-28
BP054557 95 6e-20
BP063694 84 2e-16
AV409763 79 5e-15
BP033602 60 3e-09
AV766088 58 8e-09
BP081717 55 6e-08
BP058111 55 6e-08
BI418295 54 1e-07
AV427921 54 1e-07
TC11361 46 3e-05
AV769206 45 7e-05
BP080584 45 9e-05
BP030995 42 4e-04
TC13132 similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement... 41 0.001
AV779281 39 0.004
AV777202 36 0.040
BP078157 35 0.069
AV427609 33 0.20
TC16390 similar to GB|AAM63374.1|21554299|AY086170 apospory-asso... 33 0.26
>AU088980
Length = 360
Score = 121 bits (304), Expect = 6e-28
Identities = 55/111 (49%), Positives = 81/111 (72%)
Frame = +2
Query: 738 NGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRIQSKVLENKSPYEILFGG 797
N VERKH+HILN+ RAL+F S+LPK +W +A+ HA +L++ + S +L+ + PY+ L
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFL-NN 178
Query: 798 KKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVLLDL 848
L +L+VFG+LC++ST + + KLD RARKCVFLGFK G G++++DL
Sbjct: 179 DSPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMDL 331
>BP054557
Length = 538
Score = 95.1 bits (235), Expect = 6e-20
Identities = 61/182 (33%), Positives = 89/182 (48%), Gaps = 4/182 (2%)
Frame = -2
Query: 445 IIDSGASDHICFNKLCFDTLNRIKPVSVRLPNGISIQTCYAGVVSITKQILLTHVLYVPE 504
++D+GA+ H+C N F L ++ PV + +PNG A V +++ L VLYVP
Sbjct: 537 LLDTGATHHVCHNLDVFQALRQVHPVLLSMPNGQKEIAHMADSVYFSEKFFLAXVLYVPS 358
Query: 505 FTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLKEDDLYHLVVTSPSSTCF 564
F +NLISV L K C +V + C +QE+ T K IG + LY V P+
Sbjct: 357 FNFNLISVSALVKSLNCRLVIFDECCLLQERPTLKTIGAAE-ERGGLY--AVIDPAVKAK 187
Query: 565 AIP---NVVERISDSDQLIPPGALWHFRLGHLSHDRILALNALYPSIDVSK-HFVCDICH 620
P N+ ++S + LWH RLGH S ++ L +P I + CD CH
Sbjct: 186 HPPAEGNIKTQVSHFNFCQFDCNLWHLRLGHTSSKKLAILQKNFPFITCKQITSPCDTCH 7
Query: 621 LA 622
+A
Sbjct: 6 MA 1
>BP063694
Length = 511
Score = 83.6 bits (205), Expect = 2e-16
Identities = 46/120 (38%), Positives = 65/120 (53%), Gaps = 4/120 (3%)
Frame = -2
Query: 585 LWHFRLGHLSHDRILALNALYPSIDVS----KHFVCDICHLAKLKRKMFPDSLHNAKCNF 640
LWH RLGH++ D I L+ + +DVS K C C +AK KR +F D+ A
Sbjct: 360 LWHSRLGHVNFDIIKQLHK-HGCLDVSSILPKPICCTSCQMAKSKRLVFHDNNKRASAVL 184
Query: 641 DLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFS 700
DL+H D+ GP V+S+ YF+ +DD SRF W LKRK + + F ++ +FS
Sbjct: 183 DLIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVLLRFKVFMENRFS 4
>AV409763
Length = 426
Score = 78.6 bits (192), Expect = 5e-15
Identities = 37/99 (37%), Positives = 57/99 (57%), Gaps = 9/99 (9%)
Frame = +2
Query: 53 LEPAQNPI---------SPYYIHSGENPSATVVSLPLNGRNYNAWA*SMKRVLVAKNKFK 103
+E A NP +P+Y+H +NP ++VS L G NY +W+ +M+ L+AKNK
Sbjct: 125 MEDAANPFMANANPFMANPFYLHPSDNPGISLVSQVLTGDNYRSWSRAMRMALIAKNKLG 304
