

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
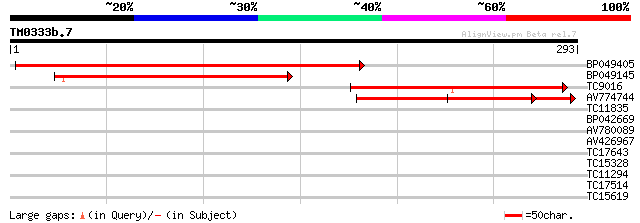
Query= TM0333b.7
(293 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP049405 333 3e-92
BP049145 165 7e-42
TC9016 weakly similar to GB|AAM10351.1|20147275|AY093727 AT3g627... 135 1e-32
AV774744 117 2e-27
TC11835 similar to UP|Q9LG00 (Q9LG00) F20N2.10, partial (35%) 28 2.4
BP042669 27 4.1
AV780089 27 5.3
AV426967 27 5.3
TC17643 26 6.9
TC15328 weakly similar to UP|Q9LP71 (Q9LP71) T1N15.15, partial (... 26 6.9
TC11294 26 9.1
TC17514 weakly similar to UP|O48683 (O48683) F3I6.9 protein, par... 26 9.1
TC15619 similar to UP|Q8GTX9 (Q8GTX9) Cell cycle control crn (Cr... 26 9.1
>BP049405
Length = 546
Score = 333 bits (853), Expect = 3e-92
Identities = 165/181 (91%), Positives = 168/181 (92%), Gaps = 1/181 (0%)
Frame = +1
Query: 4 HTSSIFFLLSSLVLSLFI-PIFCSFEDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGLDHA 62
H + FLL+ L + P CSFEDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGLDHA
Sbjct: 4 HAHILHFLLTILPSPFPLHPNICSFEDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGLDHA 183
Query: 63 DPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEAF 122
DPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEAF
Sbjct: 184 DPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEAF 363
Query: 123 AEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLL 182
AEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLL
Sbjct: 364 AEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGLL 543
Query: 183 G 183
G
Sbjct: 544 G 546
>BP049145
Length = 447
Score = 165 bits (418), Expect = 7e-42
Identities = 78/128 (60%), Positives = 97/128 (74%), Gaps = 5/128 (3%)
Frame = +3
Query: 24 FCS-----FEDVVDVDLVQFHLNLEFLEAEFYFYGTLGWGLDHADPKLAQGGPPPVGGQY 78
FCS D D+DL++F LNLE+LEAEF+ +G G GLD P L GGPPP+G +
Sbjct: 63 FCSHASAAIPDQNDLDLLEFPLNLEYLEAEFFLFGAXGHGLDVVAPNLTGGGPPPIGARA 242
Query: 79 AYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSAFGKPLNPPF 138
A LDP++RD+ QFA Q+ GHLRAI VKGFPRPLLN+SK +FAE+++SAFGKPL+PPF
Sbjct: 243 AKLDPLVRDVIYQFALQEVGHLRAIKSTVKGFPRPLLNLSKASFAEIMDSAFGKPLSPPF 422
Query: 139 DPYANSLN 146
DPYANS+N
Sbjct: 423 DPYANSIN 446
>TC9016 weakly similar to GB|AAM10351.1|20147275|AY093727
AT3g62730/F26K9_160 {Arabidopsis thaliana;}, partial
(37%)
Length = 571
Score = 135 bits (339), Expect = 1e-32
Identities = 71/125 (56%), Positives = 85/125 (67%), Gaps = 13/125 (10%)
Frame = +2
Query: 177 LVSGLLGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRNDLGK--------- 227
LV+GLLG +SGQDA+IR LLYERR L+V PY +V + T+R+S LRN LG
Sbjct: 5 LVAGLLGVESGQDAVIRALLYERRALKVHPYGNSVAKFTNRISTLRNKLGGSGLKDEGLK 184
Query: 228 ----KQAEGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNGNIAKS 283
+ AEG+ G + + YS+ Y RSPEEILRIVYG GDEHVPGGF+PKG G IA S
Sbjct: 185 VPKLQGAEGETEGNILAGDKYSMAYARSPEEILRIVYGGGDEHVPGGFYPKGAKGRIATS 364
Query: 284 YLHPN 288
YLH N
Sbjct: 365 YLHSN 379
>AV774744
Length = 439
Score = 117 bits (294), Expect = 2e-27
Identities = 68/114 (59%), Positives = 73/114 (63%), Gaps = 1/114 (0%)
Frame = -1
Query: 180 GLLGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRNDLG-KKQAEGKVVGQV 238
GLLGAKSGQDAMIRTLLYERRYLQVKPY WTVYEVTH SLLRNDLG + Q K+ +
Sbjct: 439 GLLGAKSGQDAMIRTLLYERRYLQVKPYPWTVYEVTHPPSLLRNDLGPETQLREKL*VKC 260
Query: 239 FGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNGNIAKSYLHPNAPAP 292
L L + + F KGGNGNIAKSYLHPNAPAP
Sbjct: 259 LVLTCIHLHIQGVQRKF*GLCMDQVMNMYLVASFSKGGNGNIAKSYLHPNAPAP 98
Score = 92.8 bits (229), Expect = 6e-20
Identities = 43/46 (93%), Positives = 44/46 (95%)