Query: 104 FINGEIPIAVPGDANYEAWDRCNNLIHSWILNSVTSSIA 142
F++ I + D + W R NN++ SWILNSV+ +A
Sbjct: 305 FVDASILVPDELDPLFHPWQRNNNIVASWILNSVSKDLA 421
>BP033602
Length = 533
Score = 59.7 bits (143), Expect = 3e-09
Identities = 41/134 (30%), Positives = 69/134 (50%), Gaps = 1/134 (0%)
Frame = +3
Query: 538 KKRIGLGSLKEDDLYHLVVTSPSSTCFAIPNVVERISDSDQLIPPGALWHFRLGHLSHDR 597
+K IG +K D LY+ T + + + S DQ++ LWH RLGH +
Sbjct: 153 QKMIGSAKMK-DGLYYFEDTFRNKAAQGLCGM-SCTSVRDQIM----LWHNRLGHPNFQY 314
Query: 598 ILALNA-LYPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCNFDLLHMDIWGPISVSSV 656
+ L L+ +++ S C+ C LAK +R + ++A F L+H D+WGP +++
Sbjct: 315 LRHLFPDLFKNVNCSS-LECESCVLAKNQRAPYYSQPYHASRPFYLIHSDVWGPSKITTQ 491
Query: 657 HSHRYFLTVLDDHS 670
R+F+T +DDH+
Sbjct: 492 FGKRWFVTFIDDHT 533
>AV766088
Length = 501
Score = 58.2 bits (139), Expect = 8e-09
Identities = 36/105 (34%), Positives = 52/105 (49%), Gaps = 4/105 (3%)
Frame = -3
Query: 584 ALWHFRLGHLSHDRILALNA----LYPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCN 639
+LWH RLGH S + L + Y ++ S VC+ C K R F S +
Sbjct: 310 SLWHSRLGHPSSSALRYLRSNKFISYELLNYSP--VCESCVFGKHVRLPFVSSNNVTVMP 137
Query: 640 FDLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEV 684
FD+LH D+W +SS HR+++ LDD + F+W L K +V
Sbjct: 136 FDILHSDLWTSPVLSSA-GHRFYVLFLDDFTDFLWTFPLSNKSQV 5
>BP081717
Length = 291
Score = 55.1 bits (131), Expect = 6e-08
Identities = 31/98 (31%), Positives = 49/98 (49%)
Frame = -1
Query: 543 LGSLKEDDLYHLVVTSPSSTCFAIPNVVERISDSDQLIPPGALWHFRLGHLSHDRILALN 602
L +LKE L L+ S S N+ E + + P +WHFRLGH S + ++
Sbjct: 282 LRNLKESVLRSLIGISQIS----FVNIAETVPGKNGSHPQFDMWHFRLGHPSSKVLQHIS 115
Query: 603 ALYPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCNF 640
++P + + + +CD CH+AK + FP S +K F
Sbjct: 114 KVFPYVKLHSNIICDFCHMAKQTKLPFPISDTKSKACF 1
>BP058111
Length = 570
Score = 55.1 bits (131), Expect = 6e-08
Identities = 40/116 (34%), Positives = 57/116 (48%), Gaps = 4/116 (3%)
Frame = -1
Query: 533 QEKHTKKRIGLGSLKEDDLYHLVVTSPSSTCFAIP---NVVERISDSDQLIPPGALWHFR 589
QE+ T K IG + DLY L+ P+ P N+ ++S + LWH R
Sbjct: 342 QERPTLKMIGAAE-ERGDLYALI--DPAVKVKHPPVEDNIKTQVSHFNFCQFDCNLWHLR 172
Query: 590 LGHLSHDRILALNALYPSIDVSK-HFVCDICHLAKLKRKMFPDSLHNAKCNFDLLH 644
LGH S ++ L +P I K CD CH+AK K+ FP+S+ + FDL+H
Sbjct: 171 LGHTSSKKLAILQKNFPFITCKKITSPCDTCHMAKQKKLSFPNSVTLSSEIFDLIH 4
>BI418295
Length = 523
Score = 54.3 bits (129), Expect = 1e-07
Identities = 21/95 (22%), Positives = 53/95 (55%)
Frame = +2
Query: 103 KFINGEIPIAVPGDANYEAWDRCNNLIHSWILNSVTSSIANSIVFVENACDAWRDLRDRF 162
+F+ E +A + Y W + ++++ +W+L +++ + +++ E + WR++ D F
Sbjct: 89 RFLTEEDRVAATENPAYTTWIKQDSMVFTWLLTTLSPFVLPTVINCERSSQVWREILDFF 268
Query: 163 SQGVLVRIAELQNEIGNLKQNTLSVNDYYTEIKTL 197
+ A+L++E+ N+ + + S+ +Y IKT+
Sbjct: 269 ENQCRAQSAKLRSELKNITKGSKSITEYLRRIKTI 373
>AV427921
Length = 284
Score = 53.9 bits (128), Expect = 1e-07
Identities = 30/88 (34%), Positives = 50/88 (56%)
Frame = +1
Query: 734 TPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRIQSKVLENKSPYEI 793
TPQQNG ERK++ ILN+ R+LL S LPK A++ ++ +++R + V++N P E
Sbjct: 22 TPQQNGVSERKNRTILNMVRSLLTMSGLPKSFLPEAVMWSLHILNRSPTLVVQNMMPEE- 198
Query: 794 LFGGKKIDLQELRVFGSLCFASTSSSSR 821
+ G++ + R+ L +A R
Sbjct: 199 AWSGRQPAVDHFRISRCLAYAHVPDQKR 282
>TC11361
Length = 549
Score = 46.2 bits (108), Expect = 3e-05
Identities = 30/88 (34%), Positives = 42/88 (47%)
Frame = +3
Query: 305 SSGSGYKSTGKYCTHCKKPGHTVDVCYRLHGYPTTSKSKPAFNNVSHINNIT*GYDSEDD 364
S +K CTHC GHT D CYRL G+P K +V+ +N +T G +ED
Sbjct: 6 SKSKSHKKDRPTCTHCGILGHTKDKCYRLIGFPPHYKKS---KSVASVNAVTTG--TEDS 170
Query: 365 QEESSKSQRGNGDLFTADQYKTIMAMIQ 392
+S FTA Q + ++ M+Q
Sbjct: 171 PSANSFVP------FTAQQCQVLLQMLQ 236
>AV769206
Length = 536
Score = 45.1 bits (105), Expect = 7e-05
Identities = 21/49 (42%), Positives = 31/49 (62%)
Frame = -3
Query: 822 SKLDSRARKCVFLGFKQGVKGFVLLDLNSHEIFVSRDVTFHEQILPYKS 870
+KL +++CVF+G+ KG+ LD S I++S+DV FHE PY S
Sbjct: 513 TKLSLHSKECVFIGYSINHKGYKCLD-QSGRIYISKDVLFHEHRFPYPS 370
>BP080584
Length = 507
Score = 44.7 bits (104), Expect = 9e-05
Identities = 37/102 (36%), Positives = 47/102 (45%), Gaps = 17/102 (16%)
Frame = -3
Query: 539 KRIGLGSLKEDDLYHLVVTSPSSTCFAIPNVVE------RISDSDQ----------LIPP 582
KRIG + LY L+ +P ST N V +S SD IP
Sbjct: 304 KRIGTARAVKG-LYLLMKGNPKSTTSVTCNSVSPACNSVSVSGSDSPVVDTNLACNFIPS 128
Query: 583 -GALWHFRLGHLSHDRILALNALYPSIDVSKHFVCDICHLAK 623
LWHFRLGH S + ++ ++P I +K VCDIC LAK
Sbjct: 127 MHNLWHFRLGHPSFVKEQSIRDIFPYIVCNKEQVCDICPLAK 2
>BP030995
Length = 458
Score = 42.4 bits (98), Expect = 4e-04
Identities = 34/118 (28%), Positives = 52/118 (43%), Gaps = 17/118 (14%)
Frame = -3
Query: 494 ILLTHVLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLKE----- 548
++L ++L+VP NL SV K+N F +C V+ + T++ + G+L+
Sbjct: 426 LVLNNLLHVPGVCKNLSSVSXFCKQNNVFFEFHSDSCLVKSQETEETLLQGTLRNGLCAF 247
Query: 549 DDLYHLV------------VTSPSSTCFAIPNVVERISDSDQLIPPGALWHFRLGHLS 594
D+L H V + S SS A+ V + Q LWH RLGH S
Sbjct: 246 DNLLHKVNLHSASTLSNSALLSSSSPASALSVSVFSSNKCAQADIDSRLWHSRLGHAS 73
>TC13132 similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement pol
polyprotein, partial (3%)
Length = 591
Score = 41.2 bits (95), Expect = 0.001
Identities = 15/27 (55%), Positives = 18/27 (66%)
Frame = +1
Query: 317 CTHCKKPGHTVDVCYRLHGYPTTSKSK 343
C+HC PGHT C++LHGYP K K
Sbjct: 40 CSHCGIPGHTAAKCFKLHGYPPGMKPK 120