Frame = -3
Query: 227 KKQAEGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFF 272
+ AEGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFF
Sbjct: 296 RNPAEGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFF 159
>TC11835 similar to UP|Q9LG00 (Q9LG00) F20N2.10, partial (35%)
Length = 769
Score = 27.7 bits (60), Expect = 2.4
Identities = 13/27 (48%), Positives = 17/27 (62%)
Frame = +2
Query: 114 LLNISKEAFAEVINSAFGKPLNPPFDP 140
L N+ E F +V SA GKP + PF+P
Sbjct: 170 LPNLGHEKFIKVSGSAEGKPRSYPFNP 250
>BP042669
Length = 435
Score = 26.9 bits (58), Expect = 4.1
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = -3
Query: 49 FYFYGTLGWGLDHADPKLAQGGPPPVGGQYAY 80
FYF+ T +G H+DP G P P YA+
Sbjct: 409 FYFFPTTSYGRLHSDP----GSPTPTVVPYAH 326
>AV780089
Length = 579
Score = 26.6 bits (57), Expect = 5.3
Identities = 12/30 (40%), Positives = 22/30 (73%), Gaps = 4/30 (13%)
Frame = -2
Query: 2 SLHTSSIFF----LLSSLVLSLFIPIFCSF 27
++ TSS+FF +L++++LSL + + CSF
Sbjct: 248 AIFTSSLFFKPNPILNNMLLSLVLVLLCSF 159
>AV426967
Length = 402
Score = 26.6 bits (57), Expect = 5.3
Identities = 8/23 (34%), Positives = 17/23 (73%)
Frame = -1
Query: 9 FFLLSSLVLSLFIPIFCSFEDVV 31
FF+L S +++F+P CS+++ +
Sbjct: 237 FFILRSQNINIFLPFHCSYDEFI 169
>TC17643
Length = 513
Score = 26.2 bits (56), Expect = 6.9
Identities = 19/65 (29%), Positives = 27/65 (41%), Gaps = 12/65 (18%)
Frame = +1
Query: 200 RYLQVKPYQWTVYEVTHRLSLLRNDLGKK------------QAEGKVVGQVFGLNMYSLT 247
R+L P W+ EV + +LL + K G + GQ + N+YSL
Sbjct: 133 RWLFKLPEHWSQQEVHSKQALLSLPMDPKLPLKFPT*TGISDPTGYLFGQEYDSNLYSLC 312
Query: 248 YPRSP 252
RSP
Sbjct: 313 ATRSP 327
>TC15328 weakly similar to UP|Q9LP71 (Q9LP71) T1N15.15, partial (50%)
Length = 1325
Score = 26.2 bits (56), Expect = 6.9
Identities = 14/33 (42%), Positives = 16/33 (48%), Gaps = 3/33 (9%)
Frame = +3
Query: 261 GSGDEHVPG---GFFPKGGNGNIAKSYLHPNAP 290
GSG + VPG G FP G Y+ PN P
Sbjct: 651 GSGSDLVPGPPAGVFPSSGPSIGGSMYVGPNDP 749
>TC11294
Length = 501
Score = 25.8 bits (55), Expect = 9.1
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = +2
Query: 95 QQAGHLRAITEEVKGFPRPLLNISKEAF 122
QQAGHL+ + E++ +P N+S A+
Sbjct: 116 QQAGHLQRMEFEIQTCKQPFPNLSSMAY 199
>TC17514 weakly similar to UP|O48683 (O48683) F3I6.9 protein, partial (9%)
Length = 582
Score = 25.8 bits (55), Expect = 9.1
Identities = 21/75 (28%), Positives = 37/75 (49%)
Frame = -1
Query: 113 PLLNISKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGV 172
P+L+I K+ F +V A L F N L LA ++L + + G++ + + +
Sbjct: 297 PILSIMKD-FLKVGRGAGSGLLGSKFRDLCNGL---LATFLLSSLEVIGFLDKDGEFPFL 130
Query: 173 RTRNLVSGLLGAKSG 187
V+GLLG ++G
Sbjct: 129 VEEADVNGLLGVEAG 85
>TC15619 similar to UP|Q8GTX9 (Q8GTX9) Cell cycle control crn (Crooked neck)
protein-like, partial (26%)
Length = 822
Score = 25.8 bits (55), Expect = 9.1
Identities = 13/48 (27%), Positives = 22/48 (45%)
Frame = -2
Query: 5 TSSIFFLLSSLVLSLFIPIFCSFEDVVDVDLVQFHLNLEFLEAEFYFY 52
+SSI SL L + P F + + +QF ++ + +FY Y
Sbjct: 512 SSSISLR*RSLQLCILAPSIAIFRRPLQIFFIQFSASINVTKLQFYLY 369
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,227,342
Number of Sequences: 28460
Number of extensions: 72901
Number of successful extensions: 341
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 337
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 340
length of query: 293
length of database: 4,897,600
effective HSP length: 90
effective length of query: 203
effective length of database: 2,336,200
effective search space: 474248600
effective search space used: 474248600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0333b.7