>AV779281
Length = 505
Score = 39.3 bits (90), Expect = 0.004
Identities = 16/95 (16%), Positives = 51/95 (52%)
Frame = +1
Query: 103 KFINGEIPIAVPGDANYEAWDRCNNLIHSWILNSVTSSIANSIVFVENACDAWRDLRDRF 162
+F++ E + + Y+ W++ ++ + +W+L+S++ ++ ++V + + D F
Sbjct: 202 RFLSDEDRVNAVENPAYQLWEQQDSALFTWLLSSLSPTVLPTVVNCSQSWQVREAVLDFF 381
Query: 163 SQGVLVRIAELQNEIGNLKQNTLSVNDYYTEIKTL 197
+ +L++E+ ++ + + + +DY IK++
Sbjct: 382 KAHSFAQSTQLRSELRSIAKGSKTCSDYLKRIKSI 486
>AV777202
Length = 421
Score = 35.8 bits (81), Expect = 0.040
Identities = 14/60 (23%), Positives = 34/60 (56%)
Frame = +3
Query: 103 KFINGEIPIAVPGDANYEAWDRCNNLIHSWILNSVTSSIANSIVFVENACDAWRDLRDRF 162
+F++ E + + Y+ WD+ ++ + +W+L+S++SS+ ++V + W + D F
Sbjct: 213 RFLSEEDRVNAVENPAYQVWDQQDSALFTWLLSSLSSSVLPTVVNCTQSWQIWEAVLDFF 392
>BP078157
Length = 395
Score = 35.0 bits (79), Expect = 0.069
Identities = 19/48 (39%), Positives = 27/48 (55%), Gaps = 3/48 (6%)
Frame = -2
Query: 585 LWHFRLGHLSHDRILALNALYPSI---DVSKHFVCDICHLAKLKRKMF 629
LWH RLGHLS + ++ L P D H C++C L+K++R F
Sbjct: 148 LWHSRLGHLSDKVLKVVSNLVPFTVLPDFHSH-NCNVCPLSKMRRLSF 8
>AV427609
Length = 332
Score = 33.5 bits (75), Expect = 0.20
Identities = 21/84 (25%), Positives = 35/84 (41%), Gaps = 2/84 (2%)
Frame = +1
Query: 116 DANYEAWDRCNNLIHSWILNSVTSSIANSIVFV-ENACDAWRDLRDRFSQGVLVRIAELQ 174
D YE W + + WI ++++ + +I+ A AW L D F R L+
Sbjct: 76 DPAYEQWATLDATVLQWIYSTISVDLMTTIMEPGSTAFTAWTHLADLFQDNQNARAVTLE 255
Query: 175 NEIGNLKQNTL-SVNDYYTEIKTL 197
E ++ +V Y +KTL
Sbjct: 256 QEFSTVRMEAFPTVAAYCQRLKTL 327
>TC16390 similar to GB|AAM63374.1|21554299|AY086170 apospory-associated
protein C {Arabidopsis thaliana;}, partial (34%)
Length = 532
Score = 33.1 bits (74), Expect = 0.26
Identities = 23/71 (32%), Positives = 30/71 (41%), Gaps = 7/71 (9%)
Frame = -1
Query: 283 KSLVKMAEGKKTYGKGKAPGSSSSG-------SGYKSTGKYCTHCKKPGHTVDVCYRLHG 335
KSL EG T K P SS G + ST C + P T D+ + HG
Sbjct: 328 KSL*NQVEG--TTESSKRPLHSSPGFKVIGFPTAAPSTHSMCLYSSSPKSTTDLDFFSHG 155
Query: 336 YPTTSKSKPAF 346
+ TT+ P+F
Sbjct: 154 FQTTTSGSPSF 122
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.141 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,554,661
Number of Sequences: 28460
Number of extensions: 322072
Number of successful extensions: 1987
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 1948
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1976
length of query: 1167
length of database: 4,897,600
effective HSP length: 100
effective length of query: 1067
effective length of database: 2,051,600
effective search space: 2189057200
effective search space used: 2189057200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0334a.1


